The NIAID Data Integration Roadmap

Why NIAID Data Catalog Integration Is Critical for Biomedical Research
NIAID data catalog integration enables researchers to search millions of datasets across 50+ repositories through a single, unified portal—eliminating months of manual data hunting and accelerating the path from discovery to therapeutic development. In the modern era of high-throughput biology, the bottleneck is no longer data generation, but data discovery. The National Institute of Allergy and Infectious Diseases (NIAID) recognized this shift and launched the Data Ecosystem to bridge the gap between siloed repositories and actionable insights.
Quick Answer: How to Integrate with NIAID Data Catalog
- Search existing data: Visit the NIAID Data Ecosystem Discovery Portal to search 4.5M+ datasets and 30K+ computational tools without registration. The portal uses advanced indexing to surface results from both specialized and generalist repositories.
- Register your resources: Use the Data Discovery Engine (DDE) guided wizard to create Schema.org-compliant metadata in minutes. This tool is designed for researchers who may not have extensive experience with linked data or JSON-LD.
- Access via API: Query programmatically through the Discovery API for automated workflows and federated analysis. This allows bioinformaticians to integrate NIAID discovery capabilities directly into their custom pipelines.
- Follow FAIR principles: Implement the NIAID Blueprint recommendations for metadata, persistent identifiers (PIDs), and citation standards to ensure your data is Findable, Accessible, Interoperable, and Reusable.
The problem is stark: 82% of biomedical repositories fail to implement proper metadata standards, making their datasets invisible to search engines and impossible to integrate across platforms. While 78% of generalist repositories use Schema.org markup, only 18% of specialized biomedical repositories do—creating a fragmented landscape where critical infectious disease data remains locked in infectious silos. This lack of standardization means that a researcher looking for SARS-CoV-2 transcriptomics data might find results in GEO but miss critical, related clinical trial data stored in a specialized NIAID repository.
NIAID’s solution addresses this crisis head-on. The Data Ecosystem Discovery Portal aggregates metadata from repositories like ImmPort, VDJServer, and AccessClinicalData@NIAID, alongside generalist platforms like Zenodo and Figshare. By focusing on metadata rather than centralizing the data itself, NIAID respects existing data governance and security protocols while dramatically improving findability. The portal exposes standardized metadata that links directly to source repositories, ensuring that the original data owners maintain control over access and licensing.
The impact is already measurable. The outbreak.info Research Library, built on NIAID’s integration schemas, has been accessed over 40,000 times by ~10,000 users across 140+ countries in the past year. The NIAID Systems Biology Consortium registered 345 datasets and 49 computational tools from 18 diverse repositories using the same standardization approach, proving that even complex, multi-omic datasets can be harmonized for discovery.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent over 15 years building federated genomics platforms that align with NIAID data catalog integration standards to power secure, compliant biomedical research at scale. My work on Nextflow and computational biology tools has given me deep insight into why standardized metadata matters—and how to implement it in real-world research workflows. We have seen firsthand how proper integration can reduce the time from data acquisition to insight by up to 60%.
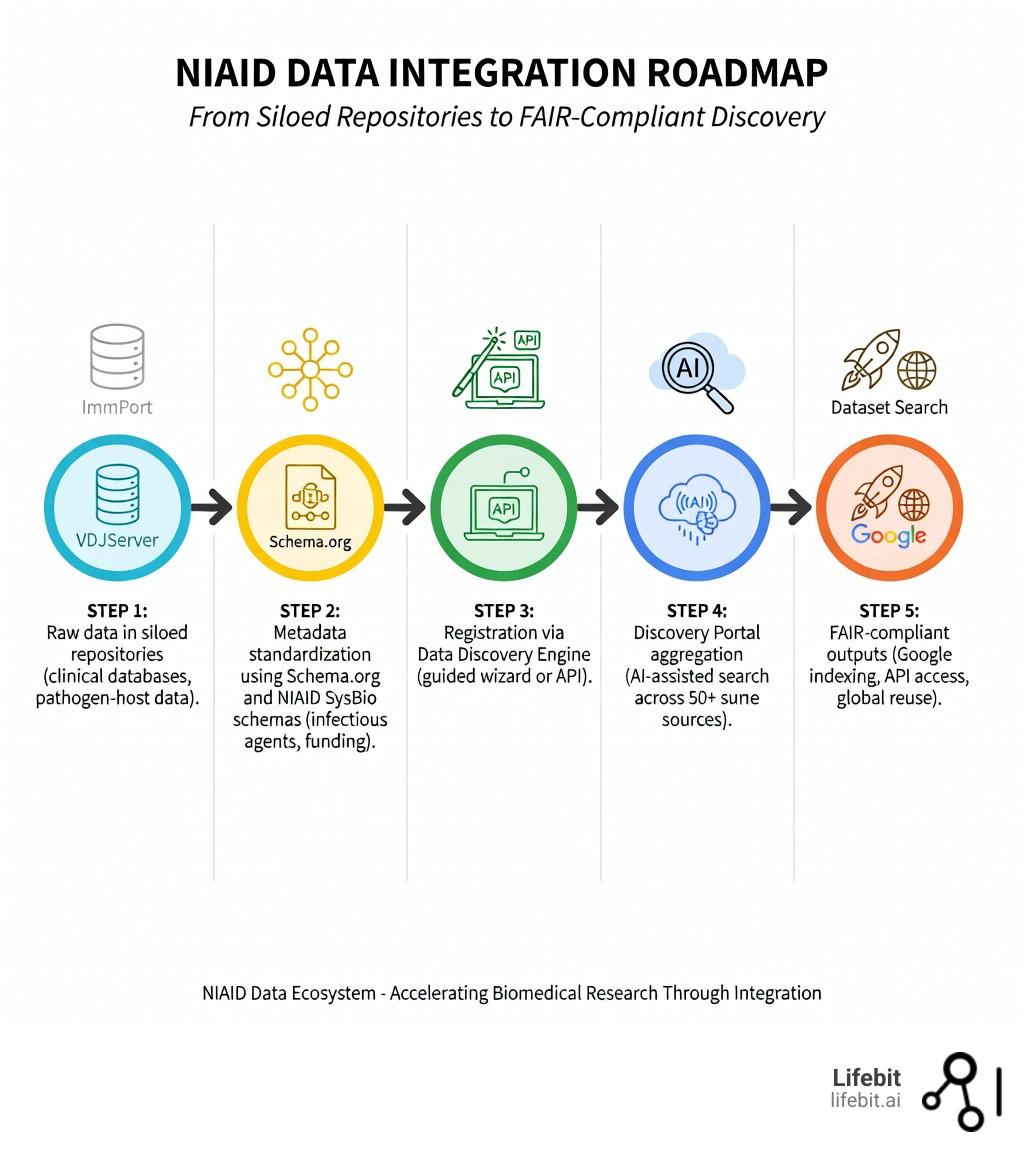
Easy niaid data catalog integration glossary:
Solving the Fragmented Data Crisis with NIAID Data Catalog Integration
Biomedical research is currently drowning in data but starving for insights. The primary culprit? Data silos. For decades, infectious and immune-mediated disease (IID) research has been scattered across hundreds of independent repositories, each with its own language, access rules, and search limitations. This fragmentation is not just a technical nuisance; it is a barrier to scientific progress. When data is siloed, meta-analyses become nearly impossible, and the potential for cross-disciplinary breakthroughs is severely diminished.
The NIAID Data Ecosystem Discovery Portal was built to solve this exact problem. It functions as a centralized metadata registry that doesn’t duplicate data but rather indexes it. By focusing on niaid data catalog integration, the portal allows a researcher to find a clinical trial dataset in one place and a related transcriptomics dataset in another, all through a single search bar. This approach respects the autonomy of original repositories while providing a “Google-like” experience for the NIAID Data Ecosystem.
Bridging 50+ Repositories into One Unified Search
The power of the portal lies in its diversity. It aggregates resources from NIAID-sponsored specialized repositories, such as ImmPort (immunology), VDJServer (repertoire sequencing), and AccessClinicalData@NIAID (clinical trials). These repositories are the lifeblood of IID research, containing high-quality, curated data that is often difficult to find through standard search engines.
However, it doesn’t stop at NIAID-funded projects. To provide a truly comprehensive view of the IID landscape, the portal also indexes generalist repositories like Zenodo, Figshare, and Dryad. By normalizing the metadata from these 50+ sources, the portal creates a unified discovery layer where over 4.5 million datasets are searchable by pathogen, host species, or measurement technique. This means that a search for “Influenza A” will return results from specialized immunology databases alongside relevant software tools from GitHub and generalist datasets from Zenodo.
Why 82% of Biomedical Repositories Fail at Data Discovery
Research shows a massive gap in how data is shared. While 78% of generalist data repositories use Schema.org—the industry standard for making web content discoverable—only 18% of specialized biomedical repositories do the same. This discrepancy is largely due to the complexity of biomedical data, which often requires specialized ontologies and vocabularies that standard Schema.org properties do not support.
Furthermore, even when repositories do use standardized schemas, they often use them inconsistently. A study found that while the Schema.org “Dataset” type specifies 124 properties, most repositories use fewer than 30. This lack of detail makes it impossible for automated systems to understand the context of the research, such as which infectious agent was studied, the specific strain of the pathogen, or the funding source. This is why metadata standardization is the backbone of the NIAID integration strategy. Without rich, standardized metadata, the “Findable” part of FAIR remains an unfulfilled promise.
Standardizing Research with NIAID SysBio and Schema.org
To fix the “invisible data” problem, NIAID developed the SysBio schemas. These are extensions of Schema.org customized specifically for the needs of infectious disease researchers. By using standard formats like JSON-LD (JavaScript Object Notation for Linked Data), NIAID ensures that data is not just findable by humans, but “machine-actionable.” This means that AI agents and automated discovery tools can parse the metadata to understand the relationships between different research outputs.
This standardization aligns with the FAIR Principles—Findable, Accessible, Interoperable, and Reusable. For software and computational tools, NIAID also leverages CodeMeta, ensuring that the scripts used to analyze data are just as discoverable as the data itself. This holistic approach ensures that the entire research lifecycle—from raw data to final analysis code—is documented and discoverable.
Key Features of the NIAID SysBio Dataset Schema
The NIAID SysBio Dataset schema takes the generic Schema.org “Dataset” and adds high-value fields for the IID community. These fields are not just optional extras; they are critical for the type of complex querying required in modern immunology. Key features include:
- InfectiousAgent: A mandatory field to identify the pathogen using standardized NCBI Taxonomy IDs (e.g., SARS-CoV-2). This prevents confusion between common names and scientific nomenclature.
- Funding: A required field to track research back to specific NIH grants and sponsors. This allows funders to see the direct output of their investments.
- Ontology Mapping: Links terms to standardized vocabularies like MONDO for diseases, Uberon for anatomy, and the Ontology for Biomedical Investigations (OBI) for assay types.
- HealthCondition: A specific property to describe the clinical state of the host, allowing researchers to filter for datasets involving specific comorbidities or disease severities.
These properties are essential for NIAID Data Harmonization, allowing datasets from different sources to be compared and combined with high confidence. For example, a researcher can query for all datasets involving “Type 1 Diabetes” and “RNA-Seq” across five different repositories, knowing that the metadata has been harmonized to the same standards.
Linking Software and Data via NIAID Data Catalog Integration
Data is only half the story; the tools used to analyze it are just as critical. The NIAID SysBio ComputationalTool schema facilitates the linkage of datasets to the software used to generate them. This promotes reanalysis and reproducibility—two cornerstones of modern science. In many cases, the “how” of a study is just as important as the “what,” and by making software discoverable, NIAID enables other researchers to validate findings or apply the same methods to new data.
By standardizing software metadata, researchers can find the exact version of a pipeline used in a specific study. We can see these schemas in action within the “Schema Playground,” a tool that allows developers to visualize and test how their metadata will appear in the ecosystem. This interactive environment helps bridge the gap between data producers and the discovery portal.
How to Register Your Resources Using the Data Discovery Engine (DDE)
If you are a researcher with a dataset sitting in a repository that isn’t Schema.org-compliant, you don’t have to move your data. This is a critical point: niaid data catalog integration does not require data migration. The Data Discovery Engine (DDE) provides a bridge. It allows you to register your metadata so it can be indexed by the NIAID portal while your actual data stays exactly where it is, whether that’s on a local server, a university repository, or a cloud bucket.
| Feature | DDE Registration | Manual Repository Migration |
|---|---|---|
| Effort | Low (minutes) | High (weeks/months) |
| Data Location | Stays in original home | Must be re-uploaded |
| Compliance | Automated Schema.org generation | Manual metadata mapping |
| Discoverability | Instant indexing in NIAID & Google | Depends on new repository |
| Governance | Maintains original access controls | Subject to new repository rules |
Step-by-Step NIAID Data Catalog Integration for Researchers
For individual researchers, the process is designed to be painless and requires no coding knowledge. The DDE acts as a “TurboTax for metadata,” guiding users through the necessary fields:
- Use the Guided Wizard: Navigate to the Knowledge Center and follow the DDE registration flowchart. The wizard will ask questions about your research in plain language.
- Fill the Fields: The wizard asks for basic info (title, creator, funding). It automatically formats this into JSON-LD. It also provides autocomplete suggestions for pathogens and diseases based on official ontologies.
- Validate: Use the compatibility checker to ensure your metadata meets NIAID standards. This step catches common errors, such as missing mandatory fields or invalid URL formats.
- Register: Once submitted, your dataset becomes discoverable in both the NIAID Portal and Google Dataset Search. The DDE generates a unique identifier for your metadata record, which can be cited in publications.
Leveraging the Discovery API for Programmatic Access
For developers and bioinformaticians, the Discovery API offers a powerful way to interact with the ecosystem. Built on the OAS3 (OpenAPI Specification) standard, the API provides dedicated endpoints for metadata retrieval and complex querying. This is particularly useful for building custom dashboards or integrating NIAID discovery into institutional data portals.
Users can track changes via the API changelog and integrate these services into their own analytical pipelines. For example, a script could be written to automatically alert a research team whenever a new dataset related to “Zika virus” is registered in the ecosystem. This programmatic access is what enables a Federated Data Ecosystem, where tools can “talk” to the NIAID catalog to find relevant data for automated reanalysis without human intervention.
Real-World Impact: From Outbreak Tracking to Global Networks
The theoretical benefits of niaid data catalog integration are clear, but the real-world impact is even more impressive. During the COVID-19 pandemic, the ability to rapidly find and reuse data was a matter of global security. The NIAID Data Ecosystem provided the infrastructure needed to turn a flood of disparate data into a coherent stream of information.
The outbreak.info Research Library is a prime example. By adapting the NIAID SysBio schemas, this project was able to harvest research from 16 different sources in real-time, including bioRxiv, medRxiv, and various clinical trial registries. This provided a unified dashboard for the global research community to track the latest variants, vaccine trials, and epidemiological shifts. Without the standardized metadata provided by the NIAID integration, this level of real-time aggregation would have required hundreds of hours of manual curation every week.
Case Study: Tracking 40,000+ COVID-19 Research Interactions
The outbreak.info platform demonstrated that when data is standardized, it moves faster. In just one year, the library was accessed by 10,000 users in over 140 countries. This wasn’t just about viewing data; it was about reuse. Because the metadata was FAIR-compliant, researchers could quickly determine if a dataset was suitable for their specific reanalysis without downloading massive files first. For instance, a researcher could filter for “Omicron variant” and “neutralization assays” and find exactly the datasets they needed in seconds.
Another success story is the NIAID Systems Biology Consortium (SBC). The SBC faced the challenge of integrating data from 18 different research centers, each using different data formats and storage solutions. By adopting the NIAID SysBio schemas and the DDE, the consortium was able to register over 300 datasets, making them discoverable to the wider scientific community. This has led to new collaborations and secondary analyses that would not have been possible if the data had remained hidden in institutional silos. This is the ultimate goal of a Federated Data Ecosystem NIAID: making the world’s research-ready data accessible at the click of a button.
Frequently Asked Questions about NIAID Data Catalog Integration
How do I register a non-Schema.org compliant repository?
You don’t need to overhaul your entire repository or change your underlying database structure. Researchers can use the Data Discovery Engine (DDE) guided wizard to manually enter metadata for individual datasets. The tool then generates the necessary JSON-LD for the NIAID portal to crawl. This “metadata-first” approach allows your data to remain in its original location while becoming fully discoverable globally. For larger repositories, NIAID provides technical support to help implement automated metadata harvesting via APIs.
What are the benefits for NIAID program sponsors?
For sponsors and funders, standardized integration is a game-changer for portfolio management and impact assessment. It allows them to automatically track research outputs and identify data gaps in real-time. Instead of relying on manual (and often delayed) self-reporting from grantees, sponsors can use the catalog to see exactly what datasets and tools have been produced by their funded projects. This transparency helps in making informed decisions about future funding priorities and ensures that public investments in research are yielding discoverable results.
Is registration required to search the Discovery Portal?
No. The NIAID Data Ecosystem Discovery Portal is completely free and requires no registration for searching or browsing. You can search millions of datasets right now. However, keep in mind that while the metadata is public, the actual data is held by individual providers (like ImmPort) who may require you to register or obtain specific approvals (like DAIT approval) before downloading sensitive clinical information. The portal acts as the “card catalog,” but you still need to visit the “stacks” to get the book.
How does NIAID ensure the quality of the metadata?
The NIAID Data Ecosystem employs both automated and manual validation processes. The DDE includes built-in validation rules that check for adherence to the SysBio schemas. Additionally, the NIAID Office of Data Science and Strategy (ODSS) works with data stewards to ensure that the indexed metadata is accurate and up-to-date. Researchers are also encouraged to provide feedback on search results to help refine the portal’s ranking algorithms and discovery capabilities.
Conclusion
The journey toward a fully integrated biomedical landscape is well underway, and niaid data catalog integration is the cornerstone of this transformation. By moving away from fragmented silos and toward a unified NIAID Blueprint, the research community is making “open science” a practical reality rather than just a theoretical ideal. The ability to find, access, and reuse data across institutional and geographic boundaries is no longer a luxury—it is a necessity for addressing the complex health challenges of the 21st century.
At Lifebit, we believe that the future of medicine depends on this kind of seamless, secure data flow. Whether it’s mastering Cloud Nine Compliance for federal data or building the next generation of federated health services, we are committed to the standards that make NIAID’s data ecosystem a success. The work being done today to standardize metadata and integrate catalogs will provide the foundation for the AI-driven discoveries of tomorrow. By adopting these integration roadmaps today, we ensure that the breakthroughs of tomorrow are built on a foundation of findable, accessible, and truly reusable data. The era of “dark data” is coming to an end, and the era of integrated, discoverable science is just beginning.

