Secure Multi-Omics Platforms That Won’t Make Your Legal Team Cry

Stop Legal Delays: Deploy Compliant Multi-Omics Analysis Tools in Weeks
Compliant multi-omics analysis tools must meet stringent regulatory standards before they can handle genomic, transcriptomic, proteomic, and metabolomic data in clinical or research settings. Here’s what you need to know:
Core compliance requirements:
- GDPR and HIPAA readiness for patient data protection
- FAIR principles (Findable, Accessible, Interoperable, Reusable) for data accessibility
- Security-first infrastructure with encryption, audit trails, and data residency controls
- Federated analysis capabilities that enable cross-institutional collaboration without moving sensitive data
Top platform categories:
- Commercial cloud platforms with built-in compliance and regulatory support
- Federated AI solutions that use privacy-preserving methods and governance to reduce data movement
- Open-source frameworks (e.g., mixOmics and other community tools) requiring additional validation for clinical deployment
Key challenges:
- Data heterogeneity across omics modalities
- Achieving interpretability for clinical decision support
- Balancing privacy protection with analytical power
- Standardizing workflows across distributed datasets
The shift from one-size-fits-all medicine to precision healthcare has created an urgent need for platforms that can integrate diverse molecular data while satisfying legal teams, regulatory bodies, and ethics committees. Many teams report significant time savings from compliant automation in multi-omics workflows, but only when those tools meet strict data governance requirements. Lifebit’s platform, for example, operates within a security-first infrastructure that conforms to privacy and compliance regulations.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where I’ve spent over 15 years building computational biology platforms and contributing to Nextflow, a workflow framework used globally in genomic analysis. My work focuses on making compliant multi-omics analysis tools accessible through federated data platforms that meet regulatory standards while empowering researchers to unlock insights from sensitive biomedical data.
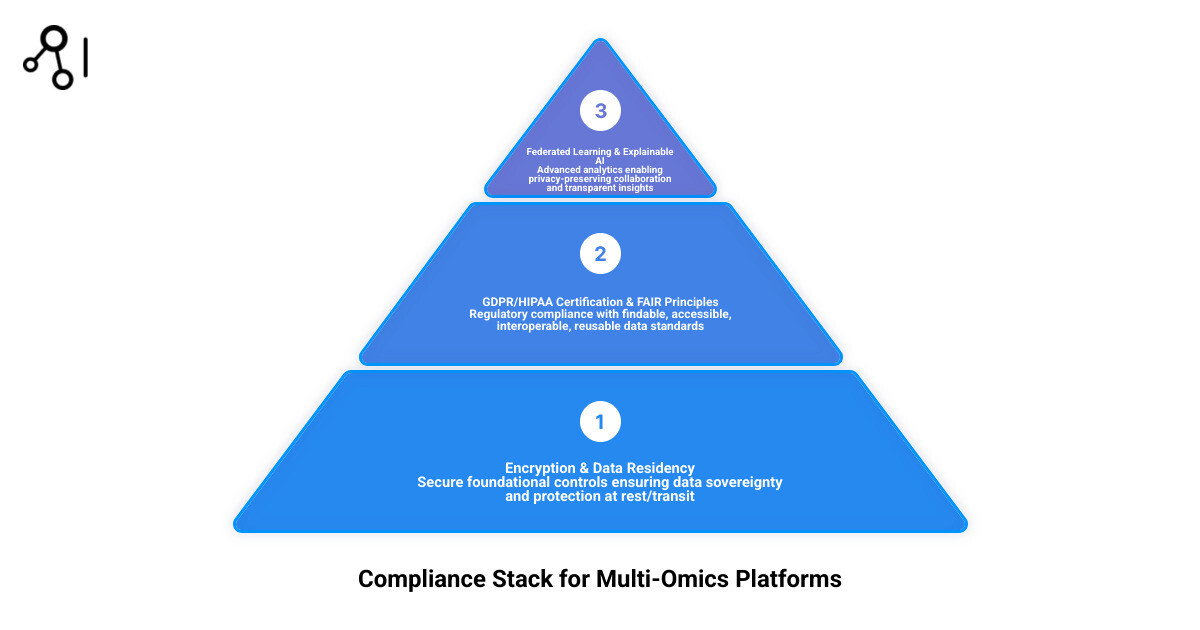
Core Requirements for Compliant Multi-Omics Analysis Tools
When we talk about multi-omics, we aren’t just talking about a bigger spreadsheet. We are talking about the integration of high-throughput datasets that capture various molecular layers—genomics, transcriptomics, proteomics, and metabolomics—within a single biological system. This complexity is a goldmine for science but a potential minefield for data privacy. To be considered “compliant,” a tool must do more than just generate a pretty heatmap. It must adhere to the FAIR Guiding Principles for scientific data management and stewardship, ensuring that data is Findable, Accessible, Interoperable, and Reusable.
The Technical Architecture of FAIR Compliance
In biopharma and clinical research, FAIR principles translate to strict data sovereignty—knowing exactly where your data lives and who can touch it. This requires a robust metadata layer that describes not just the biological content, but the provenance and legal status of the data. For instance, a compliant tool must track the specific consent forms associated with a genomic sample, ensuring that if a patient withdraws consent, the data is automatically flagged and removed from active analysis pipelines. This level of automation is essential when dealing with biobanks containing hundreds of thousands of samples.
Core features must include:
- End-to-End Encryption: Data must be protected at rest using AES-256 standards and in transit via TLS 1.3. This ensures that even if a data packet is intercepted, it remains unreadable to unauthorized parties.
- Granular Audit Trails: Every action, from a simple query to a complex model training session, must be logged for regulatory review. These logs must be immutable, providing a clear history of who accessed what data and for what purpose.
- Automated De-identification: Tools must be able to strip personal identifiers while maintaining the biological signal. This often involves techniques like k-anonymity or differential privacy to ensure that individuals cannot be re-identified through their unique molecular profiles.
As highlighted in Making multi-omics data accessible to researchers, the goal is to create an environment where data can be queried without compromising the underlying privacy of the participants. This is particularly challenging in multi-omics because the combination of different data types (e.g., a rare genotype combined with a specific metabolic profile) can act as a “biometric signature” that is harder to anonymize than a single data type.
Meeting GDPR and HIPAA Standards in Multi-Omics
For organizations operating in the USA and Europe, HIPAA and GDPR are the “Big Two.” These regulations aren’t just suggestions; they are legal mandates with significant financial penalties for non-compliance. In multi-omics, the risk of re-identification is higher because the combination of different “omics” layers can inadvertently create a “molecular fingerprint” unique to an individual.
Compliant tools solve this through security-first infrastructure. For example, Lifebit analyzes data within an environment that conforms to these strict privacy regulations. This involves robust consent management and ensuring that data residency requirements are met—keeping UK data in the UK and US data in the US. Furthermore, under GDPR, the “Right to be Forgotten” requires that tools have the capability to purge specific data points across all integrated omics layers, a task that is computationally intensive without a centralized governance framework.
Ensuring Interoperability and Data Standardization
One of the biggest headaches in omics is that every modality speaks a different language. A transcriptomics file looks nothing like a mass spectrometry output for proteomics. Standardization is the bridge. We look for tools that support:
- Metadata Tagging: Powering swift retrieval of analysis-ready datasets through standardized ontologies like SNOMED CT or HPO.
- Common Coordinate Frameworks: Essential for spatial and single-cell data integration, allowing researchers to map different omics layers onto the same physical or biological space.
- HDF5-based Formats: Utilizing scalable backends like HDF5Array or mudata to handle massive datasets efficiently without loading everything into RAM.
According to computational principles in single-cell data integration, achieving interoperability is the only way to move from raw “reads” to meaningful biological insight. Without these standards, multi-omics remains a collection of silos rather than a unified view of human health.
Trusted Platforms for Regulatory-Grade Multi-Omics Integration
The market for compliant multi-omics analysis tools is maturing, moving away from “black box” solutions toward transparent, interoperable platforms. These tools are designed to break down data silos, creating a “single source of truth” for multimodal research. In the past, researchers would download data to local servers, a practice that is now a major security risk and often a violation of data use agreements.
The Evolution of Trusted Research Environments (TREs)
At Lifebit, we specialize in this space by providing a Trusted Research Environment (TRE) that allows researchers to bring their analysis to the data, rather than moving sensitive data to the analysis. This “data-room” approach is the gold standard for maintaining compliance while enabling high-performance computing. A TRE provides a secure, walled garden where data is stored and analyzed. Researchers are granted access to specific datasets and tools, but they cannot export raw data; they can only export the results of their analysis (e.g., summary statistics or aggregate charts) after a disclosure control review.
This architecture solves several problems simultaneously:
- Security: Data never leaves the secure perimeter of the host institution.
- Governance: Every action is monitored and logged in real-time.
- Collaboration: Multiple institutions can work on the same dataset within the TRE without the need for complex data transfer agreements.
Scaling with Compliant Multi-Omics Analysis Tools
Scaling is where many tools fail. It’s easy to analyze ten samples; it’s incredibly hard to analyze ten thousand across three different continents. Advanced platforms offer tiered architectures to handle this. Lifebit’s platform, for instance, enables custom workflows and pathway analysis integration while conforming to privacy laws. These platforms use reproducible data pipelines to ensure that an analysis run today will yield the same results if audited two years from now. This is achieved through containerization (e.g., Docker or Singularity) and workflow languages like Nextflow or WDL, which package the code, dependencies, and environment together.
High-Throughput Precision Medicine Solutions
For precision medicine, speed is a life-or-death metric. We utilize Trusted Data Lakehouses to integrate omics, imaging, and real-world data (RWD) effortlessly. A Data Lakehouse combines the low-cost storage and flexibility of a data lake with the performance and ACID (Atomicity, Consistency, Isolation, Durability) transactions of a data warehouse. This allows for:
- Automated QA/QC: Using tools like MultiQC to aggregate results across 150+ tools in a single report, ensuring that only high-quality data enters the analysis pipeline.
- Clinical Decision Support: Linking multi-omic results to curated knowledgebases (like ClinVar or COSMIC) for immediate insights. This allows clinicians to see not just a mutation, but the evidence-based treatment options associated with it.
- Longitudinal Tracking: Integrating electronic health records (EHR) with omics data to see how a patient’s molecular profile changes over time in response to therapy.
Open-Source vs. Proprietary: The Compliance Gap
The debate between open-source and proprietary tools often comes down to one thing: who is responsible if something goes wrong? In a research setting, a bug in a script might lead to a retracted paper; in a clinical setting, it could lead to a wrong diagnosis.
| Feature | Open-Source (e.g., mixOmics) | Proprietary Platforms | Federated (e.g., Lifebit) |
|---|---|---|---|
| Compliance Support | Manual / Community | Built-in / Guaranteed | Built-in / Federated |
| Cost | Free (License) | High Subscription | Value-based |
| Clinical Usability | Requires Bioinfo Team | User-friendly UI | Expert-guided / Scalable |
| Data Security | User-managed | Platform-managed | Zero-trust / Federated |
| Validation | Self-validated | Vendor-validated | Continuous Validation |
While Bioconductor provides excellent software for multi-omics integration, the burden of maintaining GDPR/HIPAA compliance falls entirely on the research institution. This includes managing server security, patching software vulnerabilities, and ensuring that the data handling practices meet legal standards.
Popular Open-Source Frameworks for Research
For exploratory research where data isn’t yet “clinical,” open-source is king. These tools allow for maximum flexibility and customization:
- mixOmics: An R package that excels in feature selection and multiple data integration. It’s particularly good for datasets with many variables but few samples, using multivariate methods like PLS (Partial Least Squares).
- MUON: A multimodal omics Python framework designed for single-cell analysis. It allows for the integration of CITE-seq, RNA-seq, and ATAC-seq data in a single object.
- ASTERICS: A web-based interface for exploratory analysis, making complex statistical methods accessible to biologists without needing deep programming expertise.
- Seurat: While primarily for single-cell RNA-seq, its multimodal capabilities (WNN – Weighted Nearest Neighbor analysis) have set the standard for integrating different omics layers at the single-cell level.
Challenges in Open-Source Clinical Deployment
The “compliance gap” in open-source tools usually stems from a lack of version control and formal validation. In a clinical trial, you cannot have your analysis tool update its dependencies mid-study, as this could change the results and invalidate the trial. Best practices for RNA-seq analysis emphasize that reproducibility is a prerequisite for compliance.
Furthermore, open-source tools often struggle with “missing values” and data heterogeneity. In multi-omics, it is common for a patient to have genomic data but be missing proteomic data. Handling these gaps in a statistically sound and compliant way requires significant manual intervention from a bioinformatics team. Proprietary and federated platforms often include automated imputation and normalization routines that are pre-validated for clinical use, reducing the risk of human error.
Overcoming Integration Challenges with AI and Federated Learning
The “No Free Lunch” theorem in machine learning applies heavily to multi-omics: no single model works for every task. The sheer dimensionality of the data—where you might have 20,000 genes, 500,000 methylation sites, and thousands of proteins for only a few hundred patients—creates a “p >> n” problem (more features than observations). This is where modern AI approaches can help. Benchmarking studies across large numbers of experiments have shown that deep learning models, particularly Variational Autoencoders (VAEs) and Graph Neural Networks (GNNs), can outperform classical ML in some settings, especially when dealing with complex disease biology and highly heterogeneous datasets.
Explainable AI (XAI) in Compliant Multi-Omics Analysis Tools
Your legal team—and regulators—will hate a “black box.” If an AI predicts a patient won’t respond to a drug, it must explain why. In the context of multi-omics, this means identifying which specific biomarkers (e.g., a combination of a specific SNP and a protein expression level) drove the prediction.
Compliant multi-omics analysis tools should support explainability methods (for example, feature attribution approaches such as Integrated Gradients or SHAP values) so that biomarker discovery is grounded in biological signal and can be reviewed during audit. This is not just a regulatory requirement; it is a scientific one. Without explainability, it is impossible to validate the AI’s findings in a laboratory setting or to understand the underlying mechanism of disease.
Federated Data Access Without Patient Risk
Federated learning is the “holy grail” of compliant multi-omics analysis tools. It allows institutions to collaborate without actually sharing raw data. This is critical for rare disease research, where no single hospital has enough patients to achieve statistical significance.
- Federated architectures: Privacy-preserving designs that keep data inside each organization’s firewall. The model is sent to the data, trained locally, and only the updated model weights are sent back to a central server.
- Privacy-preserving computation: Techniques such as differential privacy add “noise” to the model updates to ensure that no individual’s data can be reconstructed from the aggregate model. Secure Multi-Party Computation (SMPC) allows different parties to jointly compute a function over their inputs while keeping those inputs private.
Research in single-cell multimodal omics shows that secure collaboration across institutions is essential. For example, by using federated learning, a consortium of hospitals can train a model to predict cancer recurrence using multi-omics data from thousands of patients without any sensitive data ever crossing a state or national border. This bypasses the need for massive data transfer and the associated legal and security risks.
Real-World Impact: Precision Oncology and Rare Diseases
Compliant tools are already changing lives. In oncology, the ability to integrate upstream data processing (raw sequencing) with downstream compliant analysis (clinical interpretation) allows for “real-time” monitoring of treatment response. This is particularly important for liquid biopsies, where multi-omic analysis of circulating tumor DNA (ctDNA) and proteins can detect cancer recurrence months before it shows up on a scan.
A prime example is NYU Langone’s MOSDIR (Multi-Omics Study Design & Data Integration Resource). They provide a structured pathway for researchers to ensure their work is compliant from day one:
- Upstream: Partner with specialized labs for raw sequencing, proteomics, or metabolomics, ensuring that data is captured in standardized formats.
- Downstream: Use MOSDIR for statistical modeling and multi-omics integration, utilizing validated pipelines that meet institutional security standards.
This rigorous approach ensures that study findings are both biologically sound and legally defensible. You can see their request form here. Such resources are becoming the blueprint for academic medical centers worldwide.
Case Study: Accelerating Rare Disease Diagnostics
In the field of rare diseases, patients often go through a “diagnostic odyssey” lasting years. Compliant multi-omics tools are shortening this path. By integrating whole-genome sequencing with transcriptomics (RNA-seq), researchers can identify “splicing variants” that are invisible to DNA sequencing alone.
Using a federated approach, researchers can compare a rare disease patient’s multi-omic profile against global databases like the Undiagnosed Diseases Network (UDN) or the Deciphering Developmental Disorders (DDD) project. Because these tools are compliant, they can handle the sensitive data of pediatric patients with the highest level of protection, ensuring that the search for a cure does not compromise the patient’s future privacy.
Future Regulatory Frameworks for AI-Driven Omics
We are seeing a global shift toward standardized policy. The “Wild West” era of omics is ending. Emerging frameworks focus on:
- Global Data Sharing Standards: Organizations like the Global Alliance for Genomics and Health (GA4GH) are developing protocols for secure, cross-border research.
- AI Accountability: New laws, such as the EU AI Act, will require high-risk AI tools (including those used in clinical decision-making) to meet strict standards for transparency, accuracy, and human oversight.
- IVDR Compliance: In Europe, the In Vitro Diagnostic Regulation (IVDR) is changing how software used in diagnostics is classified, requiring more rigorous clinical evidence and technical documentation.
The power of multimodal omics will only be fully realized if we can maintain the public’s trust through absolute transparency and compliance. As we move toward a future of “digital twins” and personalized health forecasts, the tools we use to analyze our most sensitive data must be as robust as the science they support.
Frequently Asked Questions about Compliant Multi-Omics
What are the most important security standards for omics data?
The gold standards are ISO 27001 for information security management and ISO 27701 for privacy information management. These should be implemented alongside GDPR (Europe) and HIPAA (USA) for personal health information. For multi-omics specifically, adherence to FAIR principles is crucial for data integrity, and SOC2 Type II certification is often required for cloud-based platform providers to prove their operational security.
Can open-source tools be used for clinical trials?
Yes, but they must be “frozen” in a validated environment. This typically involves using a Docker container where the exact version of the tool and all its dependencies are locked. This environment must then undergo a formal validation process (Installation Qualification, Operational Qualification, and Performance Qualification – IQ/OQ/PQ) to ensure it produces consistent results. Many organizations prefer proprietary or federated platforms for clinical trials because they provide these audit trails and validation packages out-of-the-box, saving months of bioinformatics work.
How does federated learning protect patient privacy?
Federated learning works by sending the “algorithm to the data.” Instead of Hospital A sending patient genomes to a central server, the AI model travels to Hospital A, learns from the data locally, and only sends back “mathematical weights” (summaries of what it learned). No individual patient data ever leaves the hospital’s firewall. When combined with differential privacy, it becomes mathematically impossible to reverse-engineer the original patient data from the model updates.
What is the difference between a Data Lake and a Data Lakehouse in multi-omics?
A Data Lake is a vast repository of raw data in its native format (e.g., FASTQ or BAM files). While flexible, it can become a “data swamp” without proper governance. A Data Lakehouse adds a layer of structure and management, allowing for high-performance SQL queries, versioning, and fine-grained access control. For compliant multi-omics, a Lakehouse is preferred because it allows for the rigorous data governance required by regulators while maintaining the ability to store diverse data types.
Conclusion
The era of siloed, non-compliant research is over. To stay competitive and—more importantly—to protect patient trust, organizations must adopt compliant multi-omics analysis tools that prioritize security without sacrificing scientific power.
At Lifebit, we are proud to lead this charge. Our federated AI platform, including the Trusted Research Environment and Trusted Data Lakehouse, is built specifically to handle these complexities. We enable biopharma and public health agencies to access global data in real-time, ensuring that the next breakthrough in precision medicine is both fast and legally sound.
If you’re looking to streamline your biopharma workflows, you might also want to Explore Eureka LS for AI-driven efficiency, where research teams are already saving hundreds of hours every month. The future of medicine is multimodal, federated, and—above all—compliant.

