Powering Cancer Cures

How Genomics England's Lancet-published breast cancer breakthrough ran on Lifebit
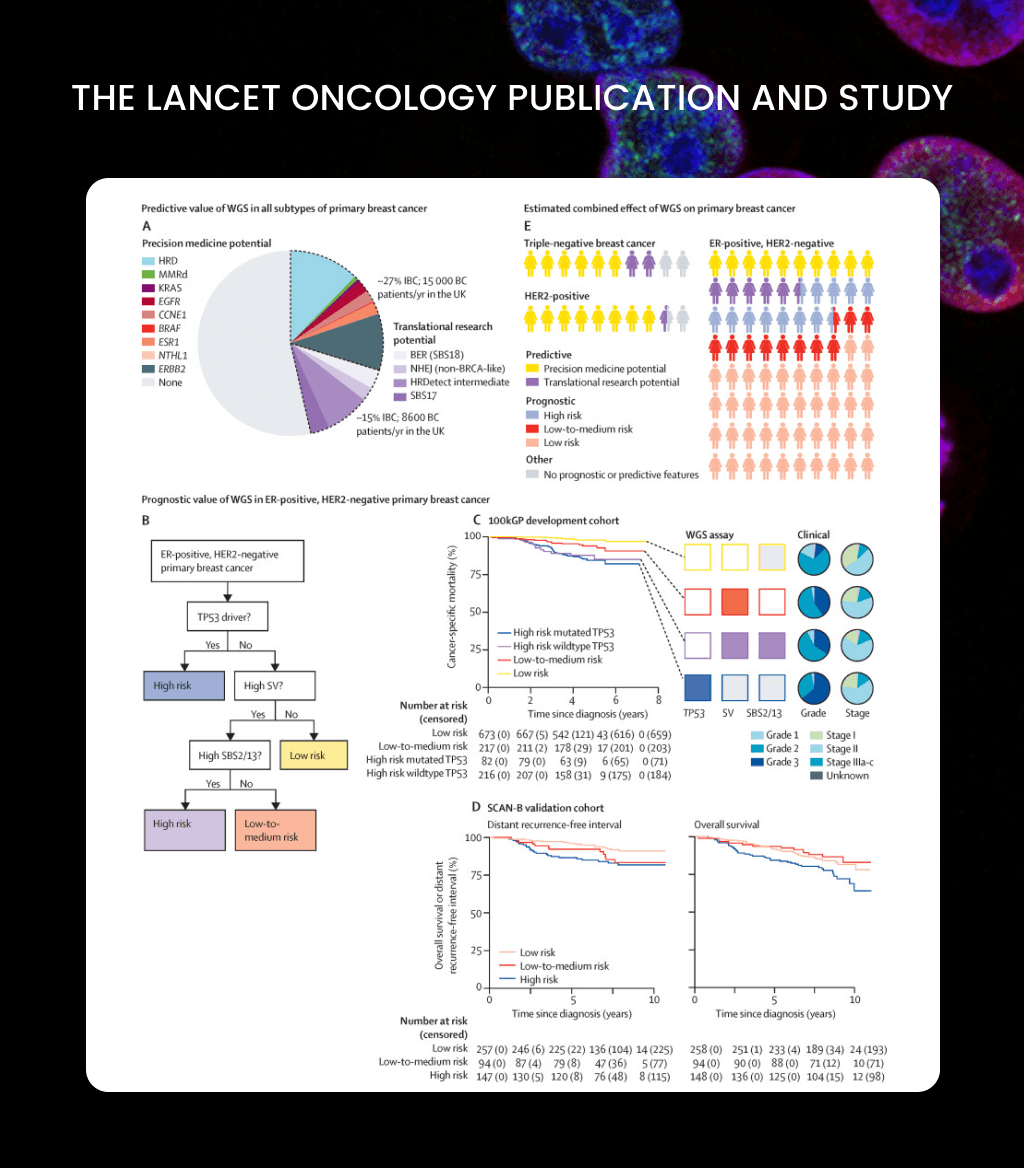
Powering a Lancet Oncology breakthrough across Genomics England, the University of Cambridge, and the NHS — analysing 2,500+ breast cancer genomes inside the research environment, without the data ever leaving the perimeter.
“Lifebit Platform has been a game-changer, driving crucial progress in the breast cancer Research Network.”
Dr Daniella Black
1st author, Lancet Oncology study
Challenges
- Massive data volumes generated per genome
- Complex, multi-stage clinico-genomic analysis pipelines
- High I/O demands and extremely long analysis runtimes
- Escalating compute costs at population scale
- Slow iteration cycles delaying scientific discovery
Outcomes
- Identification of actionable genomic biomarkers in breast cancer
- Improved patient stratification for targeted therapies
- Faster discovery cycles for genomic research teams
- Reduced computational cost for large-scale studies
- Accelerated translation from research insight to clinical application
impact
27%
Breast cancer patients
with actionable genomic findings
70%
Faster runtime
for large-scale genomic analysis workflows
2,500+
Patient genomes
analysed across England
50%
Lower compute cost
for population-scale biomarker discovery
15,000+
Women annually
who could benefit from personalised treatment
FAQs
How a federated platform powered a Lancet Oncology breakthrough
Lifebit’s federated platform underpinned the analytical infrastructure for this Lancet Oncology study — running large-scale clinico-genomic analyses across thousands of NHS breast cancer patients inside Genomics England’s research environment, without moving patient data out of custody.
Inside the study:
Population-scale genomic analysis, inside the perimeter
Whole-genome data from 2,500+ NHS breast cancer patients was analysed inside Genomics England’s secure research environment. Researchers from the University of Cambridge, Genomics England, and the NHS worked against the same governed dataset — no patient-level data ever left custody.
Pipelines that scale with the cohort
Multi-stage clinico-genomic pipelines — alignment, variant calling, annotation, and downstream biomarker analysis — ran as reproducible workflows. Runtimes that would have been prohibitive on traditional infrastructure were cut by 70% through large-scale analysis optimisation.
From research signal to clinical relevance
The study identified actionable genomic findings in 27% of patients analysed, with direct implications for stratification and targeted therapy selection — a result with potential to inform care for the 15,000+ women diagnosed with breast cancer in the UK each year.
Lower cost, broader reach
Compute cost for population-scale biomarker discovery dropped by 50%, freeing budget for the next generation of studies on the same cohort — the same federated infrastructure can now power further translational research without rebuilding the analytical stack.
Next step
Ready to power your next breakthrough?
Lifebit’s federated platform is how research teams turn population-scale genomic data into clinical impact — at Genomics England, the NIH, Boehringer Ingelheim, and beyond. Ready to do the same with yours?

