The Ultimate Guide to Accelerating Drug Repurposing with AI
Why 300 Million Patients Are Counting on Drug Repurposing AI Right Now
Drug repurposing AI is the use of artificial intelligence to find new therapeutic uses for drugs that are already approved — dramatically cutting the time and cost it takes to get treatments to patients.
Here’s what that means in practice:
| What AI Does | Why It Matters |
|---|---|
| Screens thousands of existing drugs against thousands of diseases | Covers ground no human team could manually |
| Identifies shared biological mechanisms across conditions | Finds non-obvious matches traditional methods miss |
| Predicts contraindications before clinical trials | Reduces patient risk and trial failures |
| Generates explainable rationales for each prediction | Builds clinician trust and supports hypothesis testing |
| Validates predictions against real-world patient data | Bridges the gap between computation and clinic |
The numbers tell a stark story. There are over 7,000 rare diseases affecting 300 million people worldwide. Only 5–7% of those diseases have an FDA-approved drug. Traditional drug development takes over a decade and costs billions — far too slow and too expensive for the vast majority of these conditions.
Repurposing existing drugs changes that equation entirely. Because safety profiles are already established, developers can skip early-stage trials and move faster. In fact, nearly 30% of FDA-approved drugs have already picked up at least one additional indication after their initial approval — proof that the biology supports this approach.
AI makes repurposing scalable. Where traditional methods rely on expert opinion, literature reviews, and chance observations, AI can simultaneously analyze millions of biological relationships, patient records, and molecular pathways to surface candidates that would otherwise stay hidden for years.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, and with over 15 years in computational biology, federated data infrastructure, and AI-powered biomedical research, I’ve seen how drug repurposing AI is reshaping what’s possible for patients with the fewest options. In this guide, I’ll walk you through exactly how these systems work — and how to put them to use.
Key terms for drug repurposing ai:
Why Drug Repurposing AI is the Only Hope for Rare Diseases
For the 300 million people worldwide living with a rare disease, the traditional pharmaceutical model is fundamentally broken. According to scientific research on the global burden of rare diseases, the vast majority of these 7,000+ conditions remain untreated because the “one drug, one disease” development path is too costly for small patient populations. The economic burden is equally staggering; a study by the EveryLife Foundation for Rare Diseases estimated the total economic impact of 373 rare diseases in the US alone at nearly $1 trillion annually, driven by direct medical costs and indirect costs like lost productivity for caregivers.
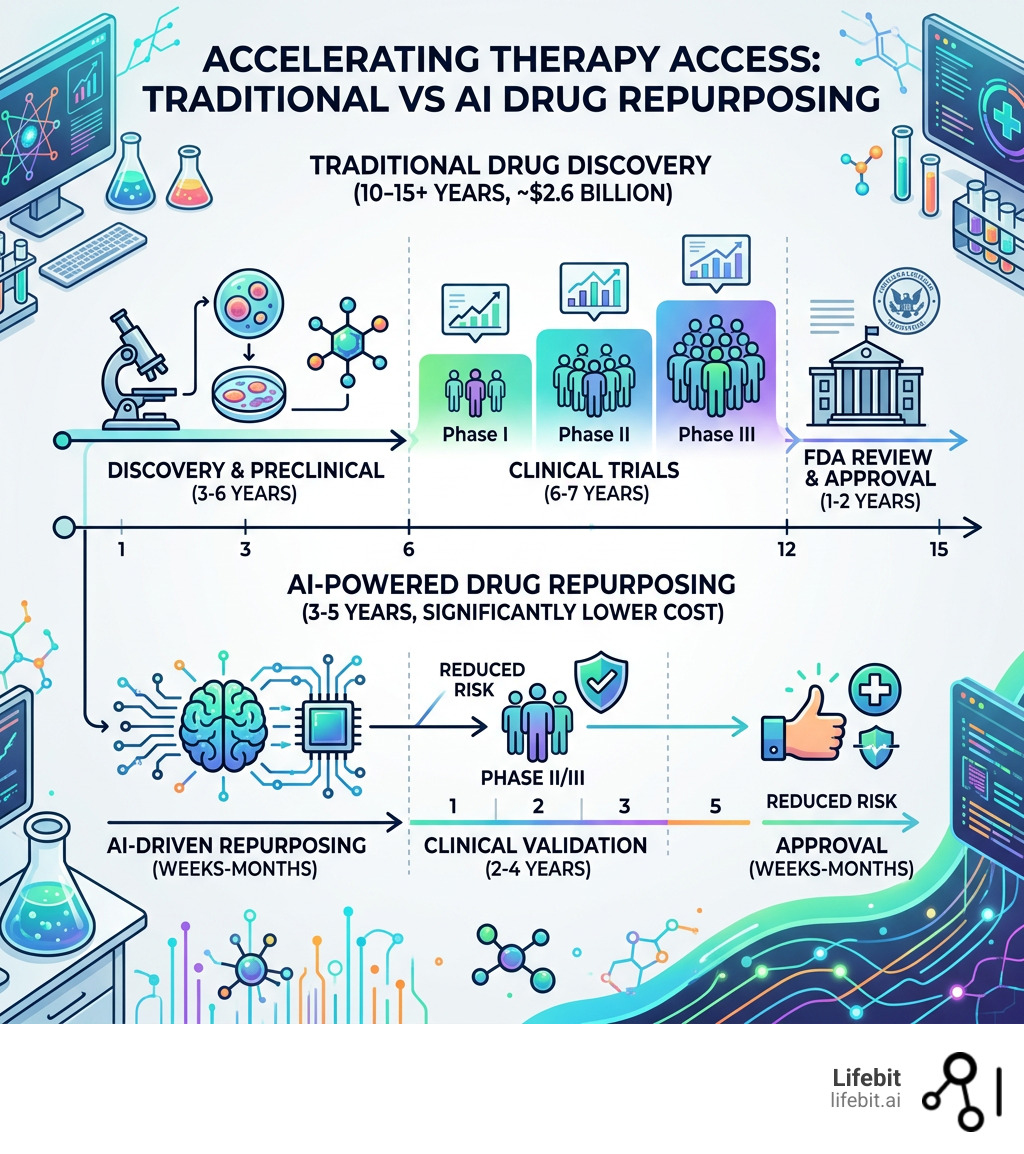
This is where drug repurposing AI steps in as a game-changer. By leveraging medicines that have already passed rigorous safety tests, we can bypass the 9-12 years and $1 billion price tag typically required for de novo drug discovery. For neglected conditions, this isn’t just a “faster” route; it’s often the only economically viable route. Because these drugs have already been through Phase I safety trials, the risk of failure due to toxicity is significantly lower, allowing researchers to focus entirely on efficacy.
A scientific review of AI in drug design highlights that AI excels at pattern recognition within “sparse data” environments. While we may not have decades of clinical trials for an ultra-rare genetic disorder, AI can look at the molecular signature of that disease and match it against the known mechanisms of thousands of existing compounds. This ability to find hidden connections across the human interactome—the complex network of all molecular interactions in a cell—is what allows us to turn the tide on the staggering human and economic toll of untreated diseases.
Overcoming the Data Gap with Drug Repurposing AI
The biggest hurdle in rare disease research is the “data gap.” Many conditions are so poorly characterized that we lack a clear map of which proteins or genes to target. To solve this, researchers use Knowledge Graphs (KGs)—massive digital webs that connect drugs, genes, diseases, and biological pathways. These graphs allow AI to perform “guilt-by-association” analysis: if a rare disease shares a biological pathway with a well-understood common disease, the AI can suggest drugs that work for the common disease as potential candidates for the rare one.
One of the most significant breakthroughs in this area is TxGNN. To build this model, researchers had to Access the Harvard TxGNN Knowledge Graph, which is a heterogeneous dataset containing:
- 123,527 nodes (representing drugs, diseases, proteins, and biological processes)
- 8,063,026 edges (representing the relationships between them, such as “drug treats disease,” “protein inhibits enzyme,” or “gene associated with phenotype”)
- 29 types of undirected edges and 10 types of nodes.
By training on this gargantuan network, drug repurposing AI can identify drug candidates for 17,080 different diseases—including those that currently have zero available treatments. It essentially “borrows” knowledge from well-studied diseases to make educated predictions about rare ones. This method, known as transfer learning, is the cornerstone of modern computational pharmacology.
Using Drug Repurposing AI to Predict Contraindications
Finding a drug that might work is only half the battle; we also need to know if it will be harmful in a new context. Traditional repurposing often relies on trial and error or anecdotal evidence, which is inherently risky for patients. AI models like TxGNN have demonstrated a 35 percent accuracy boost in predicting drug contraindications compared to previous models. This is critical because a drug that is safe for treating hypertension might have dangerous side effects when used to treat a specific rare metabolic disorder.
This isn’t just theoretical. Researchers validated these AI predictions by analyzing 1.2 million patient records from the Mount Sinai Electronic Medical Record (EMR) system. By checking the AI’s suggestions against real-world outcomes in over a million patients, we can significantly mitigate risk. This level of clinical validation ensures that when a “new” use for an old drug is suggested, it comes with a data-backed safety profile, reducing the likelihood of late-stage trial failures and ensuring that patient safety remains the top priority throughout the repurposing lifecycle.
How to Use Knowledge Graphs for Large-Scale Indication Finding

To scale drug discovery, we have to move beyond looking at one gene at a time. We use heterogeneous biological networks—knowledge graphs where different “nodes” (like a drug, a protein, or a phenotype) are connected by “edges” (like a side effect, a genetic association, or a metabolic pathway). These graphs represent the “interactome,” a holistic view of human biology that accounts for the fact that no drug acts in isolation.
According to research on representation learning for indication finding, this process involves “embedding” these complex relationships into a high-dimensional mathematical space. Think of it like a 3D map where drugs and diseases that share biological “signatures” are placed close together. If a drug node is mathematically close to a disease node it wasn’t previously associated with, that distance represents a high-probability repurposing candidate.
Strategic repurposing requires a multi-layered approach:
- Data Harmonization: Combining 10+ open datasets (like ChEMBL for bioactive molecules, PubChem for chemical structures, and DrugBank for clinical info) with proprietary clinical data. This requires advanced NLP to ensure that different naming conventions for the same disease or protein are unified.
- Multi-omic Integration: Layering DNA (genomics), RNA (transcriptomics), and protein (proteomics) interaction data to see the full biological picture. This allows the AI to see how a drug might affect gene expression or protein folding, not just its primary target.
- Strategic Mapping: Using AI to identify which existing drugs in a company’s portfolio could be “recycled” for high-need areas like oncology or neurology. This is particularly valuable for “shelved” drugs—compounds that were proven safe in trials but failed to show efficacy for their original intended use.
- Semantic Reasoning: Beyond simple proximity, modern AI can perform reasoning across the graph. It can identify that “Drug A inhibits Protein B, which is a precursor to Enzyme C, which is overactive in Disease D,” providing a logical chain of evidence for a new indication.
Step-by-Step: How TxGNN Predicts New Uses for Existing Drugs
If you want to understand the “gold standard” for drug repurposing AI, you have to look at TxGNN. As detailed in the Nature Medicine study on TxGNN, the model operates through a sophisticated multi-stage process designed to handle the complexities of rare disease data:
- The Heterogeneous GNN Encoder: This module takes the massive knowledge graph and processes the relationships between different types of nodes. Unlike standard neural networks, a Graph Neural Network (GNN) uses “message passing,” where each node gathers information from its neighbors. It learns the “context” of a drug—not just what it targets, but how that target affects an entire neighborhood of proteins and pathways.
- The Disease Similarity Decoder: For rare diseases with very little data, TxGNN uses “disease signature vectors.” It looks at the one-hop neighborhood of a rare disease (its closest biological relatives) and uses a gating mechanism to “borrow” embeddings from similar, better-characterized diseases. For example, if a rare form of muscular dystrophy shares 80% of its protein interactions with a more common variant, the AI uses the common variant’s data to fill in the blanks.
- Pretraining and Fine-tuning: The model is first pretrained on all 8 million relationships in the KG to learn general biology—essentially learning the “language” of the human body. It is then fine-tuned specifically on drug-disease pairs. This two-step strategy prevents “catastrophic forgetting,” allowing the AI to keep its broad biological knowledge while specializing in finding treatments.
- Metric Learning: This technique ensures that the AI can accurately rank which drugs are most likely to work, even when the disease is “unseen” (meaning it wasn’t in the training data). It creates a ranking system that prioritizes drugs with the strongest biological rationale, rather than just those with the most existing literature.
Interpreting Results with Drug Repurposing AI
A major barrier to AI adoption in medicine is the “black box” problem—clinicians won’t prescribe a drug just because a computer said “trust me.” If an AI suggests using a common beta-blocker for a rare neurological condition, the doctor needs to know why. TxGNN addresses this through a module called GraphMask.
GraphMask is an explainable AI tool that identifies the specific “multi-hop” paths in the knowledge graph that led to a prediction. For example, it might show a clinician: “Drug A is predicted to treat Disease B because Drug A inhibits Protein X, which is known to interact with Gene Y, a key driver of Disease B.” This transparency is vital for:
- Building Clinician Trust: Doctors can see the biological rationale and compare it against their own medical expertise.
- Hypothesis Formation: Researchers can use these paths to design lab experiments. If the AI suggests a specific protein interaction is key, the lab can test that specific interaction in a petri dish.
- Visual Evidence: The Free access to the TxGNN tool allows users to see these pathways visualized, making the data actionable for multidisciplinary teams of bioinformaticians and clinicians.
- Regulatory Support: Providing a clear biological mechanism of action (MoA) is often a requirement for moving a repurposed drug into clinical trials. GraphMask provides this MoA automatically.
Validating AI Predictions with Real-World Evidence
Does it actually work? The data suggests a resounding “yes.” In comparative tests against traditional computational methods, TxGNN was nearly 50 percent better at identifying drug candidates than other leading AI models. This performance jump is largely due to its ability to handle “zero-shot” predictions—finding treatments for diseases that have no known drugs at all.
Real-world success stories include:
- Baricitinib for COVID-19: One of the most famous examples of drug repurposing AI occurred during the 2020 pandemic. Researchers used AI knowledge graphs to identify Baricitinib—originally a rheumatoid arthritis drug—as a potential treatment for the viral “cytokine storm.” The AI identified its unique ability to inhibit AAK1, a regulator of viral entry. This prediction was made in days, led to rapid clinical trials, and resulted in FDA Emergency Use Authorization, saving countless lives.
- Anti-IL-17A Drugs: In a study of the top 50 indications ranked by AI for these drugs, 60 percent were conditions that already had positive clinical trial results. Crucially, none of the top-ranked indications were from failed trials, demonstrating the AI’s ability to filter out “noise.”
- Cancer Repurposing: AI has identified 25 candidate drugs for chondrosarcoma (including everolimus and paclitaxel) and 78 candidates for melanoma. By analyzing the metabolic pathways of tumor cells, AI can find drugs that “starve” the cancer of the specific nutrients it needs to grow.
- Breast and Lung Cancers: AI models have successfully matched drugs like sildenafil (Viagra) to liver cancer pathways and verteporfin to lung cancer treatments, often by identifying secondary effects on cellular signaling that were previously unknown.
In pilot usability studies, 12 medical experts (including MDs and pharmacists) found that the path-based explanations provided by the AI significantly increased their confidence in the predictions compared to traditional “black box” scores. This human-in-the-loop validation is essential; the AI identifies the candidates, but the medical experts provide the final sanity check before moving to the clinic. This synergy between machine intelligence and human expertise is the future of precision medicine.
Frequently Asked Questions about Drug Repurposing AI
How much faster is AI than traditional drug repurposing?
Traditional repurposing, which often relies on serendipity (like the discovery that a blood pressure medication also treats hair loss), can take 6-10 years to reach the market. Drug repurposing AI can compress the discovery and preclinical phase into months. By using federated platforms like Lifebit, researchers can access the necessary multi-omic data in real-time, cutting the total time to treatment by an estimated 6 to 7 years.
Can AI find treatments for diseases with no existing data?
Yes. Through “transfer learning” and “disease similarity” modules, AI can analyze a disease with no known drugs by comparing its genetic and protein signatures to similar conditions. TxGNN, for instance, identified drug candidates for over 17,000 diseases, many of which were previously considered “untreatable” because they were too rare to have dedicated research teams.
What is the difference between Generative AI and Drug Repurposing AI?
While Generative AI (like AlphaFold or LLMs for protein design) focuses on creating new molecules from scratch, drug repurposing AI focuses on matching existing, safe molecules to new targets. Repurposing is generally faster and lower risk because the safety data, manufacturing processes, and supply chains for these drugs already exist.
Is AI-driven drug repurposing FDA-compliant?
AI itself is a tool for discovery, but the data used to power it must be handled with extreme care. Compliance frameworks like HIPAA, GDPR, and SOC 2 are essential. Our federated approach at Lifebit ensures that sensitive patient data stays secure and compliant within its original institution while still allowing AI models to learn from it. This “data-to-model” approach is the gold standard for privacy-preserving research.
Does AI replace the need for clinical trials?
No. AI identifies the most promising candidates and provides a biological rationale, but clinical trials are still necessary to prove efficacy in humans. However, AI can help optimize those trials by identifying the specific patient subgroups most likely to respond to the drug, thereby increasing the trial’s success rate and reducing the number of participants needed.
Conclusion: How to Start Repurposing Drugs with AI in 2025
The era of relying on “serendipity” or “luck” to find new uses for old drugs is over. As lead researcher Marinka Zitnik and her team at the Kempner Institute for AI Research have shown, we now have the computational power to map the entire landscape of human disease.
At Lifebit, we believe the future of medicine is federated. Our platform provides the secure, compliant infrastructure needed to run drug repurposing AI on a global scale. By bringing the AI to the data—rather than moving sensitive data around—we enable biopharma companies and public health agencies to collaborate securely.
Ready to accelerate your drug discovery pipeline? Learn more about Lifebit’s federated AI platform and how we can help you turn your data into life-saving treatments.
Lifebit provides a next-generation federated AI platform enabling secure, real-time access to global biomedical and multi-omic data. With built-in capabilities for harmonization, advanced AI/ML analytics, and federated governance, Lifebit powers large-scale, compliant research and pharmacovigilance across biopharma and government agencies.

