Beginner’s Guide to Top AI Platforms for Biomarker Discovery

Stop Data Silos: How Lifebit’s AI Platform Finds Multi-Omic Biomarkers 10x Faster
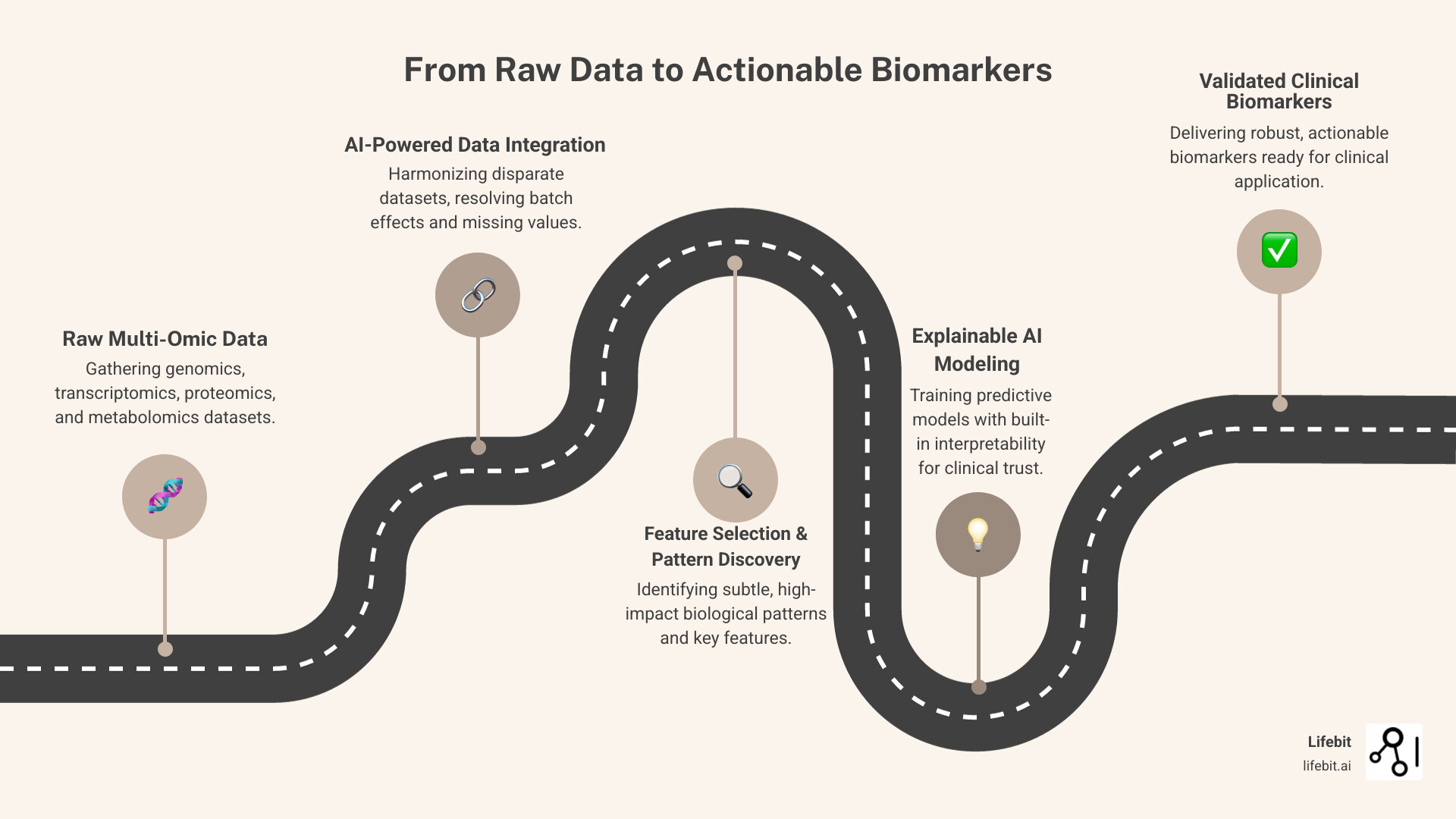
What are the top platforms using AI for biomarker findy from multi-omic data? If you’re racing to identify disease biomarkers from genomics, transcriptomics, proteomics, and other omics layers, you need platforms that integrate messy datasets, run AI models without a PhD in bioinformatics, and deliver results you can trust in the clinic. Here are the leading capabilities of the Lifebit platform:
Top AI Capabilities for Multi-Omic Biomarker Findy:
- Lifebit CloudOS – Federated, cloud-native platform for secure multi-omic analysis with explainable AI and real-time pharmacovigilance.
- Lifebit TargetID – AI-driven tool that connects genomic variants to drug targets using multi-omic evidence.
- Lifebit Cohort Browser – Interactive tool for rapid patient stratification and cohort building across diverse datasets.
- Lifebit Airlock – Ensures secure, anonymized data sharing that meets strict regulatory standards.
- Lifebit Trusted Research Environment (TRE) – A secure, scalable workspace for advanced AI/ML analytics on sensitive data.
Traditional biomarker findy is slow, expensive, and unreliable. High false-positive rates plague single-omics studies. Batch effects distort results. Missing data in metabolomics often exceeds 30%. And siloed datasets across genomics, proteomics, and clinical records make integration a nightmare.
AI changes the game. Machine learning models can integrate thousands of features from multiple omics layers, identify subtle patterns invisible to traditional statistics, and predict disease outcomes with unprecedented accuracy. But not all platforms are built the same. Some require coding expertise. Others lack explainability or regulatory compliance. And many can’t handle the messy, fragmented data that defines real-world biomedical research.
The right platform doesn’t just run models—it harmonizes data across omics types, handles missing values intelligently, and delivers interpretable results that clinicians and regulators can trust. It scales from pilot studies to global collaborations without moving sensitive data. And it empowers both bench scientists and computational biologists to extract actionable insights.
As Maria Chatzou Dunford, CEO and Co-founder of Lifebit with over 15 years in computational biology and AI-driven genomics, I’ve seen how the right infrastructure transforms multi-omic biomarker findy. In this guide, I’ll walk you through what are the top platforms using AI for biomarker findy from multi-omic data and how they solve the toughest challenges in precision medicine.
Simple What are the top platforms using AI for biomarker findy from multi-omic data? glossary:
Why Old-School Biomarker Discovery Wastes Time and Money (And How AI Solves It)
Let’s be honest: traditional biomarker discovery is a bit like looking for a needle in a haystack—while the haystack is on fire and someone keeps moving the needle. The “reproducibility crisis” is real. Many biomarkers that look promising in a small lab study fail miserably when tested on larger, more diverse patient populations. This failure rate is often attributed to the “p >> n” problem, or the curse of dimensionality, where the number of measured variables (genes, proteins, metabolites) vastly exceeds the number of patient samples, leading to models that overfit to noise rather than biological signal.
The primary reason for this failure is biological variability. A single gene or protein rarely tells the whole story. To understand complex diseases like cancer, autoimmune disorders, or neurodegenerative conditions like Alzheimer’s, we need to look at the rise of omics data integration, which combines DNA, RNA, protein levels, and metabolic signatures to see the “big picture.” Traditional statistical methods often struggle with the non-linear relationships inherent in these biological systems. For instance, a specific genetic variant might only increase disease risk if a certain protein is overexpressed and a specific metabolite is present in the gut microbiome.
Without AI, researchers are often stuck in data silos. One team has the genomics data, another has the clinical records, and a third has the proteomics. Trying to manually stitch these together leads to high false-positive rates and wasted research dollars. AI acts as a master translator, finding the hidden connections between these different data “languages” and identifying signatures that are actually robust enough for clinical use. Furthermore, AI can handle the “missing data” problem. In large-scale proteomics or metabolomics studies, it is common for 20-40% of data points to be missing due to detection limits. Advanced AI imputation techniques can predict these missing values with high accuracy, preserving the integrity of the dataset.
What Makes Lifebit a Top Platform for AI-Powered Multi-Omic Biomarker Discovery?
We built Lifebit to solve the “data deluge” problem. When people ask what are the top platforms using AI for biomarker discovery from multi-omic data?, we stand out because we don’t just provide tools; we provide a secure, federated environment. This means you can analyze data where it lives—whether that’s in a government biobank in the UK, a hospital in New York, or a private research facility in Singapore—without moving sensitive files. This “federated” approach is critical because moving petabytes of genomic data is not only expensive and slow but often legally impossible due to strict data residency laws like GDPR or HIPAA.
Our platform excels at:
- Data Harmonization: We take messy, inconsistent datasets from different sources and turn them into a unified, AI-ready format using standardized ontologies (like OMOP or SNOMED). This ensures that “Type 2 Diabetes” in one dataset is recognized as the same condition in another.
- Cross-Omics Consistency: We ensure that the signals you see in your transcriptomics data actually make biological sense when compared to your proteomics. Our AI models check for correlation across layers, filtering out technical artifacts that don’t align with biological reality.
- Horizontal and Vertical Integration: Whether you are looking at more patients (horizontal) to increase statistical power or more types of data per patient (vertical) to increase biological depth, our cloud-native architecture scales automatically to handle the compute load.

How Lifebit Handles Complex Multi-Omic Datasets with AI
Handling high-dimensional data is where the “magic” happens. We use advanced dimensionality reduction techniques like Principal Component Analysis (PCA), t-SNE, and UMAP to visualize complex relationships and identify patient clusters. But before we even get to the pretty charts, we have to deal with technical noise. Batch effects—systematic errors introduced when samples are processed on different days, by different technicians, or using different sequencing platforms—can ruin an entire study by creating false patterns.
Our AI-driven pipelines use latest research on AI for omics data analysis to perform non-linear batch correction. Unlike traditional linear methods, our AI models can identify and remove complex, non-linear distortions while preserving the underlying biological signal. This ensures that the “biomarker” you find is a real biological driver of disease, not just a technical glitch from a specific lab’s equipment.
The Engine Behind the Discovery: AI and ML Methodologies at Lifebit
What’s actually under the hood? We use a “nexus” of machine learning methodologies to ensure we catch every relevant signal. The complexity of multi-omic data requires more than just simple regression; it requires architectures that can model the hierarchical nature of biology, from the genome to the phenome.
- Supervised Learning: We train models on known outcomes (e.g., “this patient responded to treatment” vs. “this patient did not”). By using labeled datasets, we can identify the specific multi-omic features that predict drug efficacy or adverse reactions. This is the foundation of companion diagnostics.
- Unsupervised Learning: This is where we find the “unknown unknowns.” We use clustering algorithms to find entirely new subtypes of diseases. For example, what we currently call “asthma” might actually be five different diseases at the molecular level, each requiring a different treatment strategy. Unsupervised learning helps us define these molecular endotypes.
- Deep Learning: Using Convolutional Neural Networks (CNNs) for spatial transcriptomics and histopathology, and Transformers for genomic sequence analysis, we can process massive amounts of unstructured data. Transformers, in particular, are revolutionizing genomics by identifying long-range dependencies in the DNA sequence that traditional models miss.
- Reinforcement Learning: In drug discovery contexts, we use reinforcement learning to optimize lead compounds, virtually “testing” how different molecular structures might interact with a multi-omic biomarker profile to maximize binding affinity and minimize toxicity.
Key Algorithms for Multi-Omic Integration in Lifebit
We don’t believe in a “one size fits all” approach. Different diseases and data types require different tools. Our platform allows researchers to deploy a wide array of algorithms, often in ensemble configurations to improve robustness:
- Random Forest & XGBoost: These gradient-boosted decision trees are excellent for handling tabular data (like clinical records and SNP counts) and identifying feature importance. They tell us exactly which variables are the most predictive.
- Support Vector Machines (SVM): These remain highly effective for classification tasks in smaller, high-quality cohorts where the boundary between “healthy” and “diseased” needs to be precisely defined.
- Variational Autoencoders (VAEs): We use VAEs for data compression and denoising. By learning a “latent representation” of the multi-omic data, VAEs can find the core biological structures hidden beneath layers of technical noise and missing values.
- Graph Neural Networks (GNNs): Biology is a network of interactions. GNNs are perfect for modeling how proteins interact within a biological pathway or how genes regulate one another. By treating the cell as a graph, we can identify biomarkers that aren’t just single molecules, but entire dysfunctional sub-networks.
Lifebit’s platform provides a seamless interface to deploy these algorithms via pre-configured pipelines or custom scripts (Python, R, Nextflow). This ensures that researchers can move from data ingestion to model output without manual data movement, maintaining a full audit trail of every analytical step taken.
Why Lifebit Is the Go-To Platform for Multi-Omic Research
Researchers choose Lifebit because we bridge the gap between raw data and clinical insight. In the past, a multi-omic study required a massive team of bioinformaticians, data engineers, and cloud architects. We have automated the infrastructure layer so that scientists can focus on the science. Our interactive GUIs allow you to build custom AI/ML pipelines with a few clicks, while our robust API allows power users to integrate Lifebit into their existing enterprise workflows.
| Feature | Lifebit CloudOS | Traditional Tools |
|---|---|---|
| Data Access | Federated (Data stays put, code moves) | Centralized (Must move data to code) |
| Scalability | Cloud-native, serverless, unlimited | Limited by local server capacity |
| Compliance | HIPAA, GDPR, SOC 2, FedRAMP | Often manual, ad-hoc, and risky |
| Ease of Use | Interactive GUI, Low-code & API | Command line only, high barrier |
| Collaboration | Real-time shared workspaces | Emailing scripts and CSV files |
Lifebit’s AI-Driven Tools for Biomarker Discovery
We offer a suite of specialized tools designed to move you from raw data to a validated target as fast as possible. These tools are built to handle the specific nuances of multi-omic data, such as the different scales of measurement between genomics (discrete) and proteomics (continuous).
- Lifebit TargetID: This tool is a game-changer for drug discovery. It rapidly connects genetic variants (SNPs) to potential drug targets by integrating multi-omic evidence. It doesn’t just say a gene is associated with a disease; it uses AI to rank the “druggability” of that gene based on protein structure, pathway involvement, and existing clinical trial data.
- Lifebit Cohort Browser: Finding the right patients for a study used to take weeks of SQL queries. Our Cohort Browser allows you to “cut” cohorts visually. You can filter by genomic variants, clinical phenotypes, and even proteomic signatures in real-time, building complex patient groups in minutes.
- Lifebit Airlock: Security is the biggest hurdle in multi-omic research. Airlock ensures that only safe, anonymized results (like a summary statistic or a model coefficient) leave the secure environment. It prevents the export of raw sequence data, satisfying even the strictest data governors and ethics committees.
- Lifebit Trusted Research Environment (TRE): This is a secure, scalable workspace where advanced AI/ML analytics happen. It provides a “sandbox” where researchers can use high-performance computing (HPC) resources to train deep learning models on sensitive data without the risk of data leakage.
Case Study: Lifebit in Rare Disease Research. In a recent collaboration involving multiple international biobanks, Lifebit’s federated platform enabled researchers to aggregate data from over 50,000 rare disease patients. By applying AI across these diverse datasets without moving the data, the team identified three new genetic drivers for a specific pediatric condition. This process, which would have taken years using traditional data-sharing agreements and centralized storage, was completed in less than three months.
No More “Black Box”: Explainability and Regulatory Confidence with Lifebit
One of the biggest problems in clinical AI is the “black box” problem. If an AI tells a doctor, “this patient has a 90% risk of heart disease,” the doctor’s first question is “Why?” If the answer is “because the algorithm said so,” the biomarker will never be used in a clinic. Trust is the currency of medicine.
We use Explainable AI (XAI) techniques like SHAP (SHaplley Additive exPlanations) and LIME (Local Interpretable Model-agnostic Explanations). These tools “unmask” the AI, showing exactly which genes, proteins, or clinical factors contributed most to a specific prediction. For example, in a cancer recurrence model, SHAP might reveal that the prediction was driven by a combination of a specific mutation in the TP53 gene and an elevated level of a particular inflammatory cytokine. This isn’t just cool tech—it’s a regulatory requirement. To get a biomarker cleared by the FDA or integrated into clinical practice, you must show that the model is biologically grounded, reproducible, and trustworthy.
Ethical and Regulatory Strengths of Lifebit’s AI Biomarker Platform
Working with sensitive human data is a massive responsibility. Our platform is built on a foundation of privacy protection for clinical and genomic data. We go beyond simple encryption to ensure that patient privacy is maintained at every stage of the discovery process.
- Federated Governance: You control exactly who sees what. Access can be granted at a granular level, ensuring that a researcher only sees the specific data fields they need for their approved study.
- Synthetic Cohorts: We can use AI to generate “synthetic” data that mimics the statistical patterns of real biological data without containing any real patient information. This allows for collaborative research and model testing without ever exposing a real patient’s identity.
- Global Compliance: We meet the highest international standards, including HIPAA (US), GDPR (EU), and NIST 800-171. Our platform is designed to be “audit-ready,” providing a complete history of data access and analytical operations, which is essential for regulatory submissions to the FDA or EMA.
- Bias Mitigation: AI models can inherit biases present in the training data (e.g., if a dataset only contains samples from one ethnic group). Lifebit includes tools to detect and mitigate these biases, ensuring that the biomarkers discovered are effective for diverse global populations.
What’s Next: Functional Biomarkers and the Future of Precision Medicine
The future of biomarker discovery isn’t just about what is in the DNA, but how it functions in real-time. We are seeing a massive shift toward functional biomarkers and the study of Biosynthetic Gene Clusters (BGCs). While genomics tells us the blueprint, transcriptomics and proteomics tell us what is actually being built, and metabolomics tells us how the system is running.
By using AI to predict how genes will actually behave in a living system, we can move closer to the “Digital Twin” model. This involves creating a virtual version of a patient—modeled on their unique multi-omic profile—to test treatments in a digital environment before they ever receive a dose. This is particularly exciting in the field of immunotherapy, where AI is helping to predict who will respond to treatment with incredible precision. For instance, AI can analyze the interaction between a patient’s T-cells and the molecular markers on a tumor to predict the likelihood of a “cytokine storm” or other adverse events.
Furthermore, the integration of Real-World Evidence (RWE) from wearable devices and electronic health records (EHR) with multi-omic data is the next frontier. Imagine a biomarker that doesn’t just look at a blood draw, but also considers a patient’s heart rate variability, sleep patterns, and environmental exposures. Lifebit is at the forefront of this integration, providing the infrastructure to link longitudinal clinical data with deep molecular profiling. This holistic approach will allow us to move from reactive medicine (treating disease after it appears) to proactive medicine (preventing disease based on early molecular shifts).
Frequently Asked Questions about Lifebit’s AI Biomarker Platform
What are the main challenges in multi-omics data integration?
The biggest headaches are data heterogeneity (different formats and scales), high dimensionality (the “p >> n” problem), and technical noise like batch effects. In metabolomics, missing values often exceed 30%, which can lead to biased results if not handled by sophisticated AI imputation. Additionally, the lack of standardized ontologies across different “omics” layers makes it difficult to map relationships between a gene, its corresponding protein, and the resulting metabolite.
How does explainable AI (XAI) in Lifebit improve clinical trust?
XAI provides transparency by showing the “why” behind a prediction. By prioritizing features that are biologically relevant, we reduce skepticism from clinicians. If an AI identifies a biomarker that aligns with known biological pathways, it is much more likely to be adopted. XAI also helps in identifying “shortcut learning,” where an AI might be making a correct prediction based on a technical artifact (like the time of day a sample was taken) rather than actual biology.
Can Lifebit’s AI platform predict disease outcomes with high accuracy?
Yes. Platforms using these methodologies have achieved staggering results, such as high AUC (Area Under the Curve) scores for cardiovascular disease prediction and exceptional sensitivity for early-stage lung cancer detection. In rare diseases, our models have reached high accuracy in survival prediction and treatment response by integrating longitudinal clinical data with genomic profiles. However, accuracy is always dependent on the quality and diversity of the training data.
Does Lifebit support single-cell multi-omics?
Absolutely. Single-cell analysis is critical for understanding cellular heterogeneity, especially in oncology and immunology. Lifebit provides specialized pipelines for single-cell RNA-seq (scRNA-seq) and single-cell ATAC-seq, allowing researchers to identify biomarkers at the individual cell level. This level of granularity is essential for understanding how specific cell populations within a tumor might be driving drug resistance.
How does the federated model handle data security?
In a federated model, the data never leaves its original secure location. Instead of the researcher pulling the data to their local machine, the Lifebit platform sends the analytical code to the data. The analysis is performed locally, and only the results (which are non-identifiable) are sent back to the researcher. This minimizes the attack surface and ensures that sensitive genomic data is never exposed during transit.
Conclusion
The era of trial-and-error medicine is coming to an end. By leveraging the power of AI and multi-omics, we are finally able to see the full complexity of human biology. Whether you are a biopharma giant or a public health agency, the right platform can turn your “data chaos” into clinical clarity.
Ready to stop wasting time and start finding results?

