How to Link Records in Python Without Losing Your Mind

Why Messy Data Integration is Killing Your Analysis
Record linkage python is the process of identifying and connecting records that refer to the same real-world entity across different datasets, even when those records lack a shared unique identifier. Here’s what you need to know:
Quick Answer: Core Record Linkage Steps
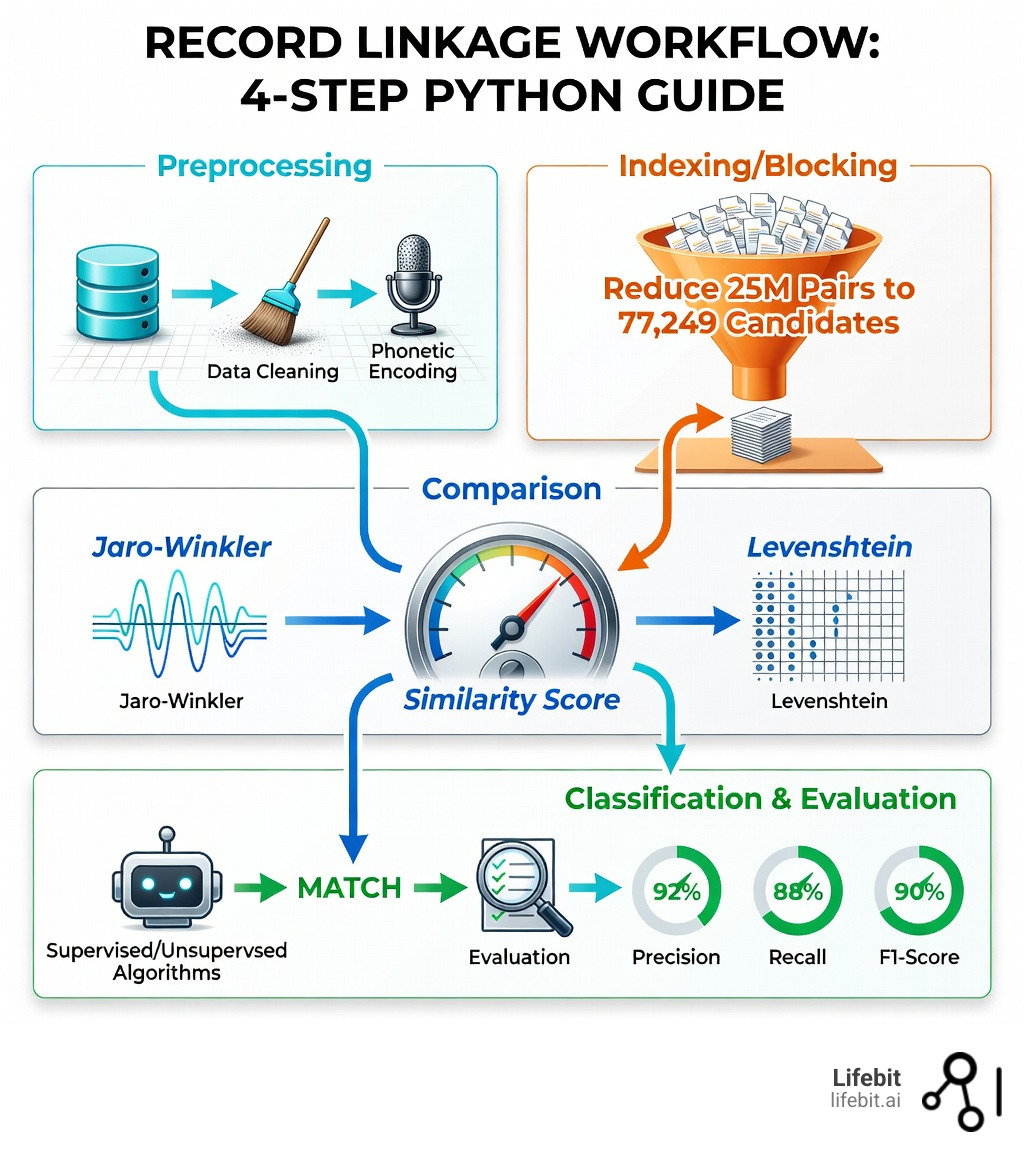
- Preprocessing – Clean and standardize your data (remove whitespace, apply phonetic encoding)
- Indexing/Blocking – Reduce comparison pairs from millions to thousands using smart filters
- Comparison – Calculate similarity scores using methods like Jaro-Winkler or Levenshtein distance
- Classification – Apply thresholds or machine learning to identify true matches
- Evaluation – Measure accuracy with precision, recall, and F1-score
The problem is real: when you’re trying to merge customer databases, integrate clinical records, or deduplicate entities across sources, you’re facing 25 million potential record pairs for just two datasets of 5,000 records each. Without proper techniques, this computational nightmare will grind your analysis to a halt.
But here’s the good news: Python libraries like recordlinkage, Splink, and hlink have solved this problem. Using blocking techniques, you can reduce those 25 million comparisons down to 77,249 meaningful candidate pairs. With the right similarity metrics and classification approach, you can achieve 90% match rates on real-world data—even when your inputs are dirty, inconsistent, or missing shared attributes.
I’m Dr. Maria Chatzou Dunford, CEO of Lifebit, where we’ve spent over a decade solving record linkage python challenges for biomedical and multi-omic research across federated, secure environments. We’ve helped pharmaceutical organizations and public sector institutions integrate siloed genomic, clinical, and claims data at massive scale—and I’ll show you exactly how to do it without the headache.
Record linkage python terms to learn:
Why Record Linkage is the Secret to Clean Data Engineering
In modern data engineering, we rarely deal with a single, pristine database. Instead, we are often tasked with merging disparate sources—think data-linking CRM data with web logs or connecting patient registries with pharmacy claims. Without a shared primary key (like a Social Security number or a universal ID), standard SQL joins fail. This is where the concept of “Entity Resolution” or “Record Linkage” becomes the backbone of your data architecture.
Record linkage allows us to create a “Single Source of Truth” or a “Golden Record.” By identifying that “Jon Doe” at 123 Main St is the same person as “Johnathan Doe” at 123 Main Street, we enable a customer 360 view that drives better business decisions and more accurate research. At Lifebit, we see this daily: data-linkage is the foundation of high-quality analytics. If the underlying data is fragmented, your AI models will be too. The economic impact of this is staggering; Gartner estimates that poor data quality costs organizations an average of $12.9 million per year. By implementing robust record linkage python pipelines, you aren’t just cleaning data; you are recovering lost revenue and preventing catastrophic analytical errors.
Why Record Linkage Python is Essential for Data Quality
Dirty data is the enemy of progress. In a typical project, we might find that 90% of records match easily, but the final 10% are “dirty”—riddled with typos, varying formats, or missing fields. Using record linkage python techniques allows us to bridge these gaps. This is particularly vital in the era of “Data Debt,” where legacy systems and rapid scaling have left organizations with massive, unorganized data lakes.
For example, in healthcare, link-claims-ehr-data-real-world-evidence is critical for understanding patient outcomes. Records might have different name spellings, transposed birth dates, or even different addresses due to patient relocation. By using probabilistic matching, we can identify these links with high confidence, turning messy inputs into data quality assets that fuel real-world evidence (RWE) studies. Without these techniques, a researcher might treat the same patient as two different individuals, leading to skewed survival rates or incorrect drug efficacy conclusions. The ability to resolve these entities at scale is what separates basic data processing from advanced data science.
The 4-Step Workflow to Link Records Without Losing Your Mind
To keep your sanity, you must view record linkage as a pipeline rather than a single function call. Following a structured workflow ensures you don’t waste computational resources or lose accuracy. This pipeline approach allows for iterative testing and refinement, which is essential when dealing with the nuances of human-generated data.
Preprocessing and Cleaning Messy Inputs
You can’t compare what you haven’t cleaned. Preprocessing involves standardizing strings to a common format. This includes:
- Whitespace removal: Stripping leading/trailing spaces and collapsing multiple spaces into one. This is often the most common cause of join failures in standard SQL.
- Case standardization: Converting all text to lowercase to ensure that “SMITH” and “smith” are treated identically.
- Phonetic encoding: Using algorithms like Soundex, NYSIIS, or Metaphone to convert names into codes based on how they sound. For instance, “Smith” and “Smyth” both become “S530” in Soundex. This is crucial for handling surnames that have multiple common spellings.
- Phone and Address normalization: Removing special characters from phone numbers and standardizing address suffixes (e.g., “St.” to “Street”). Using libraries like
usaddressorphonenumbersin Python can automate this process.
If you’re new to the logic behind this, a RegexOne refresher is a great place to start for mastering the regular expressions often used in the recordlinkage documentation.
Smart Indexing to Kill Computational Bloat
The biggest challenge in record linkage python is the “n-squared” problem. If you compare every record in Dataset A (100,000 rows) with every record in Dataset B (100,000 rows), you generate 10,000,000,000 (10 billion) pairs. Even with a fast computer, this will take hours or days.
Indexing (specifically blocking) solves this by only creating pairs that share a common attribute. For instance, if we only compare records that have the same “zip_code,” we can reduce those 10 billion pairs down to a few million. This results in a massive reduction ratio, allowing your code to run in seconds rather than hours.
Common strategies include:
- Standard Blocking: Comparing records only if they match exactly on a specific key (e.g., Year of Birth).
- Sorted Neighborhood: Sorting the data by a key and only comparing records within a specific “window” (e.g., the 10 records before and after). This is more robust to typos in the blocking key itself.
- Canopy Clustering: A more advanced technique that uses a fast, approximate distance metric to group similar records into “canopies” before performing more expensive comparisons within those groups.
Top Libraries for Record Linkage Python: Choosing the Right Tool for the Job
Not all libraries are created equal. Depending on your dataset size and technical stack, you’ll want to choose carefully. The Python ecosystem is rich with options, ranging from simple research tools to massive distributed systems.
| Library | Best For | Scaling | Key Feature |
|---|---|---|---|
| recordlinkage | Research & Mid-sized data | Single machine | Modular, easy to use, Pandas-based |
| Splink | Large-scale production | PySpark / DuckDB | Extremely fast, probabilistic, SQL-backed |
| hlink | Hierarchical data | PySpark | Configuration-driven, great for census data |
| RLTK | Academic & Complex | Multi-core | Extensible, built by USC/ISI researchers |
| Dedupe | Active Learning | Single machine | Uses machine learning to learn matching rules |
Choosing the right database-matching-software depends on whether you’re working locally or in a distributed environment. For most users starting out, the recordlinkage library provides the best balance of features and ease of use.
Probabilistic Matching with Splink and the Fellegi-Sunter Model
For those dealing with millions or billions of records, Splink is often the gold standard. It is based on the Fellegi-Sunter model, the mathematical foundation of probabilistic record linkage. This model works by calculating two key probabilities for each field:
- m-probability: The probability that the field matches given that the records are a true match (measures the quality of the data).
- u-probability: The probability that the field matches by pure chance given that the records are not a match (measures the uniqueness of the field).
By combining these, Splink assigns a “match weight” to each field. A match on a rare surname like “Cholmondeley” will receive a much higher weight than a match on a common surname like “Smith.” This allows the system to make highly nuanced decisions that simple rule-based systems would miss. Similarly, hlink provides a configuration-driven approach specifically designed for hierarchical record linkage at scale, often used for historical census data where individuals are nested within households.
Supervised vs Unsupervised Classification
Once you have your comparison scores, you need to decide if a pair is a match.
- Supervised Learning: If you have labeled data (an “answer key”), you can train a Scikit-learn model like Logistic Regression, Random Forest, or XGBoost to classify pairs. This is highly accurate but requires the manual effort of labeling thousands of pairs.
- Unsupervised Learning: If you don’t have labels (which is common), you can use the ECM (Expectation-Conditional Maximisation) algorithm. This estimates the probability of a match without needing a training set by looking at the distribution of similarity scores across the entire dataset.
Alternatively, many practitioners use a simple sum-of-scores threshold. For example, if you compare 6 features and the total similarity score is > 3.0, you mark it as a match. This is easy to implement but requires careful tuning to avoid false positives.
Step-by-Step: Implementing Record Linkage Python
Ready to get your hands dirty? Here is how you implement a basic pipeline using the recordlinkage library. This example demonstrates the core logic of indexing, comparing, and classifying.
- Install the library:
Follow the recordlinkage installation manual by runningpip install recordlinkage. - Load your data:
Import your data into two pandas DataFrames. Ensure your columns are named consistently across both datasets. -
Define an Indexer:
import recordlinkage indexer = recordlinkage.Index() indexer.block('zip_code') # Only compare records in the same zip code candidate_links = indexer.index(dfA, dfB) -
Compare attributes:
Use metrics like Jaro-Winkler (great for names) or Levenshtein (edit distance) to score the pairs. Jaro-Winkler is particularly effective for names because it penalizes errors at the start of a string more heavily than errors at the end.compare = recordlinkage.Compare() compare.string('first_name', 'first_name', method='jarowinkler', threshold=0.85) compare.string('last_name', 'last_name', method='levenshtein', threshold=0.80) compare.exact('date_of_birth', 'date_of_birth') features = compare.compute(candidate_links, dfA, dfB)
Measuring Success with Precision and Recall
How do you know if your linkage worked? You need to look at the confusion matrix. In the context of record linkage, the stakes are often high. A “False Positive” (linking two different people) can lead to privacy violations or incorrect medical treatments. A “False Negative” (failing to link the same person) leads to incomplete data.
- Precision: What proportion of predicted links were actually correct? (Avoiding “False Positives”). High precision is vital when merging financial or medical records.
- Recall: What proportion of the total actual links did your model find? (Avoiding “False Negatives”). High recall is important for investigative work or census data where you want to find every possible connection.
- F1-score: The harmonic mean of the two. This provides a single metric to evaluate the overall performance of your model.
In sensitive fields like healthcare, high precision is often prioritized. You can find deep dives into these Precision and Recall metrics on Wikipedia. To truly master this, you should plot a Precision-Recall Curve to find the optimal threshold for your specific use case.
Scaling Record Linkage Python for Massive Datasets
When you move from 5,000 records to 500,000, your laptop will likely struggle. This is where performance optimization becomes key. Libraries like Splink are built for this, utilizing distributed computing and multi-core processing to handle large-scale integration. By using PySpark or DuckDB backends, you can process millions of rows in minutes by distributing the comparison tasks across a cluster. Furthermore, consider using Privacy-Preserving Record Linkage (PPRL) techniques like Bloom Filters if you are working with sensitive data that cannot be shared in plaintext between organizations.
Frequently Asked Questions about Record Linkage
What is the difference between blocking and full indexing?
Full indexing compares every single row in Dataset A to every row in Dataset B. If both have 10,000 rows, that’s 100 million comparisons. Blocking restricts comparisons to records that share a “block” key (like a City or Year of Birth), which drastically reduces the workload and makes the process computationally feasible. It is the difference between a process taking 10 seconds versus 10 hours.
Which string similarity metric is best for names?
Jaro-Winkler is generally considered the best for names because it gives more weight to the beginning of the string (prefixes) and is more forgiving of common typographical errors. Soundex is better if you want to match names that are spelled differently but sound the same (e.g., “Catherine” vs. “Kathryn”). For long strings like addresses, Levenshtein or Monge-Elkan are often more appropriate.
How do you handle record linkage without shared unique IDs?
This is exactly what probabilistic record linkage is for! You compare multiple “soft” identifiers—like name, address, birth date, and gender—and calculate an overall similarity score. If the combined evidence is strong enough, you treat the records as a match. This mimics how a human would look at two files and decide if they belong to the same person.
Can record linkage be done in real-time?
While most record linkage is done in batch processing, real-time linkage is possible using “Index-only” lookups. When a new record enters the system, it is preprocessed and then compared against existing “candidate” blocks in the database. This is common in master data management (MDM) systems to prevent duplicate entry at the point of capture.
What is the “Golden Record” in entity resolution?
The Golden Record is the version of a record that is considered the most accurate and complete. After linking multiple records from different sources, you “collapse” them into one. For example, if Dataset A has the correct phone number and Dataset B has the correct address, the Golden Record combines both to create the most up-to-date profile of the entity.
How do you handle missing data in record linkage?
Missing data (nulls) can significantly degrade linkage quality. Most Python libraries handle this by assigning a neutral weight to missing fields, meaning they neither contribute to nor detract from the match score. However, if too many fields are missing, the record may become “unlinkable” without manual intervention.
Conclusion
Mastering record linkage python is a superpower for any data scientist or engineer. It turns fragmented, messy data into a cohesive asset that can drive groundbreaking research and business insights. At Lifebit, we use these principles to power our federated AI platform, allowing researchers to securely access and link global biomedical and multi-omic data without ever moving the data itself.
Whether you are using our Trusted Research Environment (TRE) or building your own local pipeline with recordlinkage or Splink, the key is to be systematic: clean your data, block your pairs, and evaluate your results.
Ready to take your data integration to the next level? Learn more about how we handle complex data at scale on the Lifebit platform.

