From OMOP to Salesforce: Navigating the World of Clinical Data Models

Why Healthcare Data Silos Are Costing Research Years—Not Months
Clinical data models are standardized frameworks that define how healthcare information—from patient demographics to lab results and medications—should be structured, stored, and related to enable consistent analysis across different systems and institutions. In the modern era of precision medicine, these models are no longer just technical specifications; they are the essential infrastructure for global health collaboration.
What Clinical Data Models Do:
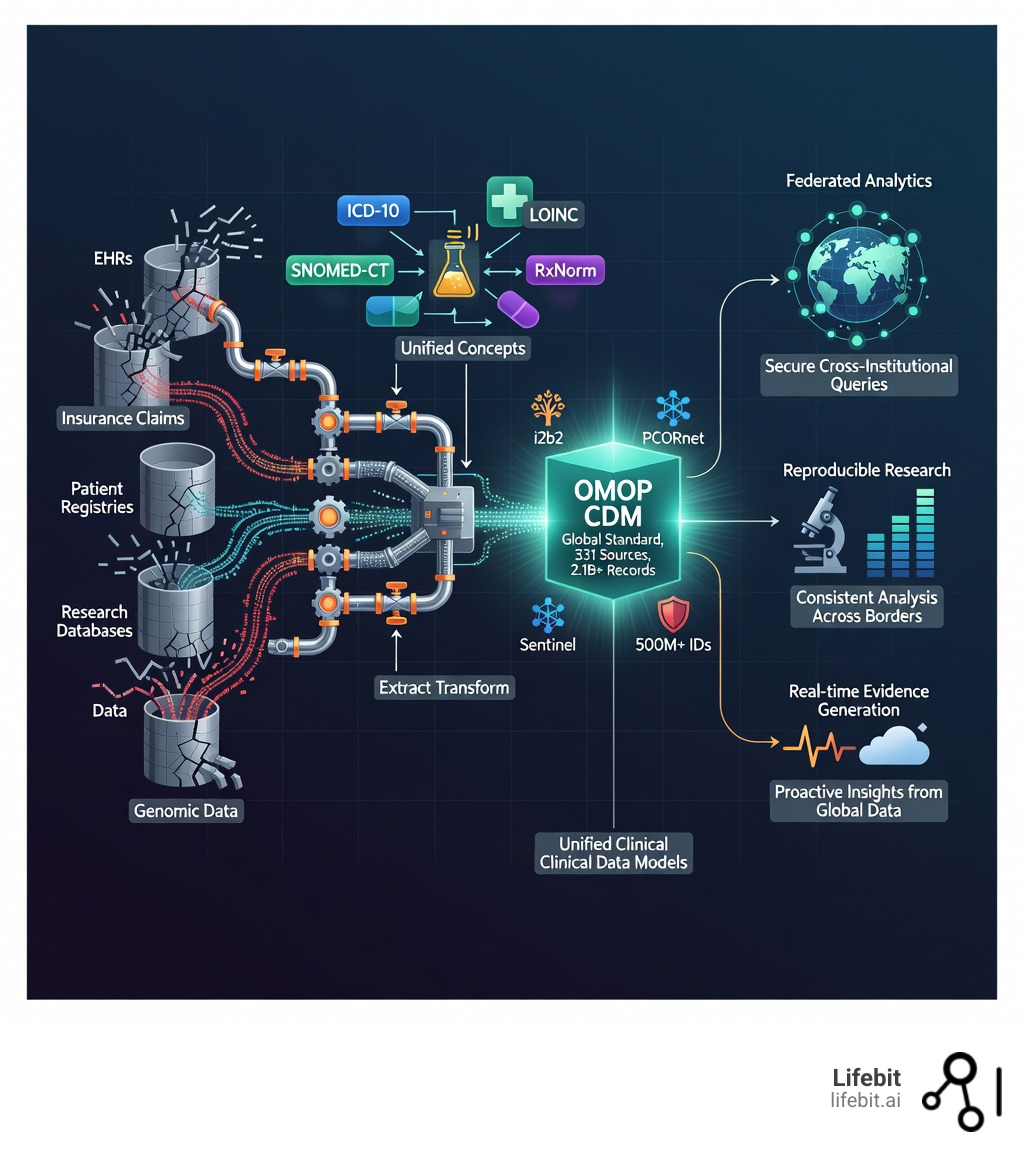
- Standardize fragmented data from electronic health records (EHRs), insurance claims, registries, and research databases into a common format, eliminating the “Tower of Babel” effect in medical informatics.
- Enable semantic interoperability by mapping different coding systems (like ICD-10, SNOMED-CT, and LOINC) to unified concepts, ensuring that a “heart attack” in one system is recognized as the same clinical event in another.
- Support federated research across institutions without moving sensitive patient data, allowing for privacy-preserving analysis that complies with GDPR and HIPAA.
- Accelerate discovery by allowing researchers to run the same query across millions of records in multiple databases simultaneously, reducing the time to generate real-world evidence from years to weeks.
Key Models Include:
- OMOP CDM – The global standard for observational research, maintained by the OHDSI community and used across 331 data sources with 2.1+ billion patient records.
- i2b2 – Optimized for cohort discovery and local institutional use, utilizing a flexible star-schema architecture.
- PCORnet – Designed for patient-centered outcomes research across 337 hospitals, focusing on clinical effectiveness.
- Sentinel – Built for FDA pharmacovigilance with 500+ million patient identifiers to monitor drug safety in real-time.
Healthcare generates massive volumes of data—a typical Phase III clinical trial alone produces 3.6 million data points. Yet this data lives in silos, coded differently across institutions, making collaborative research painfully slow. The economic cost of these silos is staggering; it is estimated that inefficient data handling adds billions to drug development costs and delays life-saving treatments. Clinical data models solve this by acting as universal translators, changing chaotic, incompatible datasets into research-ready assets that can be analyzed consistently across borders and systems.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent over 15 years building platforms that enable secure, federated analysis of genomic and clinical data models across global healthcare networks. My background in computational biology and AI has shown me how the right data model can turn years of data wrangling into weeks of meaningful discovery. By adopting these standards, we move from a reactive healthcare model to a proactive, data-driven ecosystem.
Explore more about clinical data models:
What are clinical data models and Why Do They Matter?
At its heart, a clinical data model is the blueprint for a research-ready database. While a standard database is just a container for information, clinical data models provide the logic and rules for how that information interacts. In healthcare, this is the difference between having a million individual puzzle pieces and having the picture on the box that tells you where they go. Without this blueprint, the sheer volume of healthcare data—expected to reach 2,314 exabytes by 2025—becomes an insurmountable wall rather than a resource.
Why does this matter? Because healthcare data is notoriously messy. A single patient’s journey can produce terabytes of data over their lifetime, scattered across primary care physicians, specialists, hospitals, and pharmacies. Without a common model, a “heart attack” might be recorded as “STEMI” (ST-Elevation Myocardial Infarction) in one hospital’s EHR, “410.11” in an insurance claim using ICD-9 codes, and “Myocardial Infarction” in a research registry. These discrepancies create “data silos” that prevent scientists from seeing the big picture, leading to fragmented care and missed opportunities for identifying drug side effects or disease patterns.
The Anatomy of a Clinical Data Model
A robust clinical data model typically consists of three layers:
- The Structural Layer: Defines the tables (e.g., Person, Visit, Drug Exposure) and the relationships between them. This ensures that every data point has a logical home.
- The Semantic Layer: This is where standardized vocabularies live. It maps local, idiosyncratic codes to a single, universal concept ID. This ensures that when you query for “Hypertension,” you capture every patient regardless of how their doctor coded it.
- The Metadata Layer: Provides context about the data itself—where it came from, how it was transformed, and its level of quality. This is crucial for establishing the “provenance” of research findings.
The Power of Semantic Interoperability
By using clinical data models, we achieve semantic interoperability—the ability for different systems to exchange data with shared meaning. This is a step beyond simple technical interoperability (the ability to send a file). It ensures the receiving system understands the clinical intent of the data. This allows for:
- Structural Consistency: Every database follows the same table and field naming conventions, allowing for “write once, run anywhere” analytics.
- Longitudinal Tracking: We can follow a patient’s health over time, even if they move between providers or insurance plans, by stitching together disparate records into a single timeline.
- Secondary Use of Data: We can take data originally collected for billing or clinical care and use it for life-saving research, as outlined in the American Medical Informatics Association white paper. This maximizes the value of every clinical encounter, turning routine care into a source of medical evidence.
Comparing the Heavyweights: OMOP, i2b2, and PCORnet
Choosing the right model is like choosing a vehicle; you wouldn’t take a sports car off-roading, and you wouldn’t use a tractor for a race. Each of the major clinical data models was built with a specific “mission” in mind, and understanding these nuances is critical for any research institution.
| Feature | OMOP CDM | i2b2 | PCORnet | Sentinel |
|---|---|---|---|---|
| Primary Use | Observational Research | Cohort Discovery | Patient-Centered Outcomes | Safety Surveillance |
| Network | OHDSI (Global) | SHRINE | PCORnet | FDA Sentinel |
| Data Scope | 2.1B+ Records | 200+ Institutions | 337+ Hospitals | 500M+ Identifiers |
| Strength | Advanced Analytics | Local Flexibility | Clinical Integration | Regulatory Safety |
| Architecture | Relational/Standardized | Star Schema | Relational | Distributed Tables |
OMOP: The Global Standard
The OMOP (Observational Medical Outcomes Partnership) CDM is the undisputed champion of observational research. Maintained by the OHDSI (Observational Health Data Sciences and Informatics) community, it currently spans 34 countries. Its deep reliance on standardized vocabularies means that a query written in London will run perfectly on a dataset in Singapore or New York.
OMOP is unique because it transforms all source data into a “Standardized Vocabulary” system. This means that whether your source data uses ICD-10, Read Codes, or local Japanese codes, they are all mapped to a single set of Concept IDs. This allows for massive-scale population studies, such as the LEGEND study, which compared the effectiveness of all first-line hypertension drugs across millions of patients.
i2b2: The Discovery Engine
i2b2 (Informatics for Integrating Biology and the Bedside) is a favorite for local hospital use and academic medical centers. It uses a “star schema” (a central fact table surrounded by dimension tables) that makes it incredibly fast for finding cohorts—for example, “Find me all female patients over 65 with osteoporosis who have also been treated for breast cancer.”
i2b2 is highly flexible; it allows institutions to keep their local coding systems while still enabling cross-institutional queries through the SHRINE (Shared Health Research Information Network) plugin. It is often the first step for hospitals looking to make their EHR data accessible to internal researchers.
PCORnet and PEDSNet: The Clinical Voices
PCORnet focuses on patient-centered outcomes, linking researchers directly with healthcare systems to answer questions that matter most to patients. It is designed to be “low-latency,” meaning the data is updated frequently to reflect current clinical practice. For those focused on children’s health, the CHOP PEDSNet Data Model is a specialized version of these concepts optimized for pediatric-specific research, ensuring that the unique physiological needs and developmental stages of children are represented. You can find detailed PCORnet CDM specifications to see how they handle adult hospital data, including complex elements like patient-reported outcomes (PROs).
Sentinel: The Watchdog
The Sentinel Distributed Database is the FDA’s primary tool for monitoring drug safety. It contains a staggering 1.3 billion person-years of data. Unlike other models that might be used for general research, Sentinel is purpose-built for “active surveillance.” It allows the FDA to spot rare side effects in days rather than years, potentially saving thousands of lives by identifying dangerous drug interactions or manufacturing defects shortly after a product hits the market.
Choosing the Right clinical data models for Your Study
When we help institutions at Lifebit, we tell them to consider three factors:
- Your Research Question: Are you looking for safety signals (Sentinel), long-term outcomes (OMOP), or rapid cohort identification (i2b2)?
- Data Provenance: Where did the data come from? Claims data (which focuses on billing and diagnoses) and EHR data (which includes vitals, notes, and lab results) have different hierarchies and “visit” definitions. OMOP is particularly good at harmonizing these two very different sources.
- Institutional Resources: The ETL (Extract, Transform, Load) process for OMOP is labor-intensive and requires significant expertise in medical terminologies. In contrast, i2b2 is often easier to stand up for local use but may require more work later if you want to join a global network.
How clinical data models Enable the FAIR Principles
The ultimate goal of these models is to make data FAIR: Findable, Accessible, Interoperable, and Reusable.
A prime example is the OHDSI LEGEND-HTN study. By using the OMOP CDM, researchers analyzed data from over 4.9 million patients across multiple international databases. They didn’t have to move the data; they sent the analysis package to the data, a process we champion through our federated platform. This ensures that the data remains secure within the institution’s firewall while still contributing to global medical knowledge.
The ETL Process: Turning Raw Data into Research-Ready Assets
If clinical data models are the destination, ETL (Extract, Transform, Load) is the grueling road trip to get there. It is famously labor-intensive, often consuming 80% of the time and resources in a clinical analytics project. This is where the “dirty work” of data science happens—cleaning, de-duplicating, and standardizing millions of rows of data.
The Step-by-Step Breakdown
- Extract: Pulling raw data from EHRs (like Epic, Cerner, or Allscripts), insurance claims systems, or even wearable devices. This often involves navigating complex SQL databases with thousands of tables.
- Transform: This is the “magic” phase. We map source values to Concept IDs. For example, if a source database has a column for “Gender” with values “1” and “2,” the transformation process maps these to the standard OMOP concepts for “Male” and “Female.” This phase also involves “Data Quality Profiling” to identify outliers, such as a patient recorded as being 150 years old or a lab result that is physiologically impossible.
- Load: Moving the harmonized, cleaned data into the final CDM tables. Once loaded, the data is ready for analysis using standardized tools like ATLAS (for OMOP) or specialized R packages.
The Role of Standardized Vocabularies
Without standard vocabularies, interoperability is impossible. We map data to international standards to ensure universal understanding:
- SNOMED-CT: The most comprehensive clinical terminology in the world, used for clinical findings, symptoms, and diagnoses.
- LOINC: The standard for laboratory tests and clinical observations. It ensures that a “Blood Glucose” test from a lab in Berlin is recognized as the same test as one from a lab in Boston.
- RxNorm: Used for medications and ingredients. It allows researchers to group drugs by their active ingredients, regardless of brand names or dosages.
Data Quality and Validation
Simply moving data into a model isn’t enough; you must prove the data is accurate. Tools like the OHDSI Data Quality Dashboard (DQD) run thousands of automated checks against the data. These checks look for:
- Plausibility: Are the values within a normal range?
- Conformance: Does the data follow the rules of the CDM (e.g., no null values in primary keys)?
- Completeness: Are there massive gaps in the data (e.g., a year where no drug exposures were recorded)?
Researchers can explore these standardized vocabularies on the Athena website to see how thousands of disparate codes map to a single “Standard Concept.” This ensures that a query for “Type 2 Diabetes” (Concept ID: 201820) catches every relevant patient, regardless of the original source code. This level of detail is essential for CTSA CLIC reporting on OMOP CDM, which institutions use to ensure high data quality and maintain their research funding.
Bridging the Gap: FHIR, Salesforce, and Future Directions
The world of clinical data models is evolving rapidly. We are moving away from static, “one-and-done” transformations toward real-time integration and the inclusion of increasingly complex data types like genomics and social determinants of health.
FHIR: Data in Motion
HL7 FHIR (Fast Healthcare Interoperability Resources) is the gold standard for data exchange. While CDMs like OMOP are designed for “data at rest” (large-scale population analysis), FHIR is designed for “data in motion” (real-time exchange between apps and EHRs). In our platform at Lifebit, we often see FHIR acting as the pipeline that feeds real-time clinical data into a research CDM. This allows for “Real-World Evidence” (RWE) that is truly up-to-the-minute, rather than months old.
Salesforce Health Cloud
Even CRM giants are getting involved, recognizing that clinical data is the key to patient engagement. The Salesforce Health Cloud Clinical Data Model Guide shows how FHIR standards are being used to manage patient encounters and medication requests within a business environment. This allows providers to see clinical facts alongside social determinants of health (SDoH), such as housing stability or transportation access, providing a 360-degree view of the patient.
The Next Frontier: Genomics and NLP
The future of clinical data models lies in three transformative areas:
- Genomics: Integrating multi-omic data (genomics, proteomics, metabolomics) directly into clinical tables. This allows researchers to ask questions like, “How do patients with this specific genetic mutation respond to this specific drug?”
- Unstructured Data: Using Natural Language Processing (NLP) to extract insights from free-text doctor’s notes. It is estimated that 80% of clinical value is trapped in these notes. Modern CDMs are beginning to incorporate “NLP tables” that store these extracted entities.
- Generalized Data Models (GDM): Emerging approaches that aim to preserve more of the original data nuance than traditional CDMs. While OMOP is great for standardization, some researchers worry about “information loss” during the ETL process. GDMs attempt to strike a balance between standardization and data richness.
- Federated Learning: This is the ultimate application of CDMs. Instead of moving data to a central server, we move the AI models to the data. This is only possible if every data site uses the same clinical data model, ensuring the AI is “learning” from the same structures at every location.
Frequently Asked Questions about Clinical Data Models
What is the difference between a clinical data model and a database?
Think of a database as the physical library building and the clinical data model as the Dewey Decimal System. The database (like SQL Server, PostgreSQL, or Oracle) provides the storage, the “bricks,” and the security, while the model provides the organizational logic. This logic allows any librarian (researcher) to find exactly what they need, even in a library they’ve never visited before, because they understand the underlying system of categorization.
Can an institution use multiple clinical data models simultaneously?
Absolutely! In fact, many leading research centers do. They might use a raw Epic Clarity model for local chart reviews and hospital operations, i2b2 for quick cohort discovery by clinicians, and OMOP for international collaborative studies. Our platform at Lifebit is designed to handle these hybrid environments, providing a single point of access to disparate models and allowing researchers to join different networks seamlessly.
How do OMOP and FHIR complement each other?
They are two sides of the same coin. FHIR is optimized for individual patient care, mobile apps, and real-time messaging between systems. It is “lightweight” and transactional. OMOP is optimized for large-scale population analytics and complex statistical modeling. Many modern researchers use FHIR to “extract” data from clinical systems in real-time and then “transform” it into OMOP for long-term research and evidence generation.
How long does it take to implement a clinical data model?
The timeline for an ETL project to move data into a CDM like OMOP can range from 3 to 12 months, depending on the complexity of the source data and the expertise of the team. However, once the initial mapping is done, subsequent updates are much faster. Automated tools and AI-assisted mapping are currently reducing these timelines significantly.
Are clinical data models only for hospitals?
No. Pharmaceutical companies use them to analyze clinical trial data and real-world evidence. Insurance companies use them to understand population health and cost-effectiveness. Even public health agencies use them to track disease outbreaks and vaccine safety. Any organization that handles large volumes of health data can benefit from the standardization these models provide.
Conclusion
The shift toward standardized clinical data models is more than a technical upgrade; it’s a democratization of medicine. By breaking down the silos that have traditionally trapped patient data, we are enabling a future where a breakthrough in Singapore can be validated in Canada or the UK in a matter of days. We are moving away from a world where data is a proprietary secret and toward a world where data is a shared resource for the advancement of human health.
At Lifebit, we are proud to be part of this journey. Our federated AI platform allows researchers to bridge the gap between these different models, providing secure, real-time access to global data without ever compromising patient privacy. We believe that the most significant discoveries of the next decade will not come from a single lab, but from the collective analysis of global datasets made possible by these standards. Whether you are navigating the complexities of the OMOP CDM, implementing i2b2 for your institution, or integrating real-time FHIR streams, the goal remains the same: changing raw data into life-saving insights. The infrastructure is being built; now it is time to use it to solve the world’s most challenging medical puzzles.

