The Great Debate OMOP, FHIR, and Your Data Strategy
Stop Losing Research Time: How OMOP and FHIR Data Models Fix It
OMOP and FHIR data models are the two dominant standards changing how healthcare organizations structure, exchange, and analyze patient data. But they serve fundamentally different purposes: FHIR (Fast Healthcare Interoperability Resources) excels at real-time clinical data exchange between live systems, while OMOP (Observational Medical Outcomes Partnership) Common Data Model is purpose-built for large-scale retrospective research and population analytics. The tension between these two approaches isn’t a bug—it’s a feature of a maturing digital health ecosystem.
Historically, healthcare data has been trapped in silos. The transition from paper records to Electronic Health Records (EHRs) in the early 2000s created a “Digital Tower of Babel,” where every hospital system used proprietary formats. HL7 v2 and v3 attempted to solve this, but it wasn’t until the emergence of FHIR that the industry found a truly flexible, web-based standard for interoperability. Simultaneously, the research community realized that clinical data, while useful for treating an individual, was nearly impossible to aggregate for population-level studies without a common structure. This led to the birth of the OMOP CDM, maintained by the OHDSI community.
Quick Comparison:
| Aspect | FHIR | OMOP CDM |
|---|---|---|
| Primary Purpose | Real-time clinical care and data exchange | Retrospective research and observational analytics |
| Design Philosophy | Patient-centric, modular resources via RESTful APIs | Person-centric, relational tables with standardized vocabularies |
| Temporal Precision | Millisecond-level timestamps | Date-level (YYYY-MM-DD) standardization |
| Identifiers | Complex, non-integer identifiers for clinical workflows | Integer-based keys (person_id) for de-identified research |
| Best For | EHR integration, telemedicine, cross-border exchange | Population health studies, AI training, pharmacovigilance |
| Data Structure | Hierarchical (JSON/XML) | Relational (SQL Tables) |
| Governance | HL7 (Health Level Seven International) | OHDSI (Observational Health Data Sciences and Informatics) |
Here’s the reality: healthcare data is fragmented. Every hospital, insurer, and research program uses different codes, formats, and terminologies. FHIR solves the exchange problem—getting data from point A to point B in real time. OMOP solves the analysis problem—harmonizing disparate datasets so you can run the same analytical code across millions of patient records worldwide. Without FHIR, we cannot move data; without OMOP, we cannot understand it at scale.
The challenge? Most organizations need both. They need FHIR to power clinical workflows and OMOP to open up research insights. But changing data between these standards is technically complex, involving identifier management, temporal precision alignment, vocabulary mapping, and privacy-first de-identification. This guide breaks down the technical differences, practical challenges, and real-world solutions for bridging FHIR and OMOP—including the emerging “OMOP on FHIR” approach that’s enabling real-time analytics and AI at scale.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent years building federated AI platforms that harmonize omop and fhir data models across secure, compliant environments for global pharma and public sector organizations. My background in computational biology, genomics, and health-tech entrepreneurship has taught me that the hardest part of healthcare AI isn’t the algorithms—it’s getting the data right.
Must-know omop and fhir data models terms:
Still Choosing One? Why OMOP and FHIR Data Models Need Each Other
To understand why we use both models, we have to look at their DNA. FHIR was born from the need to make Electronic Health Records (EHRs) talk to each other. Developed by HL7, it uses modern web technologies like RESTful APIs to send small “resources” (like a single prescription or a lab result) back and forth. It’s the “text message” of healthcare—fast, lightweight, and focused on the here and now.
The Structural Anatomy of FHIR
FHIR is built on the “80/20 rule”—it focuses on the 20% of data elements that satisfy 80% of clinical use cases. This makes it highly implementable. A FHIR resource, such as Observation or MedicationRequest, is a self-contained packet of information. It includes metadata, extensions for local requirements, and a human-readable narrative. This modularity is perfect for mobile apps and clinician-facing dashboards, but it creates a “graph” of data that is difficult to query for statistical trends across millions of patients.
The Relational Rigor of OMOP
OMOP, on the other hand, is the “history book.” Developed by the Observational Health Data Sciences and Informatics (OHDSI) community, the OMOP Common Data Model was designed for research. It doesn’t care about sending a single record; it cares about organizing billions of records into a standardized format so researchers can ask, “How did this drug affect patients across 30 different countries?”
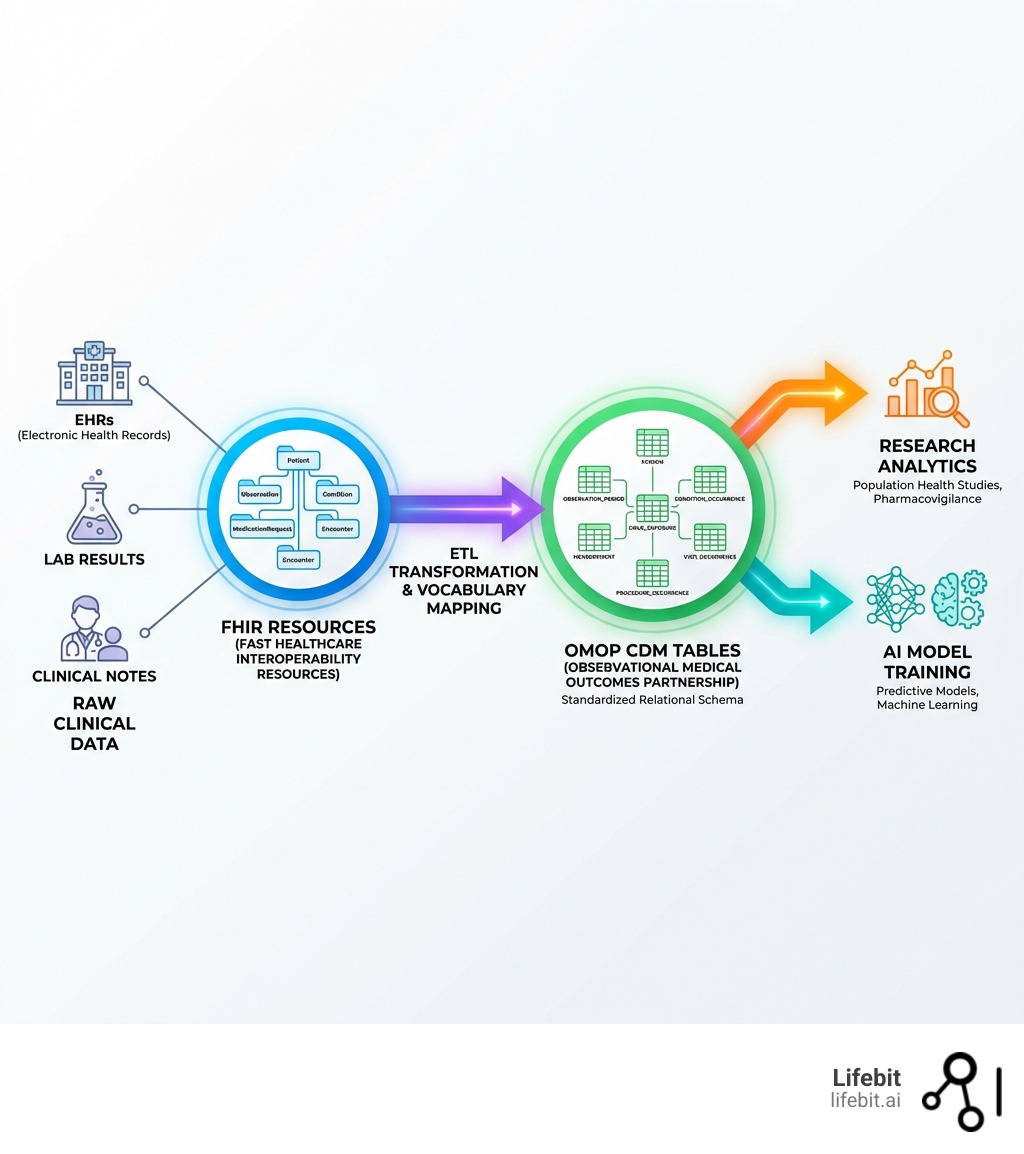
OMOP uses a star-schema relational model. At its center is the PERSON table, linked to clinical event tables like CONDITION_OCCURRENCE, DRUG_EXPOSURE, and MEASUREMENT. This structure is optimized for high-performance SQL queries. When a researcher wants to find all patients with Type 2 Diabetes who were prescribed Metformin, OMOP allows them to join these tables efficiently, regardless of whether the original data came from a hospital in Tokyo or a clinic in London.
Transactional vs. Analytical Intent
The biggest difference lies in “intent.” FHIR is transactional. When a doctor updates a patient’s allergy list, FHIR ensures that information is immediately available to the pharmacy. It supports clinical workflows where millisecond accuracy and the latest data are life-critical. It is designed for “point-of-care” interactions.
OMOP is analytical. It is built for population-level studies and evidence-based medicine. By changing disparate data into a common format, OMOP allows for systematic analysis of databases that were never meant to work together—like insurance claims and hospital EHRs. This harmonization is what makes it possible to generate real-world evidence (RWE) at scale. While FHIR tells you what is happening to a patient now, OMOP tells you what has happened to a population over time.
FHIR to OMOP Mapping: 3 Problems That Break Studies (And How to Fix Them)
If FHIR is a set of loose-leaf papers and OMOP is a structured ledger, moving data from one to the other requires a massive “Extract, Transform, Load” (ETL) effort. This isn’t just a copy-paste job; it’s a fundamental restructuring of information that requires deep domain expertise in both clinical informatics and data engineering.
When we work with omop and fhir data models, we encounter several “roadblocks” in the change process:
- Identifier Mismatch: FHIR uses complex, string-based identifiers (UUIDs or URIs) that are often specific to a single system. OMOP requires integer-based keys (e.g.,
person_id,condition_occurrence_id) for its relational tables to ensure high-speed joins. Mapping these requires a robust “Identity Management” layer that can maintain a crosswalk between the two. - Status and Intent: FHIR resources include “status” fields (e.g.,
active,completed,entered-in-error) and “intent” fields (e.g.,order,plan,proposal). Mapping a “planned” medication in FHIR as a “completed” exposure in OMOP would lead to massive errors in research, such as overestimating drug efficacy or safety risks. The ETL must filter for “finalized” or “completed” statuses to ensure research integrity. - Contextual Gaps: FHIR resources are often fragmented. A single lab result in FHIR might lack the “visit” context that OMOP requires to link that result to a specific hospital stay. To fix this, ETL pipelines must “re-stitch” the data, looking at timestamps to associate observations with the correct
VISIT_OCCURRENCErecord in OMOP.
To help developers, the FHIR to OMOP Implementation Guide provides a continuous build of standards to help bridge these gaps. Tools like OHDSI’s “White Rabbit” (for data profiling) and “Rabbit in a Hat” (for mapping documentation) are essential in this phase.
Mapping Temporal Precision in omop and fhir data models
Time is a major pain point. FHIR captures timestamps down to the millisecond—essential for knowing exactly when a heart rate spiked during surgery. However, the OMOP CDM historically standardized most fields at the date level (YYYY-MM-DD). While recent versions of OMOP have added datetime columns, the core logic of many OHDSI analytical tools still relies on the date.
While this simplicity is great for broad research, it creates “sequencing limitations.” If a patient received two medications on the same day, OMOP might not inherently know which came first without additional “time” columns. During change, we often have to use “controlled imputation”—for example, defaulting a year-only date from a legacy record to January 1st—and carefully documenting these rules to maintain data integrity. This is critical for “Temporal Phenotyping,” where the order of events (e.g., drug A before symptom B) is the entire point of the study.
Managing Identifiers and Privacy
Privacy is where the two models truly diverge. FHIR is designed for clinical care, so it naturally handles Personally Identifiable Information (PII) like names, addresses, and Social Security numbers to ensure the right patient gets the right treatment.
OMOP is designed for de-identified research. Its primary linking mechanism is the person_id, a simple integer. To maintain traceability without compromising privacy, we often build separate “data source tables” that map the internal OMOP IDs back to the original FHIR identifiers in a secure, encrypted environment. This allows us to audit the data or “re-identify” a patient for a clinical trial recruitment (with proper consent) without exposing sensitive patient details in the general research environment. This “firewall” between clinical identity and research data is a cornerstone of modern health data ethics.
OMOP on FHIR: Get Real-Time Analytics Without Breaking Your EHR
The “OMOP on FHIR” approach is the holy grail of health data. It aims to use FHIR as the “ingestion engine” and OMOP as the “analytical engine.” By leveraging FHIR Bundles for real-time data exchange and then automatically changing them into OMOP tables using tools like XSLT or specialized Python-based ETL frameworks, organizations can achieve semantic consistency across their entire data ecosystem.
This is particularly useful for the Vulcan Real-World Data IG, which defines how to pull data from EHRs for clinical research. You can track the different versions of FHIR to OMOP mappings to see how these standards are evolving to support more complex data types like social determinants of health (SDOH) and patient-reported outcomes (PROs).
Standardized Vocabularies: The OHDSI Athena Engine
The “secret sauce” of OMOP is its vocabulary system, managed through the Athena portal. In a raw EHR, one doctor might write “Heart Attack,” another writes “Myocardial Infarction,” and a third uses an ICD-10 code (I21.9). FHIR allows these to be sent as “CodeableConcepts,” but it doesn’t force them into a single standard.
OMOP maps all of these to a single, unambiguous Concept ID (e.g., 312327 for Myocardial Infarction) using standard vocabularies like:
- SNOMED CT: The primary vocabulary for clinical findings, procedures, and body structures.
- RxNorm: The standard for medications, providing a hierarchy from ingredients to branded clinical drugs.
- LOINC: The gold standard for lab tests and measurements, ensuring a “Glucose” test in one lab is the same as a “Glucose” test in another.
- ICD-10-CM/PCS: Used primarily for billing, these are mapped to SNOMED for clinical analysis.
This “source-to-concept” mapping is what allows a researcher to write one query that works across 74 different countries. Without this, global health studies would be impossible because every country uses different coding systems for their healthcare billing.
Bridging Gaps for AI Training
For AI and machine learning, omop and fhir data models are better together. AI models thrive on the structured, longitudinal data found in OMOP. To train a model to predict heart failure, you need years of history, which OMOP provides in a clean, tabular format. However, those models often need to be applied to real-time patients whose data is currently sitting in a FHIR server.
By changing real-time FHIR inputs into the OMOP format, we can perform “real-time classification.” For example, an AI model trained on OMOP data to predict sepsis can take a live stream of FHIR lab results (e.g., lactate levels, white blood cell counts), “OMOP-ify” them to match the model’s expected input features, and provide a prediction to the clinician in seconds. This creates a “Learning Health System” where research insights are immediately cycled back into clinical care.
Who’s Winning With OMOP and FHIR Data Models (FDA, All of Us, EU)
The impact of these models isn’t theoretical—it’s powering the world’s largest health initiatives. The All of Us Research Program, which has enrolled over 849,000 participants, uses OMOP as its primary data standard. By converting EHR data from over 50 health provider organizations into OMOP, they’ve created a massive, searchable database for genomic and clinical research. This allows researchers to study diverse populations that were previously underrepresented in medical studies.
Global Initiatives: EHDEN and DARWIN EU
In Europe, the European Health Data & Evidence Network (EHDEN) has built a federated network of over 180 data partners across 28 countries, all using the OMOP CDM. This network is a key component of DARWIN EU, the European Medicines Agency’s (EMA) platform for generating real-world evidence on the use, safety, and effectiveness of medicines. By using OMOP, the EMA can conduct rapid studies across multiple countries simultaneously, significantly reducing the time it takes to identify potential drug safety issues.
Use Case: Adverse Event Reporting and the FDA
The US FDA is increasingly looking at how to bridge these standards for safety. When a patient has a bad reaction to a drug, that event is often captured in FHIR-based EHR systems. By integrating this into an OMOP database, regulators can automate the generation of Individual Case Safety Reports (ICSR) in XML format. This speeds up pharmacovigilance, allowing the FDA to detect “signals” of drug toxicity months or even years earlier than traditional reporting methods.
Use Case: Genomic Data Integration (G-CDM)
Genomics adds another layer of complexity. Projects are now mapping raw genomic data (VCF files) into FHIR R4 (using the MolecularSequence resource) and then into the OMOP G-CDM (Genomic Common Data Model).
Using standardized HGNC vocabularies for genes and ClinVar for variants, researchers can finally link a specific genetic mutation (e.g., BRCA1) to a patient’s long-term clinical outcomes stored in OMOP. This is the foundation of personalized medicine: knowing not just that a drug works, but which patients it will work for based on their genetic profile. The integration of omop and fhir data models ensures that these genomic insights can move from the research lab (OMOP) back to the patient’s bedside (FHIR).
OMOP and FHIR Data Models: 5 Answers That Save You Time and Rework
1. Can FHIR replace OMOP for research?
In short: No. While FHIR is great for exchange, it isn’t optimized for “bulk” analytical queries. Trying to run a population-level trend analysis on millions of individual FHIR resources would be incredibly slow and computationally expensive. FHIR is like a library where every book is in a separate box; OMOP is like a searchable digital database of every word in those books. OMOP’s relational structure is specifically designed for these high-speed, large-scale analytical tasks.
2. What is the biggest challenge in FHIR-to-OMOP mapping?
The “Identifier Mismatch” and “Contextual Loss” are the heavy hitters. Because FHIR is modular, you often lose the “big picture” of a patient’s visit unless you have a robust ETL process that can stitch those resources back together into a coherent OMOP visit record. Additionally, mapping local “source codes” to OMOP “standard concepts” requires constant maintenance as medical terminologies evolve.
3. How does OMOP handle genomic data compared to FHIR?
FHIR handles genomics through specific profiles (like the MolecularSequence resource) meant for clinical reporting and diagnostic results. It is designed to tell a doctor, “This patient has this mutation.” OMOP uses the G-CDM extension to store genomic variants as standardized concepts, making them searchable alongside clinical data for large-scale association studies (e.g., “Do patients with this variant respond better to this drug?”).
4. What is the role of Data Quality Dashboards (DQD)?
In the OMOP world, the Data Quality Dashboard is a critical tool that runs over 3,000 checks on the data after the ETL process. It ensures that the data is plausible (e.g., no one is 200 years old), conforming (e.g., all dates are in the right format), and complete. FHIR has “Validation” profiles, but they focus on the structure of a single resource rather than the integrity of an entire dataset.
5. How do these models support Federated Learning?
Federated learning allows AI models to be trained on data without the data ever leaving its original location. By having both the source site and the central researcher use the omop and fhir data models, the AI algorithm can be sent to the data, “speak the same language” at every site, and return only the learned parameters. This is essential for maintaining patient privacy across international borders.
Stop Data Chaos: Turn FHIR Into OMOP-Ready Research Data
The debate between omop and fhir data models shouldn’t be about which one is “better.” It’s about how to use them together to build a modern data strategy. In the coming years, the organizations that succeed will be those that can seamlessly move data from the point of care (FHIR) to the point of discovery (OMOP) and back again.
At Lifebit, we help organizations move past the “Digital Tower of Babel.” Our federated AI platform, featuring the Trusted Data Lakehouse and R.E.A.L. (Real-time Evidence & Analytics Layer), provides the infrastructure to harmonize these models at scale. We understand that data harmonization is not a one-time event but a continuous process of ensuring quality, privacy, and utility.
Whether you are running global pharmacovigilance for a top-tier pharma company, managing a national biobank, or building the next generation of AI-driven diagnostics, we ensure your data is secure, compliant, and ready for analysis. Our platform automates the complex ETL processes, handles the vocabulary mappings, and provides the analytical tools needed to turn raw data into life-saving insights.
Don’t let fragmented data slow down your research or compromise patient care. The future of medicine is data-driven, and that data needs a common language. Unlock insights with a Trusted Data Lakehouse and see how we can help you bridge the gap between clinical care and life-saving discovery. Let’s build a world where data works for patients, not against them.

