Why Clinical and Omics Data Platforms are Better Together

Stop Siloing Data: Why Your Omics Data Platform Needs Clinical Integration Now
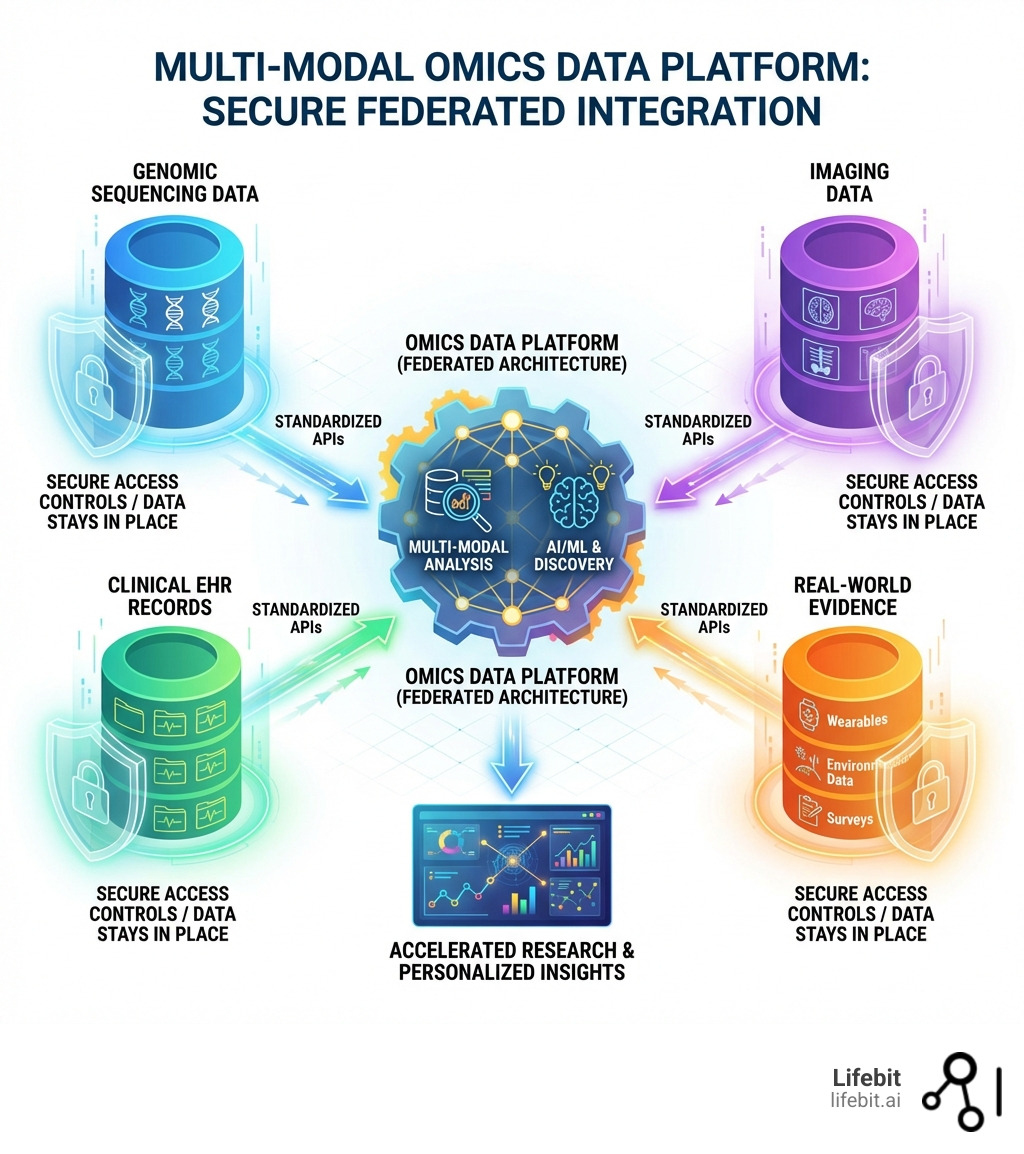
Omics data platform technology has become essential for organizations managing genomic, proteomic, and metabolomic datasets alongside clinical information. These platforms enable researchers to:
- Ingest and harmonize multi-modal datasets (omics, imaging, phenotypic, and real-world data)
- Analyze at scale using cloud-native infrastructure that handles petabytes of data
- Share securely across federated networks without moving sensitive patient information
- Comply with regulations like HIPAA and GDPR through built-in access controls and audit trails
- Accelerate discovery using AI/ML algorithms and foundation models trained on large-scale datasets
The challenge is real. Healthcare organizations face unprecedented data volume—leading platforms now process 80 petabytes of data from 3 million participants across 50 countries. Meanwhile, genomic research generates massive datasets: 1.6 million SARS-CoV-2 genomes have been processed through global federated networks for pandemic surveillance alone. The omics landscape is no longer limited to just DNA sequencing. It encompasses a vast spectrum of biological layers: Genomics (DNA), Transcriptomics (RNA), Proteomics (Proteins), Metabolomics (Metabolites), Epigenomics (Chemical modifications to DNA), and Microbiomics (Microbial communities). Each layer provides a different lens through which we view human health, but the real power lies in their integration.
But volume isn’t the only problem. Data sits in silos. Different formats don’t talk to each other. Privacy regulations restrict movement across borders. And traditional infrastructure can’t keep up with the computational demands of modern AI models like scGPT and Geneformer, which predict drug responses from single-cell omics data. Real-world data (RWD) integration involves pulling from Electronic Health Records (EHR), longitudinal patient registries, and even wearable device data to provide a 360-degree view of the patient journey. Without a unified platform, these disparate data points remain disconnected, preventing the holistic analysis required for true precision medicine.
When clinical and omics data work together, researchers can link genetic variants to real-world patient outcomes, identify therapeutic targets faster, and build more accurate predictive models. Federated networks using GA4GH open standards make this possible without compromising data sovereignty or security. This approach allows for the creation of “virtual cohorts” that span multiple institutions and countries, providing the statistical power needed to study rare diseases and complex traits that were previously out of reach.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where I’ve spent over 15 years building omics data platforms that enable secure, compliant analysis of biomedical data at scale. My work on Nextflow and federated genomic infrastructure has helped researchers worldwide connect diverse datasets while maintaining privacy by design. At Lifebit, we focus on removing the technical barriers that prevent scientists from accessing the data they need, ensuring that the next generation of life-saving treatments can be discovered more efficiently.
Crush the 80-Petabyte Bottleneck with a Scalable Omics Data Platform

When we talk about the sheer scale of modern multi-omics research, the numbers are staggering. We are no longer just dealing with “big data”; we are dealing with “astronomical data.” Industry benchmarks show that leading platforms are now managing upwards of 80 petabytes of data. To put that in perspective, if you tried to store that on standard laptops, you’d need a stack of them miles high. This data explosion is driven by the decreasing cost of sequencing and the increasing resolution of biological assays, such as single-cell sequencing, which generates millions of data points per individual cell.
The primary struggle for many organizations isn’t just storing this data—it’s the complexity. Omics data is high-dimensional, meaning for every single patient sample, there are millions of data points. This creates a significant “noise” problem. Without advanced deep learning methods for multi-omics data integration, finding a meaningful signal in that noise is like trying to find a specific grain of sand in a desert. The “High-Dimensional Low-Sample-Size” (HDLSS) problem is a mathematical nightmare for traditional statistics. In a typical omics study, you might have 100,000 features (genes, proteins) but only 500 patients. This creates a “p >> n” problem where the number of variables far exceeds the number of observations, leading to the “curse of dimensionality” and a high risk of false-positive discoveries.
We see many organizations hitting a wall with legacy infrastructure. Traditional on-premise servers or basic data warehouses simply weren’t built for the computational intensity of modern bioinformatics. This often leads to overfitting in machine learning models, where the AI “memorizes” the noise instead of learning the actual biology. Furthermore, the “technical debt” associated with maintaining legacy systems often consumes more resources than the actual research, slowing down the pace of innovation and increasing the time-to-market for new therapies.
A modern Omics data platform solves this by leveraging cloud-native, distributed processing. By using high-performance compute engines like Apache Spark and container orchestration via Kubernetes, we can run 30 million workflows or analyze 110 million single cells without the system breaking a sweat. Modern platforms utilize “Data Lakehouse” architectures, combining the flexibility of data lakes with the management capabilities of data warehouses. This allows for “schema-on-read” capabilities, where researchers can structure data at the moment of analysis rather than being locked into rigid, pre-defined formats. This scalability ensures that as your biobank grows from thousands to millions of participants, your research speed actually increases rather than slows down.
| Feature | Legacy Storage Systems | Modern Omics Data Platforms |
|---|---|---|
| Capacity | Limited by physical hardware | Virtually infinite cloud scaling |
| Data Types | Primarily structured/clinical | Multi-modal (Genomic, Imaging, RWD) |
| Processing | Sequential/Slow | Distributed/Parallel (e.g., Apache Spark) |
| Interoperability | Siloed, proprietary formats | Open standards (GA4GH, OMOP) |
| AI Readiness | Manual data prep required | Integrated ML/AI stacks |
| Cost Efficiency | High CapEx for hardware | Pay-as-you-go OpEx model |
| Data Governance | Manual and error-prone | Automated and policy-driven |
End Data Migration Nightmares: Use Federated Networks and GA4GH Standards
In the past, if a researcher in London wanted to collaborate with a team in New York or Singapore, they would have to physically move massive datasets. Not only is this slow, but it’s a regulatory nightmare. Today, data is often shared across different jurisdictions, and laws like GDPR and HIPAA make moving sensitive genetic info across borders nearly impossible. The sheer size of these datasets also makes physical transfer impractical; moving a petabyte of data over a standard high-speed connection could take weeks, if not months, and is prone to corruption or interception.
This is where the “Federated” approach changes everything. Instead of moving the data to the researcher, we move the analysis to the data. This protects data sovereignty—the data stays exactly where it was collected, behind the host organization’s firewall, while researchers run their queries remotely. This “orchestration” allows for meta-analysis across global cohorts without the data ever leaving its original jurisdiction. It effectively neutralizes “Data Gravity”—the idea that as datasets grow, they become harder to move and attract more applications and services. By bringing the compute to the data, we can leverage massive, diverse datasets for AI training and discovery without the risk of centralization.
The “glue” that holds this global network together is the Global Alliance for Genomics & Health. By implementing these open standards, an Omics data platform ensures that different systems can “talk” to each other. The GA4GH has developed a suite of protocols that serve as the “internet protocols” for genomics. These include the Workflow Execution Service (WES) for running portable workflows, the Tool Registry Service (TRS) for sharing algorithms, the Data Repository Service (DRS) for accessing data objects across different clouds, and the Beacon protocol for discovering datasets with specific genetic variants.
We’ve seen the power of this in action with initiatives like Viral AI, a global federated network that processed and shared 1.6 million SARS-CoV-2 genomes for pandemic surveillance. This wasn’t just a technical feat; it was a life-saving collaboration that allowed the world to track variants in real-time without compromising national data security. For organizations, adopting GA4GH-compliant genomics infrastructure means they are “plug-and-play” ready for international research consortiums. It transforms a local biobank into a node in a global intelligence network, enabling researchers to query across multiple biobanks simultaneously to find rare disease cohorts that would be impossible to assemble within a single institution.
From Variant to Target: How GenAI and Foundation Models Speed Up Discovery
We are currently witnessing a revolution in how we use next-generation-sequencing data. It’s no longer just about sequencing a genome; it’s about using Generative AI to “read” and “write” the language of biology. Foundation models are being trained on massive multi-omics datasets to predict things we never thought possible. These models are not just looking for simple correlations; they are learning the underlying biological “grammar” that governs how genes interact and how cells respond to their environment.
For example:
- scGPT: A foundation model for single-cell biology that can predict how a cell will respond to a specific drug. It is trained on millions of single-cell profiles, allowing it to perform “zero-shot” learning, where it can predict the behavior of a cell type it hasn’t explicitly seen before.
- Geneformer: A context-aware attention model that helps identify new therapeutic targets by understanding gene network dynamics. It uses a transformer architecture to understand how gene networks are perturbed in disease states, such as cardiomyopathy, providing insights into potential intervention points.
- Protein Synthesis: Companies are now using Large Language Models (LLMs) to generate entirely new synthetic proteins, which could lead to breakthroughs in enzyme engineering and drug design. By understanding the folding patterns and functional domains of proteins, these models can design molecules with specific therapeutic properties from scratch.
However, training these models is computationally expensive and data-hungry. A robust Omics data platform provides the “Data Intelligence” layer needed to manage this. With over 15,000 algorithms already running on top-tier platforms, the infrastructure must be able to handle the “MLOps” (Machine Learning Operations) of these models—tracking everything from data acquisition to the final explainability of the AI’s decision. The MLOps lifecycle in omics is particularly complex, requiring “Data Lineage”—the ability to trace a specific AI prediction back to the raw FASTQ files and the specific version of the alignment algorithm used. Without this, the “black box” of AI becomes a liability in clinical settings where explainability is paramount.
Stop Fighting Your Tools: A Better Omics Data Platform User Experience
One of the biggest problems in bioinformatics has always been the “technical wall.” You shouldn’t need a PhD in Computer Science just to ask a question of your data. Modern bioinformatics-platform designs are now focusing on the “IDE for Science” concept. This includes:
- Natural Language Querying: Imagine asking your data, “Show me all female patients over 50 with this specific genetic variant who responded well to immunotherapy,” and getting the answer in seconds. This uses LLMs to translate human language into complex SQL or Python queries.
- Multi-modal Integration: Weaving together genomics, transcriptomics, and proteomics with imaging and Real-World Data (RWD). This allows researchers to see the phenotypic expression of genetic variants in real-time.
- AI-Powered Assistance: Context-aware AI that helps researchers build pipelines and analyze results without writing a single line of code. This democratizes data science, allowing biologists and clinicians to lead the discovery process.
- Explainable AI (XAI): Providing visual and statistical evidence for why an AI model reached a certain conclusion, which is critical for building trust in clinical decision support systems.
By breaking down these silos, we create a “single source of truth” that allows a biologist, a clinician, and a data scientist to all work on the same dataset simultaneously, each using the tools they are most comfortable with. This collaborative environment accelerates the transition from basic research to clinical application.
Bulletproof Your Research: Essential Security for HIPAA and GDPR Compliance
When dealing with the most sensitive data on earth—human DNA—security cannot be an afterthought. It must be “built-in by design.” To ensure compliance with HIPAA, GDPR, and other international standards, an Omics data platform must provide a multi-layered security stack. This involves not just technical safeguards, but also rigorous governance frameworks that define how data can be accessed and used.
A key part of this is the omop-complete-guide to data harmonization. By mapping disparate clinical data to the OMOP Common Data Model, we ensure that privacy-preserving queries can be run across different datasets consistently. This standardization is essential for federated analysis, as it ensures that a query for “Type 2 Diabetes” returns the same clinical phenotype regardless of which hospital’s database is being queried.
But security is also about control. We look for platforms that offer the “Five Safes” framework, which is the gold standard for data access:
- Safe People: Ensuring only vetted and authorized researchers can access the platform through multi-factor authentication and identity management.
- Safe Projects: Requiring that all research goals are approved by an ethics committee or data access committee before work begins.
- Safe Settings: Utilizing Trusted Research Environments (TREs) where data is analyzed in a secure, isolated cloud environment that prevents unauthorized data extraction.
- Safe Data: Applying de-identification and anonymization techniques to ensure that individual participants cannot be identified from the research datasets.
- Safe Outputs: Screening all research results and aggregate data to ensure that no “residual disclosure” occurs when results are published.
Advanced encryption techniques, such as Differential Privacy and Homomorphic Encryption, are also becoming standard. Differential Privacy adds mathematical “noise” to datasets, ensuring that while overall statistical trends remain accurate for research, it is mathematically impossible to pinpoint an individual participant. Homomorphic Encryption allows for computations to be performed on encrypted data without ever needing to decrypt it, providing the ultimate layer of protection for sensitive genetic information.
The result of these robust controls is a measurable increase in trust. In fact, platforms that prioritize these features have achieved a 76% score on the Decision Confidence Index, meaning researchers can trust the results of their analysis enough to move forward with clinical trials or drug development. Adhering to FAIR principles (Findable, Accessible, Interoperable, and Reusable) isn’t just a best practice; it’s a requirement for modern, reproducible science that can withstand the scrutiny of regulatory bodies like the FDA or EMA.
Omics Data Management: Your Top Questions Answered
What are the primary challenges of an Omics data platform?
The “big three” challenges are volume, variety, and velocity.
- Data Complexity: Omics data isn’t just large; it’s heterogeneous. You have raw sequencing files (FASTQ), processed variants (VCF), clinical notes, and medical images (DICOM). Managing these different formats requires a sophisticated data orchestration layer.
- Standardization: Without common formats like GA4GH and OMOP, data remains in silos. Interoperability is the biggest hurdle to cross-institutional collaboration.
- Regulatory Problems: Navigating the complex web of global privacy laws is a major bottleneck. Platforms must be able to adapt to local residency requirements while still allowing for global analysis.
How do these platforms ensure data reproducibility?
Reproducibility is the bedrock of science. Modern platforms ensure this by using “containerized” workflows (like Nextflow or WDL) where the exact version of every tool and algorithm used is recorded. This is often referred to as “Software Heritage” or “Provenance.” This, combined with rigorous metadata management and scientific research on training machine learning models, ensures that if another scientist runs the same analysis on the same data, they will get the exact same result. This is critical for clinical trials where the methodology must be perfectly transparent and auditable.
Why is the federated approach better for sensitive health data?
Because it solves the “Privacy vs. Progress” paradox. You get the benefits of massive, diverse datasets for AI training and discovery without the risk of moving or centralizing sensitive information. It respects data residency laws, which often mandate that health data cannot leave the country of origin. This makes it the only viable path forward for large-scale international research consortiums, such as those tracking global health threats or studying rare genetic disorders across diverse populations.
How does an Omics data platform handle “Data Gravity”?
Data gravity is the concept that as datasets grow, they become harder to move and naturally attract more applications and services. A modern platform addresses this by bringing the compute to the data. By deploying analysis tools directly into the environment where the data resides (e.g., a national biobank’s cloud), the platform effectively neutralizes the negative effects of data gravity, allowing for rapid analysis without the need for costly and risky data transfers.
What is the role of “Synthetic Data” in omics?
Synthetic data allows researchers to develop and test algorithms on data that mimics the statistical properties of real patient data without containing any actual patient information. This is incredibly useful for the early stages of research and development, as it allows for the testing of pipelines and AI models without the need for the high-level security clearances required for real patient data. Once the model is validated on synthetic data, it can then be deployed into a secure TRE to run on the actual sensitive datasets.
Open up Precision Medicine: Start Your Multi-Modal Research Today
The future of medicine isn’t just genomic; it’s multi-modal. By bringing clinical and omics data together on a single, secure platform, we are finally able to see the full picture of human health. This shift toward “Precision Medicine 2.0” requires a move from reactive treatment to proactive prevention. By analyzing multi-modal data over time, we can identify “biomarkers of transition”—signals that a patient is moving from a healthy state to a disease state years before symptoms appear. This allows for early intervention and personalized treatment plans that are tailored to the individual’s unique biological makeup.
At Lifebit, we are proud to be at the forefront of this shift. The Lifebit Federated Biomedical Data Platform is designed to be the “operating system” for this new era of research. By combining our Trusted Research Environment (TRE) and Trusted Data Lakehouse (TDL) with our Real-time Evidence & Analytics Layer (R.E.A.L.), we empower organizations to move from variant to target in record time. Our platform is built to handle the complexities of modern biology, providing the scalability, security, and interoperability needed to turn massive datasets into actionable insights.
Whether you are a government agency managing a national biobank, a biopharma company looking to accelerate your R&D pipeline, or a clinical research organization seeking to improve patient outcomes, the goal is the same: to turn data into life-saving insights. The integration of AI and foundation models into these platforms is not just a luxury; it is a necessity for keeping pace with the rapid advancements in the field. By adopting a modern Omics data platform, you aren’t just managing data—you are unlocking the potential of precision medicine for patients everywhere, ensuring that the right treatment reaches the right patient at the right time.

