The Clinical Genomics Platform: A Deep Dive into Modern Medical Research
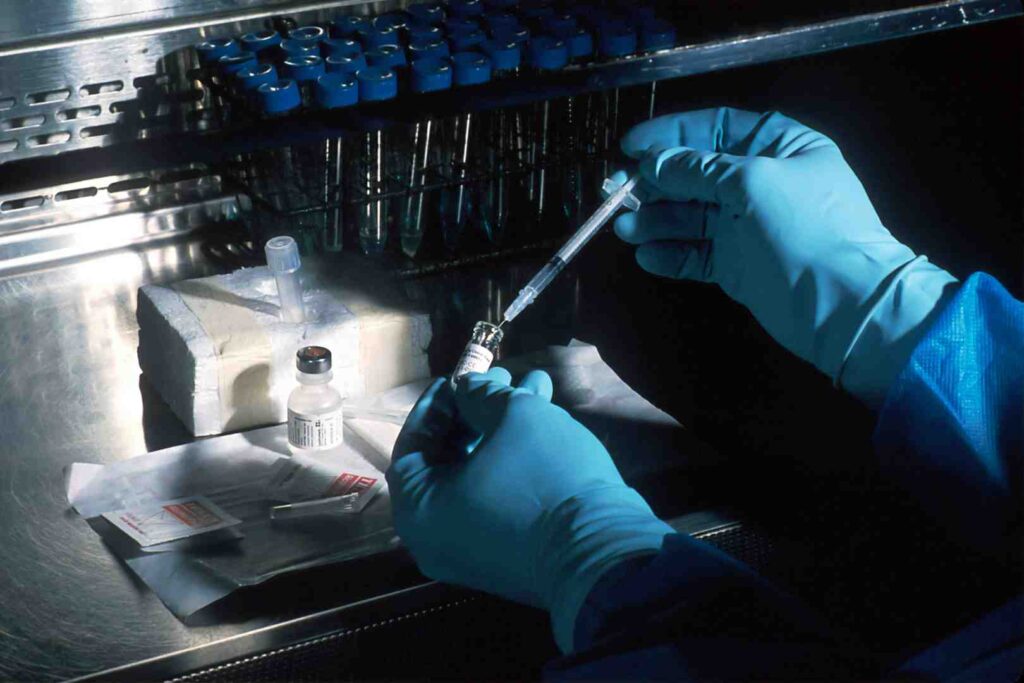
How a Clinical Genomics Platform Cuts Variant Analysis Time by 98%
A Clinical Genomics Platform is an integrated system that combines high-throughput DNA sequencing, advanced bioinformatics, and secure data management to transform raw genetic data into actionable clinical insights. These platforms enable healthcare providers and researchers to diagnose rare diseases, personalize cancer treatments, and accelerate drug discovery—all while maintaining strict data privacy and regulatory compliance. The evolution of these platforms marks a shift from the era of Sanger sequencing, which was slow and labor-intensive, to the era of Next-Generation Sequencing (NGS), where billions of DNA fragments can be sequenced in parallel.
Key capabilities of a clinical genomics platform:
- Library preparation and sequencing: This initial phase involves the fragmentation of DNA or RNA, the ligation of adapters, and the amplification of the library. Modern platforms support various modalities, including Whole Genome Sequencing (WGS), which captures the entire 3 billion base pairs of the human genome; Whole Exome Sequencing (WES), which focuses on the 1-2% of the genome that codes for proteins; and targeted gene panels, which look at specific sets of genes associated with particular conditions like hereditary breast cancer or cardiovascular disorders.
- Bioinformatics analysis: Once the sequencer generates raw data (FASTQ files), the platform applies GPU-accelerated pipelines such as GATK (Genome Analysis Toolkit) or Illumina’s DRAGEN. These pipelines perform read alignment to a reference genome (like GRCh38), duplicate marking, base quality score recalibration (BQSR), and variant calling to detect single nucleotide variants (SNVs), small insertions/deletions (indels), copy number variations (CNVs), and complex structural alterations.
- Variant interpretation: This is the most cognitively demanding step. Platforms use AI/ML tools to cross-reference detected variants against massive databases like ClinVar, gnomAD, and COSMIC. They apply phenotype matching using the Human Phenotype Ontology (HPO) and follow the rigorous ACMG/AMP (American College of Medical Genetics and Genomics / Association for Molecular Pathology) guidelines to classify variants as Pathogenic, Likely Pathogenic, Variant of Uncertain Significance (VUS), Likely Benign, or Benign.
- Clinical reporting: The final output is a clinician-ready report. These systems generate compliant PDF or JSON reports that can be automatically pushed to Laboratory Information Management Systems (LIMS) or Electronic Health Records (EHR). This ensures that the genetic findings are integrated into the patient’s broader medical history, allowing for truly informed decision-making.
- Data security and Governance: Given the sensitivity of genomic data, platforms employ federated data models, end-to-end encryption, and multi-factor authentication. They must adhere to global certifications such as CE-IVD (under the new IVDR regulations), ISO 27001 for information security, GDPR in Europe, and HIPAA in the United States.
The impact of these platforms is measurable and profound. Advanced systems have reduced analysis time by 98%, serving thousands of geneticists globally and analyzing hundreds of thousands of unique cases. Large-scale clinical services have sequenced thousands of asymptomatic adults, creating massive cohorts of clinically sequenced individuals that help us understand the prevalence of genetic risk factors in the general population. Meanwhile, national clinical genomics initiatives operate nodes across university medical faculties, bridging the gap between translational research and routine diagnostics.
These systems don’t just speed up diagnosis—they democratize access. By reducing variant analysis timelines from weeks to days, modern clinical platforms enable equitable care for underserved populations who might otherwise wait years for a diagnosis. ClinGen, a global collaborative effort spanning 74 countries and involving 2,800+ contributors, defines the clinical relevance of genes and variants, ensuring standardized, evidence-based curation worldwide. This global standardization is critical; it ensures that a variant classified as pathogenic in one part of the world is treated with the same clinical urgency elsewhere.
Why this matters now: High-throughput sequencing generates vast datasets—millions of variants per genome—that require specialized infrastructure, complex workflows, and extensive computational power. Clinical genomics platforms solve these challenges by integrating secure data management, automated analysis, and accurate test interpretation aligned with healthcare standards. They enable precision oncology by rapidly analyzing tumor mutational signatures and tumor mutational burden (TMB), support rare disease diagnosis through phenotype-driven prioritization, and facilitate pharmacovigilance through real-time safety surveillance of how genetic variations affect drug metabolism.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, a pioneering federated genomics and biomedical data platform. With over 15 years in computational biology, AI, and health-tech entrepreneurship, I’ve built tools that power Clinical Genomics Platforms for precision medicine, enabling secure, real-time access to global biomedical data across pharmaceutical and public sector organizations.

Clinical genomics platform terms at a glance:
Stop Drowning in DNA Data: Why Your Lab Needs a Clinical Genomics Platform
At its core, a Clinical Genomics Platform is the engine room of modern precision medicine. Its primary function is to manage the massive influx of data generated by high-throughput sequencing and turn it into something a doctor can actually use to treat a patient. Without these platforms, we would be drowning in billions of “A, C, T, and G” letters with no way to find the single typo causing a rare disease. The sheer scale of the data is often underestimated; a single human genome, when sequenced at high depth, can generate upwards of 100 gigabytes of raw data. For a hospital processing hundreds of patients a week, this creates a data management nightmare that only a dedicated platform can solve.
The importance of these systems lies in their ability to provide diagnostic accuracy that was previously impossible. For example, the Advancing genomic knowledge through global curation study highlights that large-scale evidence-based curation is essential for the infrastructure needed to support global medical consortia. By standardizing how we interpret genetic data, these platforms ensure that a patient in London receives the same quality of diagnostic insight as one in New York or Singapore. This consistency is vital for clinical trials and for the development of new therapies that target specific genetic markers.
Bridging the Gap with a Clinical Genomics Platform
One of the biggest problems in medicine is moving a discovery from a research lab into a hospital. This is often called the “translational gap.” A Clinical Genomics Platform acts as the bridge. It takes the latest high-throughput (HTP) techniques, such as Next-Generation Sequencing (NGS), and adapts them for routine clinical use. In the research world, a pipeline might take weeks to run and require manual intervention at every step. In a clinical setting, this is unacceptable. A clinical platform automates these workflows, ensuring they are reproducible, validated, and fast enough to meet clinical turnaround times (TAT).
National initiatives show how this works in practice. By establishing national competence centers, countries provide the support needed for clinicians to adopt HTP technologies in routine diagnostics for cancer and inherited diseases. These platforms allow doctors to Browse curated gene-disease validity data in real-time, ensuring that every medical decision is backed by the most current evidence-based medicine. Furthermore, these platforms facilitate the “Diagnostic Odyssey”—the long and often painful journey patients with rare diseases undergo, sometimes for decades, before finding a cause. By implementing WGS early in the diagnostic pathway, clinical platforms can end this odyssey in a matter of days.
The Role of Evidence-Based Medicine
In the past, genetic interpretation was often siloed within individual labs, leading to conflicting reports where one lab might call a variant “pathogenic” while another called it “uncertain.” Clinical genomics platforms solve this by integrating with global knowledge bases. They allow for the real-time sharing of phenotypic and genotypic data (in a de-identified manner), which strengthens the collective understanding of rare variants. This collaborative approach is what powers the “actionability” of the data. If a platform identifies a variant that has been successfully targeted by a specific drug in a clinical trial elsewhere, that information is immediately available to the treating physician, potentially saving the patient’s life.
From Sample to Report: The Tech Powering Modern Diagnostics
Building a high-performing Clinical Genomics Platform requires a stack of sophisticated services and technologies. It’s not just about the sequencing machine; it’s about the entire pipeline from the moment a blood sample is taken to the moment a report is generated. This pipeline is generally divided into three stages: Primary, Secondary, and Tertiary analysis.
- Library Preparation and Primary Analysis: This is the “prep work” where DNA is fragmented and tagged. Primary analysis occurs on the sequencer itself, where the machine converts physical signals (like light or pH changes) into digital base calls (A, C, T, G) and assigns quality scores to each call. This results in the FASTQ file.
- Secondary Analysis (Bioinformatics): This is where the heavy lifting happens. The platform takes the millions of short reads from the FASTQ file and aligns them to a reference genome. This is like putting together a 3-billion-piece jigsaw puzzle. Once aligned (BAM/CRAM files), the platform identifies where the patient’s DNA differs from the reference. This process, known as variant calling, produces a VCF (Variant Call Format) file. To handle this at scale, modern platforms use GPU-accelerated analysis infrastructure, which can process a whole genome in under an hour—a task that used to take days on standard CPUs.
- Tertiary Analysis (Interpretation): This is where the magic happens. The VCF file contains millions of variants, most of which are harmless. The platform filters these based on frequency (is it rare in the population?), functional impact (does it change a protein?), and clinical relevance (is it linked to the patient’s symptoms?).
- Variant Interpretation and Classification: Using AI-powered engines, the platform assists the molecular pathologist in classifying the remaining variants. It pulls in data from literature, protein structure predictors (like AlphaFold), and conservation scores (how much the gene has changed over millions of years of evolution).
Advanced Analytics within the Clinical Genomics Platform
The true power of a modern platform comes from the integration of Artificial Intelligence (AI) and Machine Learning (ML). These tools don’t just process data faster; they process it smarter, identifying patterns that are invisible to the human eye.
- Phenotype Matching: By using the Human Phenotype Ontology (HPO), platforms can match a patient’s physical symptoms (e.g., “seizures,” “hypertrophic cardiomyopathy”) to their genetic variants. This phenotype-driven prioritization significantly increases the diagnostic yield, especially in complex rare disease cases where multiple variants might be present.
- Polygenic Risk Scores (PRS): While traditional genomics looks for single “smoking gun” mutations, PRS looks at the cumulative effect of thousands of small variations. Clinical platforms are increasingly using PRS to predict a person’s risk for common diseases like heart disease, type 2 diabetes, or breast cancer, allowing for preventative interventions years before symptoms appear.
- Automated Prioritization: Platforms can now Access clinical actionability curations to automatically flag which genetic findings have clear treatment options. For example, if a patient with lung cancer has an EGFR mutation, the platform will immediately highlight the specific tyrosine kinase inhibitors (TKIs) that are effective against that mutation.
| Feature | Whole Genome Sequencing (WGS) | Whole Exome Sequencing (WES) |
|---|---|---|
| Coverage | Entire DNA (3 billion base pairs) | Protein-coding regions (1-2% of DNA) |
| Diagnostic Yield | Highest (can find non-coding and structural variants) | High (finds ~85% of known disease-causing variants) |
| Data Size | Very Large (~100GB+ per sample) | Moderate (~10GB per sample) |
| Cost | Higher (but decreasing rapidly) | Lower (more accessible for routine use) |
| Clinical Utility | Best for undiagnosed rare diseases | Standard for many oncology panels |
Cut Costs and Scale: Solving the High-Throughput Sequencing Headache
Implementing high-throughput sequencing isn’t without its headaches. The sheer volume of data is staggering—one human genome can take up as much space as 100 high-definition movies. For a large healthcare system, the storage requirements quickly reach the petabyte scale. Managing this requires specialized equipment, high-speed networking, and complex workflows that many hospitals simply aren’t equipped to handle on their own. This is often referred to as the “data deluge” in genomics.
A Clinical Genomics Platform addresses these challenges through scalability and cost reduction. By centralizing the computational power in a secure cloud-based or hybrid genomic data center, we can make advanced diagnostics affordable for everyone, not just those at elite research institutions. This is vital for ensuring equitable care and global competitiveness. Cloud-native platforms allow for “elastic” computing, where resources are only used when a sample is being processed, significantly lowering the overhead costs compared to maintaining massive on-premise server farms.
Furthermore, these platforms solve the “silo” problem. In many institutions, data is trapped in individual departments, making it impossible to perform large-scale analysis. A unified platform provides a single source of truth. For instance, researchers can Explore somatic cancer variant databases to see how specific mutations respond to different drugs across their entire patient population. This allows for “precision oncology” that targets the tumor’s specific genetic makeup, reducing the use of ineffective, toxic treatments and ultimately lowering the total cost of care for the healthcare system.
Another cost-saving aspect is the reduction in “re-sequencing.” When data is stored in a standardized, accessible format within a clinical platform, a patient’s genome can be re-analyzed as new medical knowledge emerges without needing to draw blood or run the sequencer again. This “living data” approach means that a negative result today could become a life-saving diagnosis two years from now, simply by re-running the interpretation pipeline against an updated database.
HIPAA, GDPR, and Beyond: Securing Your Patient’s Genetic Blueprint
When you are dealing with a person’s entire genetic blueprint, security isn’t just a “nice to have”—it’s a legal and ethical requirement. Genetic data is uniquely identifiable; unlike a credit card number, you cannot change your DNA if it is leaked. Therefore, the security architecture of a Clinical Genomics Platform must be robust, resilient, and transparent. We take this responsibility incredibly seriously, implementing “security by design” at every layer of the software stack.
A robust platform must comply with international standards like HIPAA (Health Insurance Portability and Accountability Act) in the USA and GDPR (General Data Protection Regulation) in Europe. But we go further. Leading clinical platforms are CE-IVD Class C certified under the new In Vitro Diagnostic Regulation (IVDR) and hold ISO 27001 (Information Security Management) and HDS (Health Data Hosting) certifications. These systems undergo rigorous third-party auditing and penetration testing to ensure the highest levels of data protection and regulatory compliance.
The Federated Revolution
To solve the problem of data “silos” and the legal restrictions on moving medical data across borders (data residency requirements), we use federated data models. Traditionally, to analyze data from different hospitals, you would have to move all that data to a central server. This creates a massive security risk and often violates privacy laws. In a federated model, the data stays exactly where it was created—inside the hospital’s secure firewall. Instead of moving the data to the analysis, we move the analysis to the data.
This “federated AI” approach ensures that sequencing data never leaves the home institution. Only the non-identifiable “insights” or aggregate results are shared. This maintains strict data ownership and sovereignty while still allowing for global collaboration. For example, a researcher in Brazil can run a query across genomic datasets in the UK, Germany, and Japan to find other patients with a specific rare mutation, without any of those patients’ raw DNA files ever being transferred. This is the only way to achieve the scale needed for precision medicine while respecting the stringent privacy demands of the 21st century. It also protects against “data gravity,” where the cost and time of moving massive datasets become a barrier to scientific progress.
Frequently Asked Questions about Clinical Genomics
How do these platforms improve cancer and rare disease treatment?
Clinical genomics platforms are game-changers for oncology and rare diseases. In cancer, they enable tumor profiling to identify “actionable” mutations—those for which a specific targeted drug or clinical trial already exists. This moves us away from a “one-size-fits-all” chemotherapy approach to precision oncology. For rare diseases, platforms help identify de novo mutations (new mutations not inherited from parents) that are often the root cause of undiagnosed conditions. Efforts like the Medical Genome Initiative are actively working to expand access to clinical whole genome sequencing (cWGS) to end the “diagnostic odyssey” for these patients, often providing answers in days for critically ill infants in neonatal intensive care units (NICU).
What role does AI play in genomic analysis?
AI is the “super-expert” that never sleeps. It assists geneticists by automating the most tedious and error-prone parts of the job, such as variant prioritization and literature review. Machine learning models can recognize patterns in multi-omic data—combining genomics with transcriptomics (RNA) or proteomics (proteins)—to find subtle clues about disease progression that a human might miss. AI can also predict the functional impact of “variants of uncertain significance” (VUS) by simulating how a mutation might change the shape and function of a protein. This speeds up diagnosis from weeks to days, which is critical for patients with aggressive cancers where every day counts.
How is patient data protected in large-scale sequencing?
Protection is built into the architecture using a “defense in depth” strategy:
- Encryption: Data is encrypted using AES-256 standards both at rest (on the server) and in transit (while being moved).
- Granular Access Control: Using Role-Based Access Control (RBAC), only authorized personnel can see specific files, and every access event is logged in an immutable audit trail.
- Federated Governance: Data stays within its original jurisdiction (e.g., a hospital in London or a lab in Israel), and only the “insights” are shared, satisfying data residency laws.
- De-identification: Identifiable information (name, DOB) is stripped away and replaced with a unique ID before any large-scale research begins, ensuring that researchers can study the DNA without knowing the patient’s identity.
What is Pharmacogenomics (PGx) and why is it included?
Pharmacogenomics is the study of how genes affect a person’s response to drugs. Many clinical genomics platforms now include PGx reporting. This tells a doctor if a patient is a “slow metabolizer” or an “ultra-fast metabolizer” for certain medications. For example, if a patient lacks a specific enzyme (CYP2D6), common painkillers like codeine won’t work for them, or certain antidepressants might cause severe side effects. By checking the patient’s genetic profile before prescribing, doctors can choose the right drug and the right dose the first time, avoiding the dangerous “trial and error” approach to medication.
The Future of Precision Healthcare: Multi-Omic Integration
The future of the Clinical Genomics Platform is moving toward total multi-omic integration. We are beginning to realize that DNA is just the blueprint; it doesn’t tell the whole story. To truly understand human health, we need to look at how that DNA is expressed through RNA (transcriptomics), how those instructions are turned into proteins (proteomics), and how those proteins interact with the environment and our metabolism (metabolomics). This holistic view will allow for population health management on a scale we’ve never seen, identifying health risks before they become illnesses.
Imagine a world where your clinical genomics profile is a living document, integrated with your wearable device data and your electronic health record. If your platform detects a slight change in your proteomic signature that suggests early-stage inflammation or the very first signs of a tumor, it could alert your doctor to intervene months or even years before a traditional test would catch anything. This is the shift from reactive medicine (treating the sick) to proactive medicine (keeping people healthy).
At Lifebit, we are proud to be at the forefront of this revolution. Our federated AI platform provides the secure, real-time access to global data that researchers need to solve the world’s most complex health challenges. We are working with organizations like Genomics England and the NIHR to ensure that the most advanced genomic tools are available to clinicians on the front lines. By breaking down data silos while keeping security at the center, we are making the promise of precision medicine a reality for patients across five continents. The goal is simple: the right treatment, for the right patient, at the right time, every single time.

