Genomic Giants: The Best Platforms for Population-Scale Analysis

Why Genomic Giants Are Racing to Solve Population-Scale Analysis
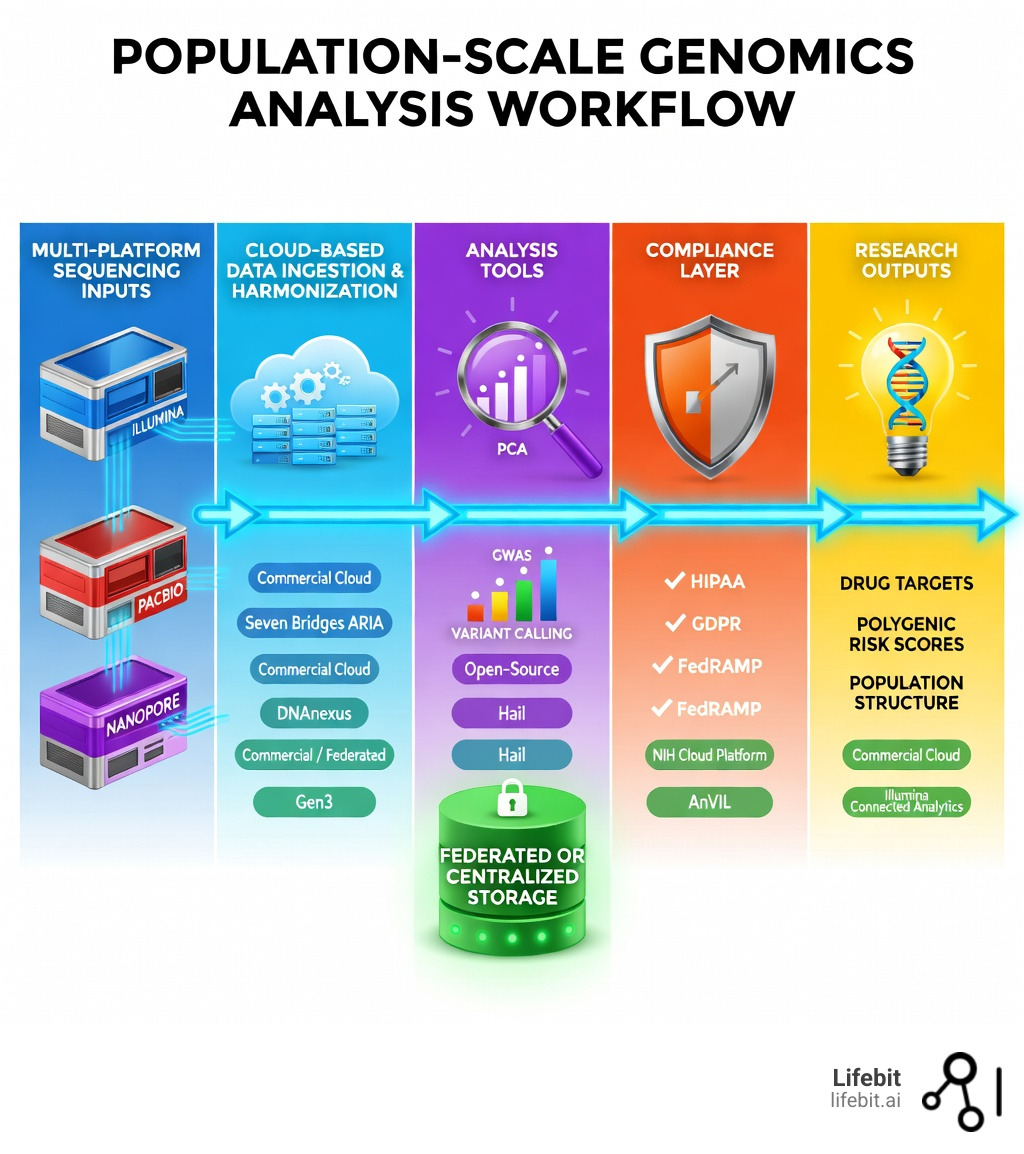
The platforms for population-scale genomics analysis you choose can mean the difference between a breakthrough and a bottleneck. At million-sample scale, most teams end up choosing between two operating models:
- Centralized cloud workspaces: bring data into one controlled environment and analyze at scale.
- Federated analysis: keep data where it already lives (by country, institution, or cloud) and send workflows to it.
Genomic sequencing projects are no longer small experiments. They now routinely span hundreds of thousands – even millions – of participants. The UK Biobank alone holds data on half a million individuals. AnVIL hosts over 5PB of data from more than 287,000 participants across 79 studies. Projects like gnomAD and the GREGoR Consortium have analyzed tens of thousands of individuals to uncover rare variants invisible in smaller cohorts. The transition from the initial Human Genome Project, which took over a decade and cost billions, to today’s reality where we sequence thousands of genomes per week, has created a “data deluge” that threatens to overwhelm traditional bioinformatic infrastructures.
The scale is staggering. In the early 2000s, a single human genome was a monumental achievement. Today, we are looking at the “Century of the Genome,” where the goal is to sequence entire nations. This shift is driven by the realization that rare variants—those occurring in less than 1% of the population—often hold the key to understanding complex diseases like Alzheimer’s, Type 2 Diabetes, and various cancers. To find these variants with statistical significance, researchers need cohorts in the hundreds of thousands.
This creates a real problem for researchers and institutions. Traditional approaches – downloading data locally, running analyses on in-house servers – simply can’t keep up. Data volumes are too large. Compute demands are too unpredictable. And the security stakes around sensitive genomic data are too high. When a single whole-genome sequence (WGS) file can be 100GB, moving 100,000 samples across the internet is not just slow; it is physically and economically impossible for most institutions.
The result? A new generation of platforms built specifically to handle population-scale genomic analysis in the cloud, with compliance, collaboration, and scalability built in from the ground up. These platforms are designed to handle the “Four Vs” of genomic big data: Volume (petabytes of raw reads), Velocity (the speed at which new sequences are generated), Variety (integrating DNA with RNA, proteomics, and clinical records), and Veracity (ensuring data quality and provenance).
This article explains what to look for in modern platforms for population-scale genomics analysis – and how to evaluate architectures (centralized vs. federated) based on your research goals.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit, and a key contributor to Nextflow – one of the most widely used workflow frameworks in genomic data analysis worldwide. Over 15 years working in computational biology, federated data infrastructure, and precision medicine gives me a grounded perspective on what actually works when evaluating platforms for population-scale genomics analysis. Let’s dig in.
Simple platforms for population-scale genomics analysis word guide:
Why Population-Scale Genomics is the New Standard for Precision Medicine

Population-scale genomics is no longer a luxury; it is the engine driving modern Genomics. By analyzing the Genomes of entire populations rather than isolated individuals, we can uncover rare genetic variants that remain invisible in smaller studies. This is critical for identifying novel drug targets, understanding complex diseases, and tailoring treatments to specific genetic profiles.
In medicine, these massive datasets allow us to stratify patient populations with surgical precision. For example, the discovery of PCSK9 inhibitors—a revolutionary class of cholesterol-lowering drugs—was made possible by studying individuals with rare loss-of-function mutations in the PCSK9 gene. Without large-scale population data, such “natural experiments” in human genetics would go unnoticed. Furthermore, population-scale analysis is essential for the development of Polygenic Risk Scores (PRS). PRS aggregate the effects of thousands of common variants to predict an individual’s risk for conditions like coronary artery disease or breast cancer. To calibrate these scores accurately, researchers need data from millions of individuals across diverse ancestral backgrounds to ensure the scores are equitable and effective for everyone, not just those of European descent.
In agriculture, population sequencing helps identify genetic markers for yield and disease resistance, accelerating breeding programs through genomic selection. By sequencing thousands of varieties of a crop like rice or wheat, breeders can identify the specific alleles that allow certain plants to thrive in drought conditions or resist specific pests. Even in evolutionary biology, we can now reconstruct migration patterns and admixture events with unprecedented detail, tracing the history of our species through the subtle variations in our DNA.
However, the traditional model of genomic data sharing—where researchers download data to analyze it locally—is fundamentally broken. As datasets cross the petabyte threshold, moving data is too slow, too expensive, and too risky. The industry is moving toward a model where we invert the model of genomics data sharing, bringing the compute to the data. This shift is the foundation for use case: population genomics strategies that actually scale. This “Data Gravity” effect means that as datasets grow, they become increasingly difficult to move, necessitating platforms that can deploy analysis pipelines directly into the storage environment, whether that is a public cloud bucket or a secure government server.
Overcoming the 5PB Hurdle: Challenges in Platforms for Population-Scale Genomics Analysis
When we talk about “population-scale,” we are talking about a data tsunami. Platforms must now manage datasets exceeding 5PB, such as those hosted on AnVIL, which include over 287,000 participants. Handling this volume presents four primary challenges that require sophisticated engineering solutions:
Data Volume & Storage: Storing millions of whole-genome sequences (WGS) requires elastic cloud infrastructure. Traditional on-premise servers simply run out of room, and the maintenance costs of physical hardware become prohibitive. Modern platforms utilize object storage (like AWS S3 or Google Cloud Storage) which offers virtually infinite scalability and high durability. However, managing the lifecycle of this data—moving infrequently accessed raw reads to “cold” storage while keeping analysis-ready variant files in “hot” storage—is a complex orchestration task.
Compute Scalability: Running a Genome-Wide Association Study (GWAS) on millions of participants requires massive, on-demand parallel processing. A single GWAS might involve testing 10 million variants against a phenotype across 500,000 individuals. This requires thousands of CPUs running in parallel. Without cloud-native auto-scaling, an analysis that should take hours can take months, delaying critical research and drug discovery timelines.
Data Movement Costs: Egress fees from cloud providers can bankrupt a project if you are constantly moving data between regions or out of the cloud entirely. For instance, moving a petabyte of data out of a major cloud provider can cost upwards of $50,000 to $80,000 in transfer fees alone. Platforms must be designed to minimize data movement, utilizing “zero-copy” architectures where multiple researchers can access the same underlying data without creating redundant, expensive copies.
Security & Privacy: Genomic data is the most personal data there is; it is the ultimate identifier. Platforms must ensure that even if an account is compromised, the risk of re-identification is minimized. This involves the physical and logical separation of administrative identifiers (names, IDs) from genomic files. Furthermore, as data privacy laws like GDPR and the CCPA evolve, platforms must support “sovereign data” models where data remains within a specific geographic region while still being accessible for global meta-analysis.
To navigate these hurdles, organizations are forecasting the future of genomic management by moving toward federated and cloud-native architectures that eliminate the need for data transfer entirely. This involves using technologies like Docker and Kubernetes to package analysis tools so they can be shipped to the data, rather than the other way around.
Leading Frameworks and Platforms for Population-Scale Genomics Analysis
Choosing between open-source and commercial platforms often comes down to the balance between customization and “out-of-the-box” efficiency. Both are essential for scaling genomics in clinical environments.
Leveraging Open-Source Tools and Frameworks for Population-Scale Genomics Analysis
Open-source tools provide the transparency and flexibility that academic researchers crave, allowing for deep inspection of algorithms and community-driven improvements.
- Hail: A Python-based library built on Apache Spark, hail-is/hail is widely used for scalable genomic analysis. It has been used in large cohort contexts such as gnomAD and UK Biobank GWAS efforts. Hail’s core innovation is its ability to represent genomic data as a “MatrixTable,” allowing researchers to perform complex filtering, population structure correction (like PCA), and variant annotation across very large variant-by-sample matrices using distributed computing. This allows a researcher to query millions of variants across hundreds of thousands of samples in seconds rather than days.
- AnVIL: This NIH-funded “lab-space” provides a cloud environment to access key NHGRI datasets like the Centers for Common Disease Genomics (CCDG). It integrates tools like Terra (developed by the Broad Institute), Galaxy, and Bioconductor into a unified ecosystem. AnVIL is particularly powerful because it provides a “Trusted Research Environment” where the data is already pre-loaded, saving researchers the cost and time of data ingestion.
- Gen3: Specifically designed for building data commons, Gen3 uses open APIs and modular services to help organizations host and share complex datasets with tiered access. It focuses on data discovery and governance, ensuring that researchers can find the data they need while adhering to strict consent protocols. It has been used to build over 15 data commons globally, including the BloodPAC Data Commons and the Kids First Data Resource.
Commercial Cloud Solutions and Platforms for Population-Scale Genomics Analysis
Commercial platforms often provide a more streamlined experience, reducing the “bioinformatics burden” on researchers by providing managed services and intuitive user interfaces.
- Cohort Browsers: These are high-level interfaces that allow clinicians and researchers to navigate thousands of phenotype fields (e.g., blood pressure, smoking status, MRI results) and large variant datasets without writing a single line of code. This democratizes data access, allowing non-computational experts to generate hypotheses and visualize trends.
- Jupyter & Spark Integration: On-demand clusters that spin up for compute-intensive jobs and spin down automatically to save costs. This “serverless” approach to genomics means researchers only pay for the exact seconds of compute they use.
- Workflow Acceleration: Optimized execution for common pipelines (for example, alignment/variant calling and cohort-scale association testing) to reduce turnaround time. Many commercial platforms offer “accelerated” versions of standard tools like GATK, which can run up to 10x faster than the open-source versions by utilizing specialized hardware like FPGAs or GPUs.
- Data Harmonization and Ingestion at Scale: One of the biggest challenges in population genomics is that data often comes from different sources in different formats. Commercial platforms provide automated tooling to standardize phenotypes (using ontologies like HPO or SNOMED), sample metadata, and analysis-ready formats (like VCF or Zarr) so teams can re-run analyses consistently across disparate cohorts.
For organizations that must collaborate across jurisdictions or data owners, federated models can be a decisive advantage: instead of moving sensitive data, you dispatch approved workflows to wherever the data lives and return only governed, aggregated outputs. This is particularly relevant for international consortia where data cannot leave a country due to national security or privacy laws.
Essential Features for High-Throughput Genomic Workflows
A platform is only as good as the workflows it supports. For population-scale work, the following capabilities are non-negotiable for ensuring scientific rigor and operational efficiency:
- GWAS & PCA: The ability to run linear and logistic regression while correcting for population structure using HWE-normalized Principal Component Analysis (PCA). In large cohorts, failing to account for population stratification (the subtle genetic differences between sub-populations) can lead to false-positive associations. Modern platforms must handle this at the scale of millions of samples.
- Multi-Omics Integration: Moving beyond just DNA to integrate proteomics (proteins), transcriptomics (RNA), and imaging data into a “single source of truth.” The future of precision medicine lies in understanding how a genetic variant affects protein expression, which in turn affects cellular function and ultimately manifests in clinical symptoms. Platforms must be able to join these massive, heterogeneous datasets efficiently.
- Methylation Profiling: Epigenetics—the study of how behavior and environment cause changes that affect the way genes work—is becoming central to disease research. Modern platforms now support HiFi sequencing (from PacBio), which allows for detecting methylation as a “fifth base” without the need for harsh bisulfite treatment. This provides a much clearer picture of the epigenome and its role in aging and cancer.
- Pan-Genome Support: For decades, genomic analysis relied on a single “reference genome” (like GRCh38), which was largely derived from a small number of individuals. This introduced “reference bias,” where variants not present in the reference were harder to detect. Platforms are now moving toward pangenome references—graph-based structures that capture the full genetic diversity of human populations, including large structural variations that are common in certain ancestries but absent in the standard reference.
- Workflow Standards: Support for WDL (Workflow Description Language) and Nextflow ensures that analyses are reproducible and portable. These domain-specific languages allow bioinformaticians to define complex pipelines that can run on any cloud (AWS, Google, Azure) or on-premise cluster without modification. This prevents “vendor lock-in” and ensures that scientific results can be independently verified by other teams.
This technical flexibility is a hallmark of federated technology in population genomics, where workflows can be dispatched to where the data lives, regardless of the underlying hardware or cloud provider. By standardizing the workflow layer, researchers can focus on the science rather than the plumbing of the infrastructure.
Security and Compliance: Protecting Million-Genome Datasets
When handling Genomics England or UK Biobank level data, security is not just a feature—it is the primary concern. A single data breach could undermine public trust in genomic research for a generation. Platforms must adhere to stringent global standards and implement a “defense-in-depth” strategy:
- HIPAA & GDPR: Ensuring patient privacy and the “right to be forgotten.” In the US, HIPAA sets the standard for protecting sensitive patient data, while in Europe, GDPR provides even stricter protections, including the requirement for explicit consent and the right for individuals to have their data deleted. Platforms must have automated tools to manage these consent strings at scale.
- FedRAMP & ISO27001: Meeting the high security bar required for government and institutional data. FedRAMP is a US government-wide program that provides a standardized approach to security assessment, authorization, and continuous monitoring for cloud products and services. ISO27001 is the international standard for information security management systems.
- Trusted Research Environments (TRE): These are “air-locked” environments where researchers can analyze data but cannot download individual-level raw files. In a TRE, the data stays within a secure perimeter. Researchers log in to a virtual workspace, perform their analysis, and can only export the final results (like a summary table or a graph) after it has been reviewed for potential privacy leaks. This is often referred to as the “Five Safes” framework: Safe People, Safe Projects, Safe Settings, Safe Data, and Safe Outputs.
- Data Sovereignty and Residency: Many countries now mandate that genomic data of their citizens must remain within national borders. This creates a massive challenge for global research. Federated platforms solve this by allowing a central “orchestrator” to send analysis tasks to local nodes in different countries. The data never moves, but the insights are aggregated centrally, allowing for global-scale science that respects local laws.
At Lifebit, we advocate for Lifebit CloudOS genomic data federation, which allows for secure collaboration without the data ever leaving its home jurisdiction. This “federated governance” model is the only way to comply with strict data residency laws while still enabling global scientific collaboration. It allows institutions to maintain full control over their data while still participating in the global effort to cure rare diseases and improve human health.
Frequently Asked Questions about Platforms for Population-Scale Genomics Analysis
What are the typical cost models for these platforms?
Most platforms operate on a “pay-as-you-go” model for compute and storage, similar to utility billing. However, the economics of population-scale data are nuanced. Many platforms offer “free hosted datasets” where the platform (or a sponsor like the NIH) covers the storage costs of public data (like the 1000 Genomes Project or the Cancer Genome Atlas), and the user only pays for the compute they use to analyze it. Data transfer (egress) is often the hidden cost; if you run an analysis in AWS US-East but your data is in Google Cloud Europe, you will be hit with massive transfer fees. This is why keeping compute in the same cloud region as the data is critical for budget management.
How do platforms handle different sequencing technologies like PacBio and Nanopore?
Leading platforms are technology-agnostic. While Illumina remains the dominant force for short-read sequencing (ideal for finding small variants), modern platforms provide optimized pipelines for PacBio’s HiFi (long-read) and Oxford Nanopore data. Long-read sequencing is essential for resolving complex regions of the genome, such as structural variants and repetitive elements that short reads often miss. For instance, HiFi sequencing is increasingly used for straightforward epigenome analysis because it detects methylation signals directly during the sequencing run, eliminating the need for chemical conversion steps that can damage DNA.
What is the difference between cloud-native and hybrid approaches?
Cloud-native platforms are built entirely within a provider like AWS or Google Cloud, offering maximum elasticity—the ability to scale from 1 to 10,000 cores in minutes. Hybrid approaches allow an organization to keep some data on their own local servers (perhaps for legacy or extreme security reasons) while “bursting” to the cloud for heavy compute tasks. The trend is moving toward “federated” approaches, which provide a unified interface across both local and cloud environments, making the underlying hardware location transparent to the researcher.
How do these platforms support AI and Machine Learning?
AI is becoming a core component of genomic analysis, from variant calling (using tools like DeepVariant) to predicting protein folding (AlphaFold). Modern platforms provide integrated environments where researchers can easily access GPUs and pre-trained models. They also provide the “data plumbing” necessary to feed massive genomic datasets into ML training pipelines, ensuring that the data is properly formatted and labeled for supervised learning tasks.
Conclusion: Choosing the Right Platform for Your Research
Selecting from the available platforms for population-scale genomics analysis is a strategic decision that affects your research speed, budget, and compliance posture. Whether you are building a national data commons using Gen3, running rapid GWAS with Hail, or managing a Population Genomics Platform Hong Kong Genome project, the goal remains the same: turning massive data into meaningful medicine. The choice often depends on where your data is located and who needs to access it.
As we move toward the goal of sequencing millions of genomes, the limitations of centralized data silos become more apparent. The future of genomics is not about building bigger and bigger central databases, but about creating a connected ecosystem of data that can be queried securely and efficiently. This requires a shift in mindset from “owning the data” to “governing access to the data.”
At Lifebit, we believe the future is federated. Our Lifebit Federated Biomedical Data Platform is designed to solve the data silo problem by bringing AI-driven analytics directly to the data. This approach ensures that whether you are working with the Genomics England Research Environment or a private biopharma dataset, you have the real-time insights you need without compromising security. By removing the barriers to data access, we can accelerate the pace of discovery and bring the promise of precision medicine to patients faster than ever before.
Ready to scale your analysis? Let’s build the future of precision medicine together.

