A Systematic Review of Federated Learning: The Cure for Isolated Health Data

Federated Learning in Healthcare: Access 80% More Data Without Privacy Risks
Federated learning in healthcare is a privacy-preserving approach that trains AI models across multiple institutions without centralizing patient data. Instead of moving sensitive records, the model travels to the data—learning locally and sharing only encrypted updates. This addresses the tension between the need for large-scale training data and the necessity of patient confidentiality. For decades, the medical field has struggled with the “Data Paradox”: the more data we need to train accurate AI, the more risk we incur by aggregating it. Federated learning (FL) resolves this by ensuring that the raw data remains behind the hospital’s firewall, while the collective intelligence of the network grows.
Key benefits of federated learning in healthcare:
- Privacy by design: Raw data never leaves the institution, reducing breach risks and eliminating the need for risky data transfers.
- Regulatory compliance: Meets GDPR, HIPAA, and the EU AI Act requirements by avoiding cross-border data transfers and maintaining data residency.
- Collaborative AI: Enables multi-institutional research without lengthy data-sharing agreements (DSAs) that can often take years to negotiate.
- Better models: Access to diverse datasets improves accuracy and generalizability across different ethnicities, age groups, and clinical settings.
- Faster deployment: Eliminates bottlenecks related to data centralization, de-identification, and the complex ETL (Extract, Transform, Load) processes required for central repositories.
Traditional healthcare AI is hindered by fragmented data. For decades, medical records have been locked in isolated silos due to legal barriers, technical incompatibilities, and institutional competition. Historically, the transition to digital systems in the 2000s—accelerated by the HITECH Act in the US—often worsened this by creating proprietary “walled gardens.” These systems were designed for billing and local clinical care, not for large-scale machine learning. Consequently, researchers often had to rely on small, single-center datasets, leading to models that performed well in one hospital but failed in another—a phenomenon known as “overfitting” or “algorithmic bias.”
Federated learning flips the script. Research across diverse studies shows federated models achieve 99% of centralized model quality while keeping data at its source. A landmark study involving 10 pharmaceutical companies created a massive chemical compound library through collaboration that was previously unthinkable due to intellectual property concerns. This allows institutions to maintain “data sovereignty” while contributing to collective intelligence. By decentralizing the training process, we are finally able to tap into the 80% of healthcare data that is currently “dark” or inaccessible to the broader research community. This “dark data” often includes longitudinal patient records, rare disease presentations, and diverse genomic sequences that are too sensitive to move but too valuable to ignore.
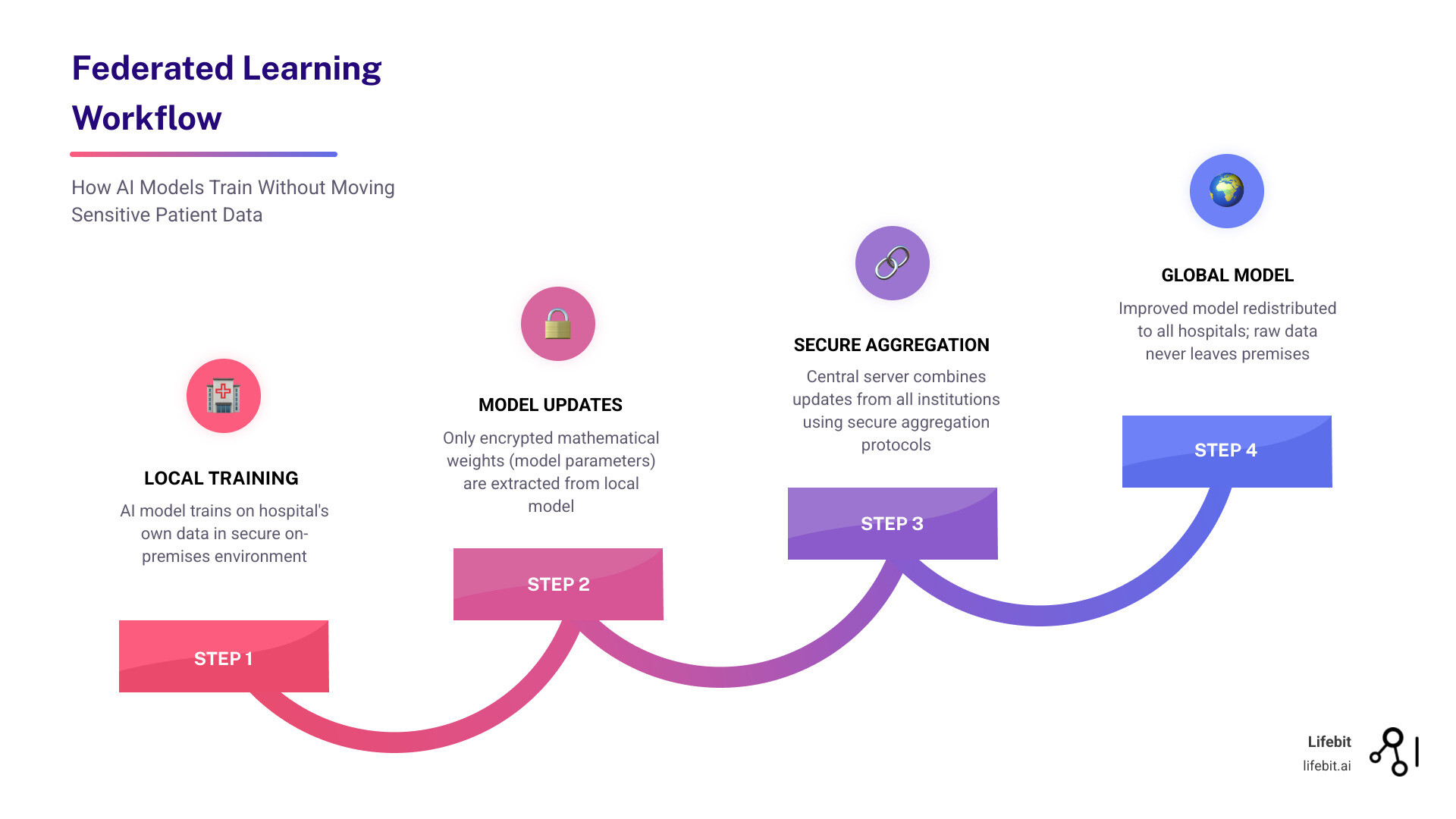
The technical process is elegant in its simplicity: hospitals train local model copies on their own hardware and share only encrypted mathematical weights (gradients) with a central aggregator. The global model improves iteratively without any institution seeing another’s records. Privacy techniques like differential privacy ensure that even model updates cannot leak sensitive information. This iterative cycle continues until the model reaches a desired level of accuracy, at which point the final, robust model is shared with all participants. This architecture effectively solves the “Cold Start Problem” for smaller clinics, allowing them to benefit from high-quality AI models that were trained on the vast datasets of larger academic medical centers without ever needing to possess that data themselves.
Real-world applications are already thriving. The Federated Tumour Segmentation (FeTS) initiative involves 30 international institutions improving brain tumor detection. The HealthChain project predicts cancer treatment responses without images leaving their home hospitals. During the COVID-19 pandemic, researchers developed severity prediction models in just two months using multimodal data that never crossed institutional boundaries. These examples demonstrate that FL is not just a theoretical framework but a practical solution for the most pressing challenges in modern medicine.
As CEO of Lifebit, I have spent 15 years building tools that make federated learning practically deployable. The shift from isolated data to collaborative intelligence is solving problems that have stalled medical AI for decades, allowing us to move from reactive care to proactive, precision medicine. By creating a secure bridge between data owners and data consumers, we are unlocking the potential of global health data to save lives at an unprecedented scale.
Cut Data Sharing Delays by 12 Months with Federated Compliance
Traditional Collaborative Data Sharing (CDS) requires shipping raw data to a central server, which is technically difficult and a regulatory nightmare. Data Use Agreements (DUAs) can take over a year to finalize, delaying critical research. Furthermore, the EU AI Act mandates rigorous governance for high-risk healthcare AI, which centralized models struggle to meet without compromising anonymity. Centralized repositories also create a “honeypot” effect, where a single security breach can expose millions of patient records, leading to catastrophic legal and reputational damage. In a federated setup, the risk is distributed; even if one node is compromised, the attacker only gains access to a fraction of the total network’s data, and the global model remains protected by encryption.
Federated learning in healthcare enables federated data analysis where analysis happens on-site. This respects “Data Gravity”—the principle that large datasets are easier to process where they reside. In genomics, where individual sequences can exceed 100GB, moving data is often physically impossible or prohibitively expensive due to egress fees and bandwidth limitations. By keeping data at the source, institutions also simplify HIPAA compliance, as the data never enters “transit” in a way that would require complex de-identification protocols. This is particularly relevant for the European Health Data Space (EHDS), which seeks to facilitate the cross-border exchange of health data while maintaining strict privacy controls.
| Feature | Centralized Machine Learning | Federated Learning |
|---|---|---|
| Data Location | Moved to a central repository | Stays at the local institution |
| Privacy Risk | High (Data can be intercepted or leaked) | Low (Only mathematical weights are shared) |
| Compliance | Difficult (Complex transfers, GDPR/HIPAA) | Easier (Aligns with residency laws) |
| Data Diversity | Limited by movement and cost | Massive (Can scale to 100+ hospitals) |
| Model Quality | Baseline (Gold standard for accuracy) | ~99% of centralized quality |
| Security | Single point of failure | Distributed security architecture |
This shift is essential for precision medicine, as highlighted in The future of digital health with federated learning. To understand the supporting infrastructure, explore what is a federated data platform.
Overcoming the Privacy-Utility Trade-off
One of the biggest hurdles in medical research is the trade-off between data utility (how useful the data is for training) and patient privacy. Advanced privacy-preserving technologies (PPTs) prevent “membership inference attacks” where models might be reverse-engineered to reveal whether a specific patient’s data was used in the training set. This is a critical concern for pharmaceutical companies who must protect the identity of clinical trial participants.
- Differential Privacy (DP): This is a mathematical framework that adds a calculated amount of “noise” to the model updates. By doing so, it ensures that the presence or absence of a single individual in the dataset does not significantly affect the outcome, making it impossible to identify specific patients. This is vital for protecting patients with rare diseases, where even a few data points could be identifying. DP provides a provable guarantee of privacy that is increasingly required by regulatory bodies.
- Secure Aggregation: This protocol ensures the central server only sees the sum of updates from all participating hospitals, never individual contributions. This prevents the aggregator from being a single point of failure or a source of data leakage, as the server never has access to the specific gradients of a single hospital. It uses cryptographic techniques to ensure that the individual updates are only revealed when combined with others.
- Homomorphic Encryption: This allows calculations to be performed on encrypted weights. The server performs “blind” computation, never seeing the underlying numbers. While computationally intensive, it provides the highest level of security for sensitive pharmaceutical research, allowing for the training of models on highly proprietary chemical structures without risk of exposure.
- Trusted Execution Environments (TEEs): These are hardware-level “secure enclaves” (like Intel SGX) that isolate the model training process from the rest of the system. Even the administrator of the hospital’s server cannot see what is happening inside the TEE, providing an additional layer of defense against internal threats.
At Lifebit, we use federated data sharing to guarantee security. Research like Privacy-first Health Research with Federated Learning proves these methods produce scientific results as accurate as traditional methods while maintaining a much higher security posture. This multi-layered approach to security ensures that federated learning is not just a privacy feature, but a comprehensive security framework for the future of medicine.
99% Accuracy, 0% Data Movement: How Federated AI Trains on Private Records
The core of the operation is the aggregation algorithm, which determines how the central model incorporates the learning from each local node. The most common is FedAvg (Federated Averaging), where the server takes a weighted average of the model parameters from multiple hospitals. However, in the real world, hospitals have different computing power and network speeds. To handle these “stragglers” (slower nodes), we use FedProx, which allows for variable amounts of local work and adds a proximal term to the objective function to stabilize the training process. This ensures that the global model is not biased toward hospitals with the fastest internet connections or the most powerful GPUs.
Techniques like FedCurv prevent “catastrophic forgetting,” a common issue where a model loses previous knowledge while adapting to new data from a specific site. We also see the rise of Incentive Mechanisms like “Federated Shapley Values.” These are used to quantify each hospital’s contribution to the final model’s performance, allowing for fair compensation or credit in multi-institutional research collaborations. This is crucial for encouraging participation from smaller institutions that may have high-quality data but fewer resources.
Alternatively, Peer-to-Peer (Decentralized) architectures are gaining traction. These allow hospitals to sync models directly via gossip protocols, removing the need for a central server entirely. This further decentralizes power and reduces the risk of a central point of failure. For more, see our ultimate guide to federated analytics.
Integrating Deep Learning Models
Federated learning is not limited to simple statistics; it supports the most advanced Deep Learning (DL) models currently used in medicine, enabling complex pattern recognition across diverse datasets:
- CNNs (Convolutional Neural Networks): These are the gold standard for medical imaging. Federated CNNs can detect tumors in MRI scans or identify anomalies in X-rays without moving heavy files. By learning from global variations in pathology, these models become much more robust than those trained on a single hospital’s imaging archive. They can account for differences in lighting, contrast, and resolution across different imaging centers.
- RNNs and LSTMs: Recurrent Neural Networks and Long Short-Term Memory networks are used for analyzing EHR sequences. They can predict mortality, sepsis onset, or disease progression over time by identifying patterns in patient timelines, all without sharing the sensitive timelines themselves. This is particularly useful for chronic disease management where long-term patient history is critical.
- Transformers and ViTs (Vision Transformers): These are the latest breakthrough in AI, capable of handling massive datasets and finding complex patterns in pathology slides. Transformers are particularly good at “attention mechanisms,” focusing on the most relevant parts of a medical image or record, and they remain robust across different clinical environments. Their ability to handle multi-modal data (e.g., combining text and images) makes them ideal for comprehensive patient diagnostics.
- GANs (Generative Adversarial Networks): In a federated context, GANs can generate synthetic data to augment local datasets. This is incredibly helpful for training models on rare cases where a single hospital might only have one or two examples. The GAN learns the distribution of the rare disease and creates “fake” but realistic data to help the model learn without exposing the original patient’s identity.
As noted in Federated learning for medical imaging, these models consistently outperform single-site AI by learning from a wider variety of global edge cases, such as rare pathologies or unusual presentations of common diseases. Furthermore, Federated Personalization allows the global model to be fine-tuned for specific local populations, ensuring that a model trained on global data still performs optimally for the specific demographic of a local clinic.
Governance and Ethical Oversight
Technology alone is not enough; it requires human guardrails to remain ethical, transparent, and unbiased. Federated learning governance is typically structured in three layers to ensure accountability:
- Procedural Governance: This involves the contractual agreements, Data Use Agreements (DUAs), and the adherence to FAIR principles (Findable, Accessible, Interoperable, Reusable). It defines who owns the resulting model and how the intellectual property is shared. It also establishes the legal framework for cross-border collaboration.
- Relational Governance: Data Stewards at each hospital oversee the local nodes. They are responsible for ensuring data quality and verifying that the local training process adheres to the institution’s privacy policies. They act as the human link between the technology and the clinical staff, ensuring that the AI’s goals align with patient care.
- Structural Governance: Oversight bodies or consortium boards perform “Model Auditing.” This involves testing the global model for algorithmic bias against protected groups (e.g., ensuring a skin cancer model works equally well on all skin tones) before it is cleared for clinical deployment. This layer is essential for maintaining public trust in AI-driven healthcare.
Our complete guide to federated governance explains how these layers ensure compliance with laws in the USA, Europe, and beyond, creating a framework of trust that is essential for large-scale medical collaboration.
How 10 Pharma Giants Cut Drug Discovery Time Using Federated AI
Federated learning is no longer a laboratory experiment; it is actively accelerating medical discovery and saving lives. One of the most prominent examples is the MELLODDY consortium (Machine Learning Ledger Orchestration for Drug Discovery). This project involved ten rival pharmaceutical companies, including giants like Janssen and Novartis, training a model on a combined library of 10 million molecules. By using federated learning, they identified drug leads significantly faster while keeping their proprietary chemical structures secret from one another. This set a new benchmark for industry collaboration, proving that even direct competitors can work together for the common good. The project demonstrated that the predictive power of models increases significantly when trained on the combined chemical space of multiple companies. Read more about MELLODDY drug discovery here.
Similarly, the FeTS initiative (Federated Tumour Segmentation) united 30 institutions across several continents to improve brain tumor segmentation. By using the largest and most diverse dataset of glioblastoma cases ever assembled, they built a model that generalizes across different MRI scanner manufacturers and imaging protocols. This is a major step forward, as traditional AI often fails when moved from a Siemens scanner to a GE scanner. The FeTS project showed that federated models could achieve a Dice similarity coefficient (a measure of overlap) that was comparable to models trained on centralized data, but with much higher robustness to site-specific variations. This work is supported by Federated learning in medicine.
In the field of genomics, our work on federated technology in population genomics helps map the human genome across borders. This is critical for identifying rare genetic variants that would otherwise remain hidden in isolated databases. By connecting genomic data from different countries, we can identify the genetic drivers of disease in diverse populations, leading to more equitable healthcare outcomes. For example, a variant that is rare in Europe might be more common in East Asia; federated learning allows researchers to find these patterns without the legal hurdles of moving genomic data across continents.
Case Study: Predicting Treatment Response
The HealthChain project demonstrates the power of the federated data lakehouse in life sciences. By connecting four major French hospitals, researchers were able to predict treatment responses for breast cancer and melanoma using high-resolution histology slides. Training the models locally avoided the massive bandwidth costs and security risks associated with transferring terabytes of image files.
This project specifically solved the “Generalizability Gap,” where AI trained at one site fails at another due to differences in how slides are stained or digitized. By training on data from all four sites simultaneously, the model learned to ignore these “site-specific artifacts” and focus on the actual biological markers of cancer. This efficiency addresses the concerns raised in Diagnosing pharmaceutical R&D decline regarding the escalating costs of drug development and clinical trials. The project resulted in a 15% improvement in prediction accuracy compared to models trained on single-site data.
Additionally, the EXAM model (Electronic Medical Record (EMR) Chest X-ray AI Model) used data from 20 global institutes to predict COVID-19 oxygen needs. Developed in just two weeks during the height of the pandemic, it achieved an Area Under the Curve (AUC) of over 0.94. By combining X-ray images with clinical lab values (like C-reactive protein levels), the model provided clinicians with a powerful tool for triaging patients. This proved that federated learning is not just for long-term research but is a critical component of emergency medical response, allowing the global community to pool resources instantly during a crisis. The EXAM model was deployed across multiple continents, providing a standardized tool for clinicians facing a rapidly evolving virus. This rapid deployment was only possible because the federated framework bypassed the months of legal negotiations typically required for international data sharing.
Federated Learning in Healthcare: 6 Answers on Security, Cost, and ROI
Does federated learning perform as well as centralized machine learning?
Yes. Extensive research shows that federated models typically achieve 99% of the quality of centralized models. In many cases, they actually perform better in real-world scenarios because they are trained on more diverse, representative data. This prevents “overfitting,” where a model becomes too specialized to the specific equipment or patient population of a single hospital and fails when deployed elsewhere. By exposing the model to a wider variety of “noise” and edge cases during training, federated learning creates a more resilient and generalizable AI.
How does FL ensure compliance with GDPR and HIPAA?
Federated learning is often described as “compliance by design.” Since the raw patient data never moves, it doesn’t cross protected borders (a key requirement of GDPR) and remains under the direct control of the covered entity (a key requirement of HIPAA). Institutions maintain full sovereignty over their data, and the sharing of only encrypted, non-identifiable mathematical weights eliminates the need for the aggressive de-identification that often strips data of its clinical utility. This allows researchers to work with high-fidelity data while remaining fully within the bounds of the law.
What are the main technical barriers to deploying FL in hospitals?
There are three primary challenges that must be addressed for successful deployment:
- Communication Costs: Sending large model updates over the internet can be slow. We use techniques like quantization (reducing the precision of the numbers) and sparsification (only sending the most important updates) to reduce update sizes by up to 90%. This allows FL to function even in environments with limited bandwidth.
- Hardware Requirements: Hospitals need sufficient computing power (GPUs) for local training. This is often solved by utilizing secure cloud enclaves or Edge AI devices that can be easily integrated into existing hospital infrastructure without requiring a total overhaul of their IT systems.
- Data Standardization: For a model to learn effectively, data must be mapped to a common format. Using standards like OMOP or HL7 FHIR ensures that the model understands that “heart rate” in one system is the same as “pulse” in another. This process, known as data harmonization, is often the most time-consuming part of the setup but is essential for accuracy.
Who owns the resulting AI model?
Ownership is typically defined by the consortium agreement. In most cases, participating institutions have joint ownership of the global model, while each institution retains 100% ownership and control of its local data. This “shared intelligence, private data” model is a major incentive for hospitals to participate in research, as they can benefit from the collective knowledge of the network without losing their competitive advantage or compromising their patients’ trust.
Can federated learning be hacked?
While FL is significantly more secure than centralized sharing, it is not immune to all risks. Potential threats like “model poisoning” (where a malicious actor tries to influence the model by sending bad updates) exist. We counter this with robust aggregation techniques that can detect and ignore anomalous updates, as well as model watermarking to track the provenance of the AI. Additionally, the use of Secure Multi-Party Computation (SMPC) ensures that even the central aggregator cannot see the individual updates from each node.
How does FL handle different data qualities across hospitals?
We use weighted aggregation, which gives more influence to updates coming from larger or higher-quality datasets. Furthermore, automated quality checks at the local nodes ensure that “garbage in” doesn’t affect the global model. If a hospital’s data is too noisy or poorly formatted, the system can automatically flag it for review before it is used for training. This ensures that the global model remains high-quality even when the participating nodes have varying levels of data maturity.
Stop Siloing Data: Start Your Federated Research Environment Today
The era of isolated health data is ending. Federated learning proves we don’t have to choose between security and innovation. At Lifebit, we provide the tools that allow researchers to collaborate without compromise, ensuring that the next generation of medical breakthroughs is built on a foundation of privacy and trust. The transition from centralized to decentralized AI is not just a technical shift; it is a cultural one that prioritizes patient rights while accelerating scientific discovery.
Our platform provides the Trusted Research Environment (TRE) and Trusted Data Lakehouse (TDL) necessary to turn these theories into reality. With our R.E.A.L. (Real-time Evidence & Analytics Layer), biopharma companies and public health agencies can gain insights from global data in seconds rather than years. This infrastructure supports the entire lifecycle of federated learning, from data discovery and harmonization to model training and deployment.
We are moving toward a future where rare diseases are no longer mysteries because patterns are found across a global network of hospitals. The future of healthcare is federated, secure, and collaborative. By integrating federated learning with emerging technologies like Swarm Learning and Edge Computing, we are creating a truly decentralized intelligence network for global health. Start building your federated research environment today and join the network that’s curing isolated health data and accelerating the path to a healthier world for everyone. The potential to transform patient outcomes is within our reach, provided we have the courage to collaborate securely.

