How to stop worrying and love your TRE platform

Why Secure Biopharma Research Data Is Broken — And What Actually Fixes It
Trusted research environment biopharma is the answer to one of the most costly problems in drug development today: sensitive genomic, clinical, and real-world data locked in silos, wrapped in compliance requirements, and impossible to analyze at scale.
Here is what you need to know quickly:
| Problem | What a TRE Solves |
|---|---|
| Data can’t leave the hospital or country | Analysis runs where data lives — no movement required |
| Harmonizing datasets takes 12 months | AI-powered harmonization compresses this to 48 hours |
| Compliance slows every collaboration | Built-in HIPAA, GDPR, FedRAMP, and ISO controls from day one |
| Siloed genomic and clinical data | Federated access connects multi-omics, EHR, and imaging in one secure workspace |
| Reproducibility gaps in research | Audit trails, version control, and code-to-data models ensure full traceability |
A Trusted Research Environment (TRE) is a secure, controlled digital workspace that lets approved researchers analyze sensitive biomedical data without ever moving it. Instead of copying data to a central location — which creates re-identification risks and regulatory headaches — TREs bring the computation to the data.
Over 60% of new drugs now originate externally through partnerships with biotech, academia, and consortia. That means secure, cross-institutional data collaboration isn’t a nice-to-have. It’s the pipeline.
Yet most biopharma organizations are still wrestling with fragmented infrastructure, unstructured records (80% of healthcare data is unstructured), and data harmonization timelines that stretch across an entire year. The result: delayed trials, missed targets, and lost competitive advantage.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit, and I’ve spent over 15 years at the intersection of computational biology, federated data infrastructure, and precision medicine — including foundational work on Nextflow, the genomic workflow framework used by research programs worldwide. My work building federated trusted research environment biopharma solutions has shown me exactly where the bottlenecks are, and more importantly, how to eliminate them.
Simple trusted research environment biopharma word guide:
What is a Trusted Research Environment Biopharma Professionals Actually Need?

When we talk about a trusted research environment biopharma solution, we are describing more than just a “folder in the cloud.” A true TRE is a highly secure, controlled computing infrastructure designed to give approved researchers remote access to sensitive biomedical data while strictly protecting patient confidentiality.
Often referred to as “data safe havens” or “secure data environments,” these platforms serve as a middle ground. They allow the data to remain under the control of the original provider (like a hospital or a national biobank) while giving biopharma researchers the high-performance computing power they need to run complex analyses. According to scientific research on secure computing environments, TREs are essential to meet the growing global need for secure health data sharing.
The Evolution of Data Access in Life Sciences
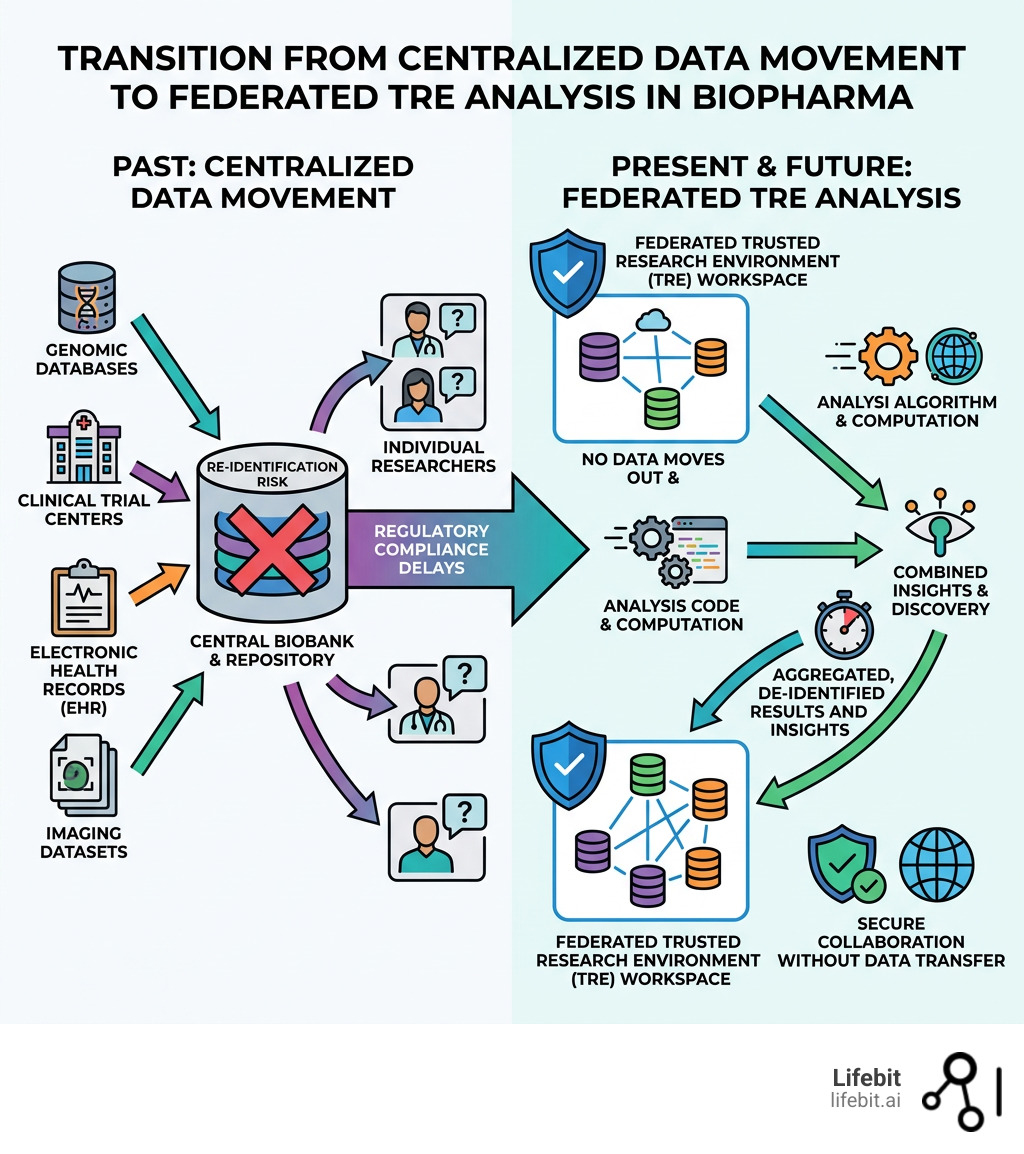
Historically, biopharma research relied on a “data transfer” model. If a pharmaceutical company needed to analyze a cohort of 10,000 patients from a specific hospital system, the data was de-identified, encrypted, and physically or digitally moved to the company’s internal servers. This model is no longer sustainable for three primary reasons:
- Volume: Modern datasets, particularly whole-genome sequencing (WGS) and high-resolution medical imaging, are petabytes in size. Moving them is logistically impossible and prohibitively expensive.
- Sovereignty: Nations are increasingly passing laws that forbid genomic data from leaving their borders. A TRE allows the data to stay in its home country while allowing international researchers to query it.
- Security: Every time data is copied, the attack surface increases. A TRE eliminates the need for copies, keeping the data behind the provider’s firewall.
Key TRE features include:
- Remote Access: Researchers log in via secure portals; data never downloads to their local machines.
- Air-Gapped Analysis: The environment is isolated from the open internet to prevent data leaks.
- Integrated Tooling: Built-in support for R, Python, Jupyter Notebooks, and specialized bioinformatics pipelines.
- Identity Management: Integration with enterprise SSO (Single Sign-On) to ensure only authorized personnel can enter the environment.
TRE vs. Traditional Data Environments
In the old way of doing things, if a biopharma company wanted to analyze a dataset from a partner, they had to move it. This “data movement” model is increasingly dead. Why? Because moving 500,000 whole genome sequences is expensive, slow, and a massive security risk. Once data leaves its original perimeter, the provider loses control over how it is used.
| Feature | Legacy Data Environments | Modern TRE (Federated) |
|---|---|---|
| Data Location | Centralized (Moved to researcher) | Local (Stays with provider) |
| Security Risk | High (Vulnerable during transit/storage) | Low (Data never leaves its home) |
| Cost | High egress and storage fees | Optimized (Compute goes to data) |
| Governance | Manual and fragmented | Automated and unified |
| Scalability | Limited by storage capacity | Unlimited via federated networks |
The Five Safes Framework for Biopharma
To ensure trust, we adhere to the “Five Safes” framework. This isn’t just a checklist; it’s a philosophy for secure TRE governance.
- Safe People: Only researchers who have been vetted and trained are granted access. This often involves institutional agreements and individual background checks.
- Safe Projects: Data use is restricted to ethically approved research aims. Researchers must submit a proposal detailing exactly what they intend to study, which is reviewed by a Data Access Committee (DAC).
- Safe Settings: The environment itself is hardened against unauthorized access and data exfiltration. This includes disabling copy-paste functions, print-screen capabilities, and external internet access within the workspace.
- Safe Data: Researchers only see de-identified or pseudonymized data, reducing re-identification risks. Advanced TREs use k-anonymity and differential privacy to further protect individual identities.
- Safe Outputs: All analysis results are screened (often via an “Airlock”) before they are allowed to be exported, ensuring no patient-level data accidentally slips out. This prevents “re-identification by inference,” where a small cohort size could reveal a patient’s identity.
Why Biopharma R&D Demands a Federated TRE Approach
The future of drug discovery isn’t happening in one lab; it’s happening across a global network. However, data sovereignty laws (like GDPR in Europe or specific genomic data regulations in Singapore) often prevent sensitive health data from crossing borders.
This is where a federated TRE becomes a game-changer. Instead of trying to pull all the world’s data into one “master silo,” we use a federated architecture. This software process allows multiple databases to work together as one. The data stays in its original jurisdiction, but the researcher can query across all of them simultaneously. This is how we enable cross-border collaboration without ever violating data residency requirements.
The Technical Mechanics of Federation
In a federated trusted research environment biopharma setup, the “compute goes to the data.” When a researcher runs a query—for example, looking for a specific genetic variant across three different national biobanks—the query is sent to each local environment. The analysis is performed locally, and only the aggregated, non-identifiable results are sent back to the researcher’s central dashboard. This ensures that the raw, sensitive data never moves, yet the researcher gains the statistical power of a massive, multi-national dataset.
Accelerating Drug Discovery with a Trusted Research Environment Biopharma Solution
Time is the enemy in R&D. Every day a drug spends in development is a day it isn’t helping patients. A dedicated trusted research environment biopharma platform accelerates the pipeline in several ways:
- Target Identification: By accessing diverse, multi-omic datasets (genomics, proteomics, transcriptomics) in a TRE, researchers can identify novel disease markers with higher confidence. This reduces the likelihood of Phase II failures by ensuring the biological target is sound.
- Clinical Trial Optimization: Use real-world evidence (RWE) to design better trials, identify patient cohorts in seconds rather than months, and improve recruitment diversity. By analyzing historical patient data, companies can predict which patients are most likely to respond to a therapy.
- Biomarker Discovery: Seamlessly integrate clinical trial results with external biobank data to find predictive biomarkers for treatment response. This allows for a more personalized approach to medicine, ensuring the right drug reaches the right patient.
Our drug target solutions allow teams to move from a raw variant to a validated target in a fraction of the time traditional methods require.
Solving the 12-Month Data Harmonization Nightmare
The biggest “dirty secret” in biopharma is that 80% of a researcher’s time is spent cleaning data, not analyzing it. Because healthcare data is often unstructured and stored in different formats (EHR, DICOM, VCF), harmonizing it for a meta-analysis usually takes 12 months of manual labor.
Data harmonization involves mapping disparate data fields to a common standard. For instance, one hospital might record “Gender” while another records “Sex,” and a third uses numerical codes. Without harmonization, automated analysis is impossible.
We’ve fixed this. Using AI-powered harmonization, we can transform disparate datasets into standard formats like OMOP (Observational Medical Outcomes Partnership) in as little as 48 hours. This biopharma data integration ensures that when you run a query, you are comparing apples to apples across every site in your network. This speed is critical for responding to emerging health threats or staying ahead in highly competitive therapeutic areas like oncology and rare diseases.
Essential Compliance and Security for Global Research
In trusted research environment biopharma, compliance isn’t just a “nice to have”—it’s the foundation of the business. If a platform can’t prove it’s secure, it can’t be used. The regulatory landscape is a minefield of overlapping requirements that vary by country and data type.
We maintain the highest global standards to ensure secure health data sharing. This includes:
- HIPAA & GDPR: Ensuring patient privacy and data rights are protected in the US and Europe. This includes the “Right to be Forgotten” and strict requirements for data processing agreements.
- FedRAMP: Meeting the rigorous security requirements for US federal government agencies, which is essential for collaborations involving the NIH or VA.
- ISO 27001 & SOC II: Industry-standard certifications for information security management and operational privacy. These certifications require regular third-party audits to verify that security controls are functioning as intended.
- GxP Compliance: Ensuring that the software and processes used in the TRE meet the “Good Practice” guidelines required by regulatory bodies like the FDA for data used in clinical trials.
Comprehensive Audit Trails and Transparency
Every action taken within our TRE is logged. These audit trails provide a transparent record of who accessed what data, what analysis they performed, and what results they attempted to export. This level of granularity is critical for:
- Regulatory Submissions: Providing evidence to the FDA or EMA that data was handled correctly.
- Internal Governance: Monitoring for any unusual activity that might indicate a security breach or policy violation.
- Reproducibility: Allowing other researchers to see exactly how a result was reached, which is the cornerstone of scientific integrity.
Automated Airlocks and Secure Data Export
The most vulnerable point in any research workflow is the “export.” How do you ensure that a researcher taking a graph or a summary table out of the TRE isn’t accidentally taking patient-level data with them? In the past, this was a manual, slow process that created a massive bottleneck.
Traditionally, this required a manual review by a data “gatekeeper,” which could take weeks. Our research efficiency tools include Automated Airlocks. These use privacy-enhancing technologies to automatically scan export requests for sensitive information.
How the Automated Airlock works:
- Request: A researcher submits a file (e.g., a CSV or PNG) for export.
- Scanning: The system checks for PII (Personally Identifiable Information) and runs statistical checks to ensure the data is sufficiently aggregated.
- Thresholding: If the file contains data from a cohort smaller than a pre-defined threshold (e.g., fewer than 10 patients), it is automatically flagged.
- Approval: If the output is safe (e.g., an aggregated statistic), it’s approved instantly. If it’s risky, it’s flagged for human review. This reduces the gatekeeper’s workload by up to 90%, allowing research to move at the speed of thought.
Real-World Impact: From Biobanks to Clinical Innovation
The impact of a trusted research environment biopharma setup is best seen in large-scale population health programs. These initiatives are the engines of modern precision medicine, but they only work if the data is accessible and secure. Platforms like ours manage over 275 million records across major deployments in the UK, Singapore, and for the NIH in the USA.
Case Study: UK Biobank and Large-Scale GWAS
- UK Biobank: This massive resource contains whole genome sequences from 500,000 individuals, linked to their medical records and lifestyle data. Using a TRE allows researchers from around the world to run GWAS (Genome-Wide Association Studies) on this data without the data ever leaving the UK’s secure perimeter. This has led to breakthroughs in understanding the genetic basis of diseases like Type 2 diabetes and Alzheimer’s. Without a TRE, the UK Biobank would have to ship hard drives around the world, a process that would be both insecure and incredibly slow.
Case Study: Rare Disease Research and Global Federation
- Rare Disease Research: Rare disease data platforms use TREs to connect small, disparate patient groups across the globe. For a rare disease that affects only 1 in 100,000 people, a single hospital might only have two patients. By federating data across 50 hospitals in 10 countries, researchers can find 100 cases—enough to power statistically significant studies that were previously impossible. This global connectivity is only possible because the TRE ensures that each hospital maintains control over its own patient data.
The Power of Multi-Omic Integration
Modern pharma research requires multi-omic health data access to understand the “360-degree view” of a patient. This involves linking:
- Genomics: The patient’s DNA sequence.
- Transcriptomics: How genes are being expressed.
- Proteomics: The proteins present in the body.
- Clinical History: Longitudinal data from Electronic Health Records (EHR).
- Imaging: MRI, CT scans, and pathology slides.
By integrating these layers within a trusted research environment biopharma platform, researchers can identify “molecular signatures” of disease. This allows for the development of companion diagnostics, which can predict which patients will respond best to a specific drug, thereby increasing the success rate of clinical trials and improving patient outcomes.
Frequently Asked Questions about Trusted Research Environment Biopharma
How does a TRE handle multi-modal data at scale?
Modern biopharma data software must handle more than just text. Our TRE is built to support imaging (DICOM), clinical notes (EHR), and massive genomic files (VCF/BAM). We use a “Trusted Data Lakehouse” architecture that allows these different data types to be queried together using scalable cloud compute. This means you can correlate a specific genetic mutation with a specific feature on an MRI scan across thousands of patients. The system automatically scales compute resources up or down based on the complexity of the task, ensuring that even the most intensive AI workloads are handled efficiently.
Can a TRE support AI and machine learning workflows?
Yes. In fact, a TRE is the ideal place for AI/ML. Because the environment is secure and high-performance, you can spin up GPUs to train large-scale predictive models directly next to the data. This eliminates the need to move data to a separate AI environment, which is a major security risk.
We also support federated learning, where an AI model “travels” to different data sites, learns from the local data, and then brings the insights back to a central model—all without the raw data ever being shared. This is particularly useful for training models on rare disease data or sensitive clinical trial results where data sharing is strictly prohibited.
What is the difference between a TRE and a Data Clean Room?
While the terms are sometimes used interchangeably, a TRE is generally more robust for scientific research. A “Data Clean Room” is often used in marketing to join two datasets for simple overlap analysis (e.g., matching a customer list with an ad platform).
A trusted research environment biopharma platform, however, provides a full suite of analytical tools (like Nextflow, RStudio, and Jupyter), deep research governance (the Five Safes), and the ability to perform longitudinal tracking over years of patient data. TREs are designed for complex, iterative scientific discovery, whereas Clean Rooms are typically designed for simpler, one-off data matching tasks.
Is a TRE cloud-agnostic?
Yes, a modern TRE should be cloud-agnostic, meaning it can run on AWS, Google Cloud, Azure, or even on-premise servers. This is crucial for biopharma companies that may have data stored across multiple cloud providers or need to comply with specific institutional requirements. A cloud-agnostic approach ensures that the research environment can be deployed wherever the data lives, providing maximum flexibility and avoiding vendor lock-in.
How do TREs improve research reproducibility?
Reproducibility is a major challenge in biomedical research. A TRE solves this by providing a “frozen” environment where the exact versions of software, libraries, and data used in an analysis are recorded. Because every step of the workflow is logged and the environment is standardized, another researcher can re-run the same analysis and achieve the same results. This is essential for validating findings and meeting the transparency requirements of regulatory agencies and top-tier scientific journals.
Conclusion: Choosing the Right Partner for Secure Discovery
The era of centralized data silos is over. To compete in modern drug discovery, biopharma teams need a solution that is flexible, cloud-agnostic, and built for the reality of global, federated data.
At Lifebit, we provide the next-generation federated AI platform that makes this possible. By bringing the analysis to the data, we eliminate the security risks of data movement and the “bottleneck” of manual harmonization. Whether you are identifying new drug targets, optimizing clinical trials, or generating real-world evidence, our Lifebit Trusted Research Environment provides the security, speed, and AI-driven insights you need to turn data into life-saving discoveries.
Stop worrying about data silos and start loving your research again. The data is out there—we just help you reach it safely.

