The Robot Will See Your Molecule Now: AI in Pharma R&D

How AI in Pharmaceutical R&D Cuts Preclinical Costs by 30% and Saves 7 Years
AI in pharmaceutical R&D is reshaping how we find, design, and bring new drugs to market—but the change is still unfolding. Here’s what you need to know:
Key Applications of AI in Pharma R&D:
- Target Discovery: Identifying disease-causing proteins from the 89% of the human proteome not yet mapped to small molecules
- Molecular Design: Navigating the vast chemical space of 10^60 possible compounds to find promising drug candidates
- Virtual Screening: Predicting which molecules will bind to targets without expensive lab tests
- ADMET Prediction: Forecasting absorption, distribution, metabolism, excretion, and toxicity early in development
- Clinical Trial Optimization: Matching patients to trials, predicting outcomes, and reducing failure rates from 95% to more manageable levels
- Protein Structure Prediction: Tools like AlphaFold 2 achieving 92.4% accuracy in predicting how proteins fold in 3D
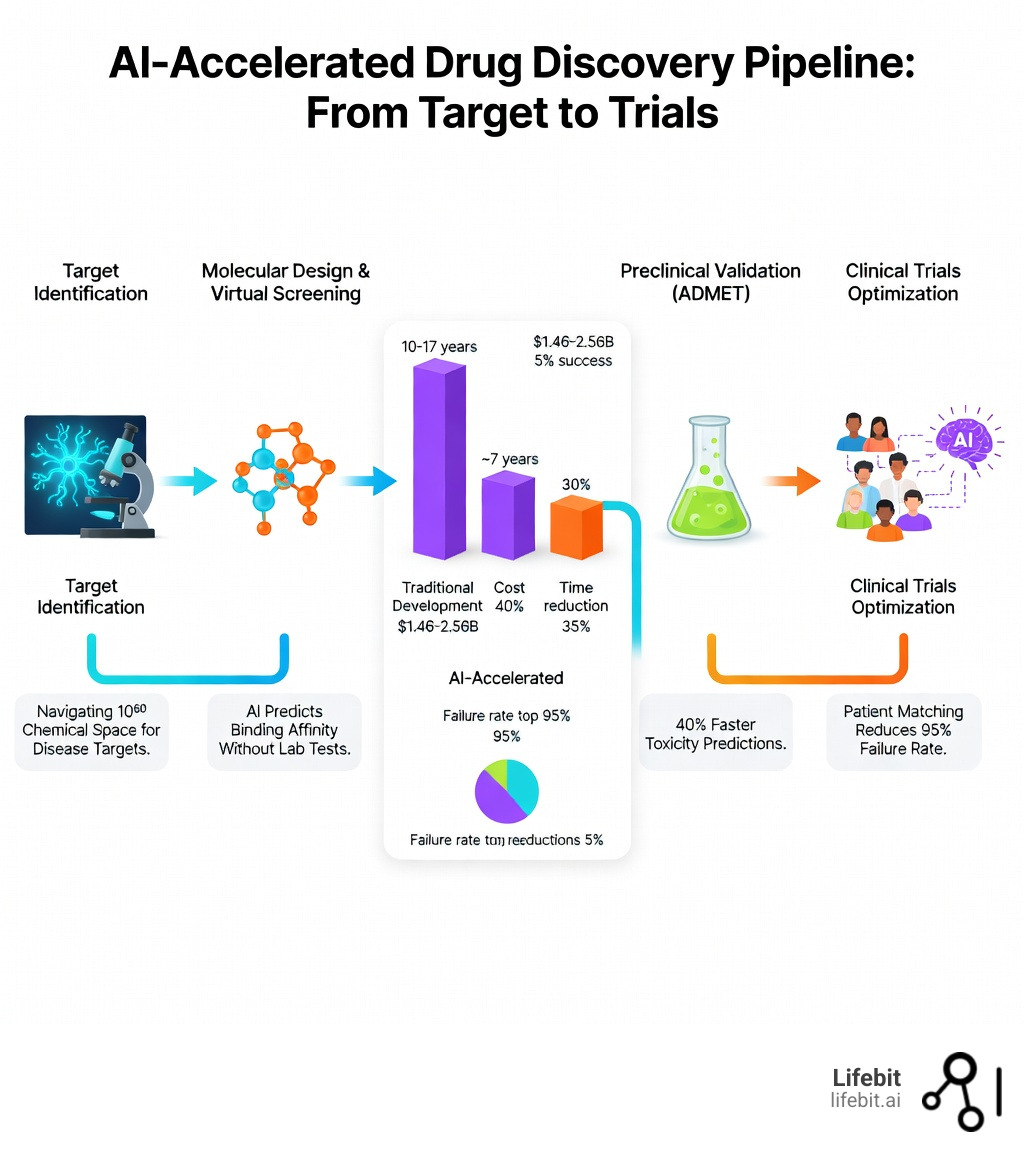
The Stakes: Traditional drug development costs between $1.46 billion and $2.56 billion per drug and takes 10-17 years. Even after passing Phase I trials, only 5% of candidates reach the market. AI promises to cut these timelines by 40% and costs by 30% in preclinical stages alone.
Current Reality: Over 500 drug submissions with AI components have reached the FDA since 2016. While no AI-generated drug has received full FDA approval yet, dozens of AI-designed molecules are advancing through clinical trials—including candidates for idiopathic pulmonary fibrosis, fragile X syndrome, and inflammatory diseases.
The challenge isn’t whether AI works in pharma R&D—it’s how to scale it across siloed datasets, integrate wet and dry lab workflows, and build trust in black-box models while navigating evolving regulatory frameworks.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent over a decade building federated platforms that enable secure, compliant AI in pharmaceutical R&D across distributed health and genomic datasets. Our work powers real-time analytics for global pharma and public sector organizations without moving sensitive data.
Stop Guessing: The 3 Data Pillars Powering AI in Pharmaceutical R&D
To understand how we moved from Penicillin (discovered via a moldy petri dish and a bit of luck) to AI-designed inhibitors, we need to look at the three fundamental elements driving AI in pharmaceutical R&D: data, computation, and algorithms.
- Data Quality and Multi-Omics Integration: This is the “fuel.” It includes everything from public databases like the Protein Data Bank (PDB) to proprietary R&D datasets and Real-World Evidence (RWE). Modern AI requires more than just chemical structures; it needs “multi-omics” data—genomics, proteomics, transcriptomics, and metabolomics. Without high-quality, harmonized data that accounts for biological variability, even the best algorithms produce “hallucinations” or biologically irrelevant molecules.
- High-Performance Computation and Quantum Readiness: Processing 10^60 possible small molecules requires massive GPU power. Cloud-based resources now allow us to run simulations in hours that used to take years. Furthermore, the industry is preparing for the “Quantum Leap,” where quantum computing could theoretically simulate molecular interactions at the sub-atomic level with perfect accuracy, a feat currently impossible for classical computers.
- Advanced Algorithms and Generative Chemistry: We’ve evolved from simple Quantitative Structure-Activity Relationship (QSAR) models—which basically tried to correlate a molecule’s shape with its effect—to deep learning models that can “invent” entirely new chemistry. These models don’t just search a database; they use latent space representation to navigate the “chemical dark matter” where no human chemist has ever ventured.
Historical Evolution: From Serendipity to Strategy
Before the 1980s, drug discovery was largely a game of random screening. You’d throw thousands of compounds at a disease and see what stuck. By the 1990s, we began using Computer-Aided Drug Design (CADD) and high-throughput screening. However, the real “paradigm shift” occurred around 2018, when AI transitioned from a theoretical concept to a practical tool capable of generating clinical-ready molecules.
According to Scientific research on the future of AI in pharmaceuticals, this evolution is finally addressing the “Grand Challenge” of R&D productivity. By heavily investing in AI, the pharmaceutical industry could see a return on investment (ROI) increase of more than 45%. This is critical because Eroom’s Law (Moore’s Law spelled backward) suggests that drug discovery has become exponentially more expensive and slower over time; AI is the first technology with the potential to reverse this trend.
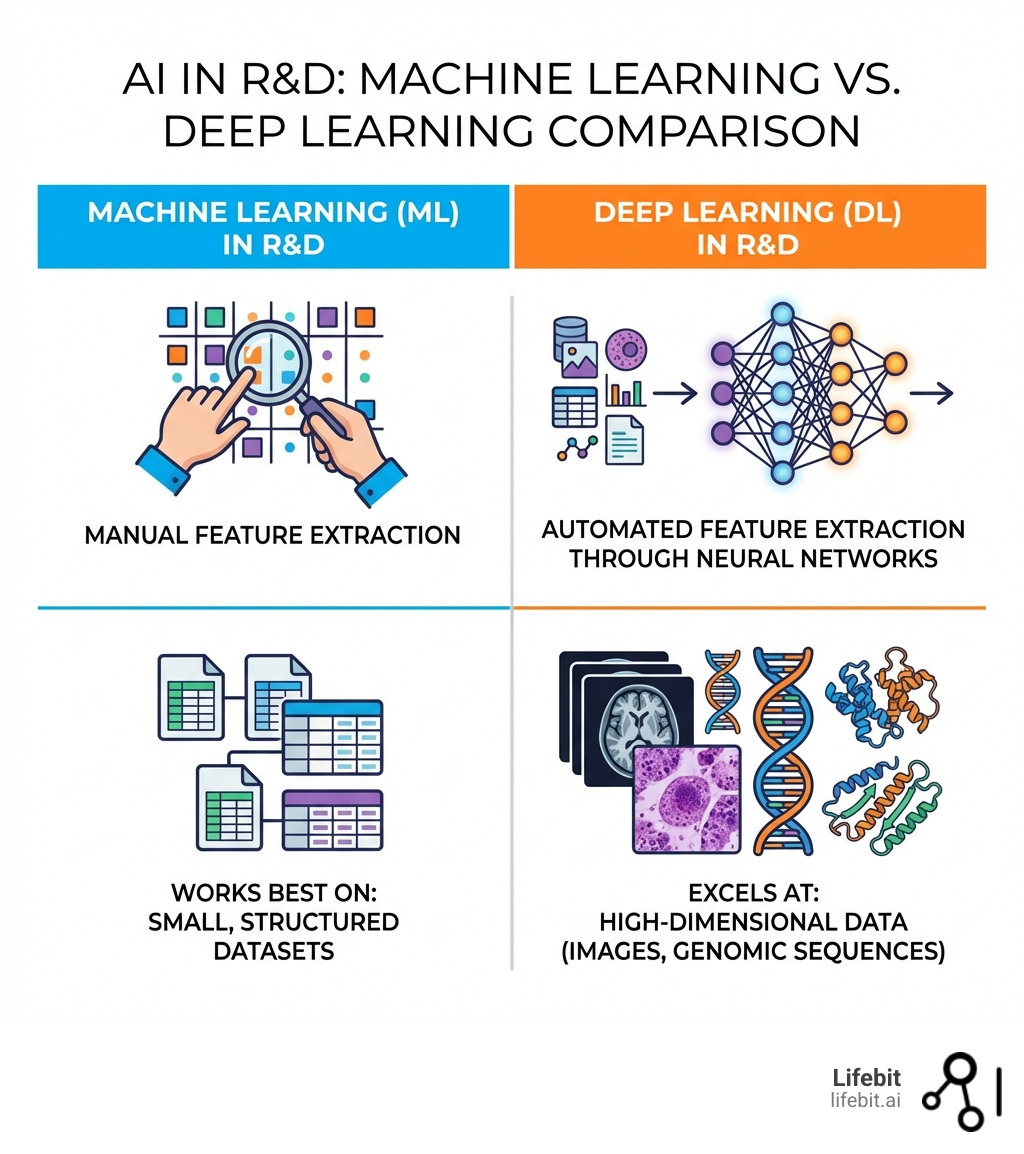
Deep Learning Architectures for AI in Pharmaceutical R&D
Deep learning (DL) is the “brain” behind the modern robot scientist. Unlike traditional machine learning, which requires humans to tell the computer which “features” of a molecule matter (like the number of carbon atoms), DL architectures learn directly from the raw data.
- CNNs (Convolutional Neural Networks): Mostly used for image-based data. In R&D, this means analyzing pathology slides at a speed and precision no human can match, or identifying subtle “phenotypic” changes in cells treated with experimental drugs.
- RNNs and Transformers: Excellent for sequential data. Since molecules can be represented as strings of text (called SMILES strings), these models help “read” and “write” new chemical formulas. Transformers, the tech behind ChatGPT, are now being used to create “Chemical Language Models” that understand the grammar of molecular stability.
- GNNs (Graph Neural Networks): These are the current gold standard. They treat molecules as graphs where atoms are nodes and bonds are edges. This captures the actual spatial 3D relationship of a molecule better than a text string. Scientific research on graph neural networks shows how these models are revolutionizing molecular property prediction by accounting for the geometric constraints of the binding pocket.
- GANs and Diffusion Models: Imagine two AIs—one trying to “counterfeit” a drug molecule and another trying to catch the fake. Diffusion models, similar to those used in AI art generators like Midjourney, are now being used to “denoise” random atomic arrangements into highly stable, novel drug candidates.
Target Discovery and Small Molecule Design
The human proteome is like a massive library where 89% of the books are written in a language we don’t fully understand yet. Only about 11% of human proteins have been successfully “annotated” or targeted with small molecule probes. This leaves a massive “undruggable” space that AI is beginning to unlock.
AI helps us bridge this gap through:
- Virtual Screening: Instead of testing 1 million compounds in a “wet” lab (which would cost millions and take years), we use AI to screen them “in silico” (on a computer). This narrows the field to the top 100 candidates with the highest binding affinity.
- Lead Optimization: AI tweaks the structure of a promising molecule to make it more effective or less toxic. For instance, it might suggest replacing a specific atom to improve how the drug dissolves in the bloodstream.
- ADMET Prediction: One of the biggest reasons drugs fail is that they look great in a dish but turn out to be toxic to the liver or don’t absorb well in the human gut. Scientific research on ADMET prediction platforms highlights how new platforms like ADMETlab 3.0 provide decision support early in the pipeline, potentially saving billions in failed trials by identifying “toxicophores”—structural patterns known to cause adverse reactions—before a single dose is manufactured.
Solve the 50-Year Protein Folding Mystery in Seconds with AI
If you want to stop a disease, you usually need to find a protein “lock” and design a molecular “key” to fit it. But proteins are messy—they are long chains of amino acids that fold into complex, tangled 3D shapes. For 50 years, predicting that shape based solely on the amino acid sequence was one of biology’s greatest mysteries, known as the “Protein Folding Problem.”
Then came AlphaFold 2, developed by Google DeepMind. At the CASP14 competition, it achieved a median Global Distance Test (GDT) score of 92.4, essentially solving the problem. This wasn’t just a technical win; it was a Nobel Prize-winning breakthrough that gave us the structures for nearly all 200 million proteins known to science. Before AlphaFold, determining a single protein structure required years of arduous work using X-ray crystallography or Cryo-Electron Microscopy.
AlphaFold 3: The Next Frontier in Molecular Interaction
While AlphaFold 2 focused on single proteins, AlphaFold 3 represents a massive leap forward. It can predict how proteins interact with a wide array of other biological components, including DNA, RNA, and—most importantly for R&D—ligands (drug molecules). Scientific research on AlphaFold 3 upgrades explains how this model covers 98.5% of the human proteome, allowing researchers to model the entire “interactome” of a cell. This means we can now predict not just if a drug will hit its target, but if it will accidentally hit 50 other proteins, causing side effects.
De Novo Protein Design
Beyond just predicting existing shapes, AI is now being used for de novo design—creating proteins that have never existed in nature. Using tools like ProteinMPNN and RFdiffusion, scientists can specify a desired function (e.g., “bind to this specific virus spike”) and the AI will generate the exact amino acid sequence required to build that protein from scratch. This is opening the door to entirely new classes of biologics and vaccines that are more stable and potent than anything found in the natural world.
Stop Losing $1.4B Per Trial: How AI in Pharmaceutical R&D Fixes Clinical Failure
The “Valley of Death” in pharma is the clinical trial phase. This is where 95% of oncology drugs fail, often at a loss of $800 million to $1.4 billion per failed Phase III trial. AI in pharmaceutical R&D acts as a safety net here, transforming trials from a high-stakes gamble into a data-driven science.
Precision Patient Matching and Recruitment
One of the primary reasons trials fail or are delayed is poor recruitment. AI uses Natural Language Processing (NLP) to scan millions of Electronic Health Records (EHRs) and match the right patients to the right trials in seconds. This is particularly vital for rare diseases, where finding a single eligible patient can be like finding a needle in a haystack. AI can also predict “patient churn”—identifying which participants are likely to drop out of a study so that investigators can intervene early.
Digital Twins and Synthetic Control Arms
Perhaps the most revolutionary application in clinical trials is the use of Synthetic Control Arms (SCAs). Traditionally, a trial requires a control group of patients who receive a placebo. This is expensive, ethically complex in terminal illnesses, and difficult to recruit for. AI can generate a “Digital Twin” of a patient based on historical data, allowing researchers to simulate the control group. This reduces the number of human participants needed, slashes costs, and accelerates the path to approval.
Drug Repurposing: New Life for Old Molecules
AI can find new uses for old drugs, a process known as drug repurposing. Since these drugs have already passed safety tests, they can skip straight to Phase II or III trials, saving nearly a decade of development time. A famous example is Sildenafil, which was originally a heart drug before data analysis repurposed it for erectile dysfunction. More recently, Scientific research on AI for cancer drug discovery shows how AI is identifying “hidden” signatures in genomic data to predict which patients will actually benefit from immunotherapy, allowing drugs that failed in general populations to succeed in specific, genetically-defined subgroups.
Real-World Impact: AI Success Stories in Pharma R&D
Is this all just hype? Not anymore. We are seeing “first-in-class” molecules designed entirely by AI entering human trials at an unprecedented pace.
- Idiopathic Pulmonary Fibrosis (IPF): Insilico Medicine used AI to identify a novel target and design a new drug candidate in under 18 months—a process that usually takes five years and tens of millions of dollars. That drug is now in Phase IIa trials, proving that AI-generated molecules are safe for human consumption.
- NLRP3 Inhibitors: These are crucial for treating inflammatory diseases like Alzheimer’s and Parkinson’s. Scientific research on AI-driven NLRP3 inhibitors details the discovery of SN3-1, a potent inhibitor found via deep learning that shows massive potential for treating chronic inflammation by crossing the blood-brain barrier more effectively than previous candidates.
- Wilson’s Disease: AI platforms have screened oligonucleotide candidates and moved them into the pipeline in record time, demonstrating that AI is equally effective for complex biologics as it is for small molecules.
Access Global Health Data Without Moving It: Solving the AI Trust Gap
Despite the excitement, AI faces a significant “trust gap.” Many models are “black boxes”—they give a prediction, but they can’t explain why they reached that conclusion. In medicine, “because the computer said so” isn’t a valid reason to give a patient an experimental drug or to invest $500 million in a clinical trial.
The Problem of Bias and Data Silos
If an AI is trained on data that mostly represents one demographic (e.g., men of European descent), its predictions for everyone else will be biased. For example, the “gender data gap” often leads to AI systems that poorly estimate drug safety for women because they weren’t represented in the training sets. Furthermore, the world’s most valuable health data is locked in silos—hospitals and research centers are rightfully hesitant to share sensitive patient data due to GDPR, HIPAA, and other privacy regulations.
Scientific research on AI bias and pharmaceutical solutions suggests that Explainable AI (xAI) is the only way to build the transparency needed for regulatory approval. xAI techniques like SHAP (SHapley Additive exPlanations) allow scientists to see which specific molecular features led the AI to predict toxicity, turning the “black box” into a “glass box.”
How Lifebit Fixes the “Data Problem” with Federated AI
At Lifebit, we believe the answer isn’t moving data into one giant, risky pile. Instead, we use Federated AI. This approach allows the AI model to travel to the data, rather than the data traveling to the model.
Our platform creates Trusted Research Environments (TREs) where researchers can run their AI models directly where the data lives—whether that’s in a government biobank in London, a hospital in New York, or a genomic center in Singapore. This architecture provides three key benefits:
- Privacy: Sensitive patient data never leaves its original secure environment.
- Compliance: It automatically adheres to local data residency laws.
- Diversity: By connecting disparate datasets globally, we can train AI on the most diverse populations possible, eliminating demographic bias and ensuring that the next generation of drugs works for everyone, regardless of ethnicity or gender.
AI in Pharmaceutical R&D: Your Top Questions Answered
How does AI reduce the cost of drug development?
AI reduces costs by failing early and failing “cheaply.” By predicting toxicity (ADMET) and efficacy before a single molecule is synthesized in a lab, companies avoid spending hundreds of millions on clinical trials that are destined to fail. AI-enabled workflows can save up to 30% of total preclinical costs and reduce the time spent in the discovery phase by several years.
What is the success rate of AI-designed drugs in clinical trials?
While it’s too early for a definitive industry-wide percentage (since many are still in Phase I/II), early data suggests that AI-selected candidates have a much higher “hit rate” in the lab. The goal is to move the Phase I-to-market success rate from the current 5% toward 20% or higher. If AI can even double the success rate to 10%, it would fundamentally change the economics of the entire industry.
Can AI predict the toxicity of new drug candidates?
Yes. Deep learning models like DeepTox have already won international challenges by accurately predicting the toxicity of over 12,000 compounds. These models analyze chemical structures to find “toxicophores”—structural patterns known to cause adverse reactions. This allows researchers to eliminate dangerous compounds before they ever reach animal or human testing.
Will AI replace human chemists and biologists?
No. AI is a tool that augments human expertise. While AI is excellent at pattern recognition and navigating vast data spaces, human scientists are still required to set the parameters, interpret complex biological contexts, and make the final ethical and strategic decisions. The future belongs to the “augmented scientist.”
How do regulators like the FDA view AI in R&D?
The FDA is actively encouraging the use of AI while maintaining rigorous safety standards. They have released a discussion paper on AI/ML in drug development and are working on frameworks for “predetermined change control plans,” which would allow AI models to update and improve in real-time as they receive more data, provided they stay within certain safety bounds.
The 45% ROI Boost: Why AI in Pharmaceutical R&D Is the New Standard
The future of AI in pharmaceutical R&D isn’t about replacing scientists; it’s about giving them “superpowers.” We are moving toward a world of Self-Driving Labs, where AI plans the synthesis of a molecule, robotic arms mix the chemicals, and the system uses real-time sensors to learn from its own mistakes. If a reaction fails, the AI immediately analyzes why and tries a different approach, running 24/7 without human intervention.
This shift is not just a luxury; it is a necessity. As diseases become more complex and the “low-hanging fruit” of drug discovery is exhausted, the industry must embrace these technologies to survive. Regulators like the FDA and EMA are already adapting, creating risk-based frameworks to ensure these technologies are safe and effective. For pharma companies, the potential is clear: a 45% increase in ROI and the ability to bring life-saving treatments to patients in years rather than decades.
At Lifebit, we provide the infrastructure to make this possible. By connecting global biomedical data through our Trusted Data Lakehouse and federated analytics, we ensure that the next blockbuster drug isn’t hidden in a data silo. We are helping the industry move from a model of “discovery by chance” to “discovery by design.”

