A Practical Guide to Driving Innovation in Omics Data Analysis

Cut Drug Findy Timelines by 30-50%: A Guide to Omics Innovation
Driving innovation in omics data analysis requires a fundamental shift away from traditional, siloed methods. Organizations that master three key pillars consistently achieve superior disease subtyping, find more robust biomarkers, and accelerate drug findy timelines by 30-50%.
The path forward involves:
- Integrating Multi-Omics Data: Moving from isolated genomic studies to integrated investigations of genomics, proteomics, and metabolomics. Research shows this delivers higher accuracy and better disease classification.
- Democratizing Access with AI: Empowering clinicians and domain experts to query complex datasets using natural language, removing the bioinformatics bottleneck.
- Building Secure, Collaborative Environments: Using federated platforms and Trusted Research Environments (TREs) to enable teamwork across institutions without compromising data privacy.
While high-throughput technologies generate massive omics datasets, most organizations struggle to extract actionable insights. Biopharma and public sector organizations face fragmented data, overwhelmed bioinformatics teams, and a gap between data generation and findy. This paradox—more data, slower translation—can only be solved by rethinking our approach.
I’m Maria Chatzou Dunford, CEO of Lifebit. With over 15 years in computational biology and AI, I’ve seen how combining federated data access with AI-powered analytics in secure environments is the key to open uping the full potential of precision medicine.
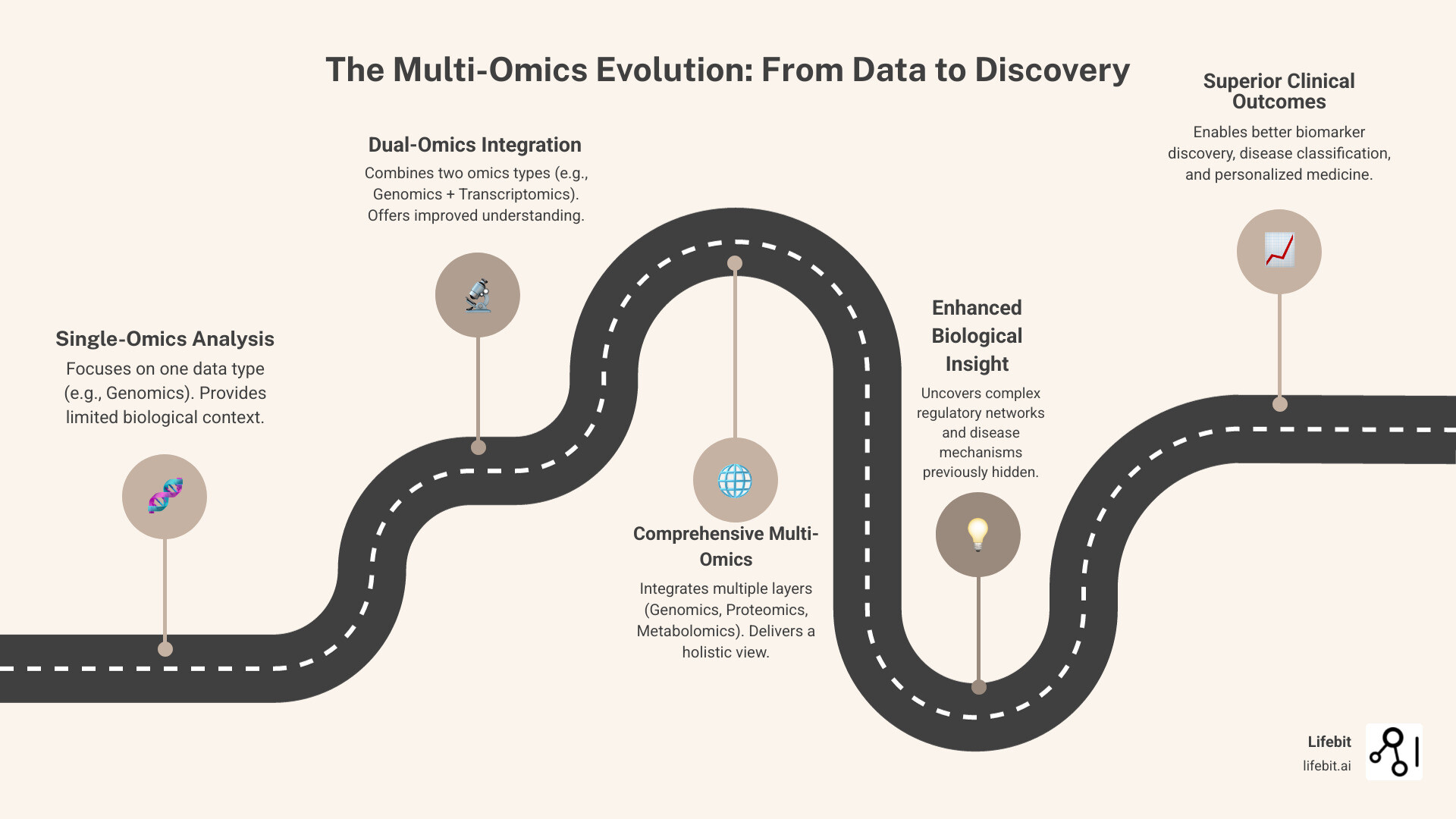
The Multi-Omics Imperative: Why Integrated Analysis Delivers Superior Insights
Analyzing genomics data in isolation is like trying to understand a car by only looking at the engine—you miss how all the interconnected parts work together. Systems biology teaches us that information flows through multiple layers simultaneously. Single-omics analysis cannot capture this complex web of interactions. How to Drive Innovation in Omics Data Analysis begins with embracing this principle.
Multi-omics integration combines data from different biological layers to create a holistic view that reflects how biology actually works. These layers include:
- Genomics: The complete set of DNA, representing the fundamental blueprint of an organism.
- Epigenomics: Modifications to DNA (like methylation) that don’t change the sequence but regulate which genes are turned on or off.
- Transcriptomics: The set of all RNA molecules, revealing which genes are actively being expressed at a given moment.
- Proteomics: The entire complement of proteins, representing the functional machinery that carries out cellular processes.
- Metabolomics: The collection of small molecules (metabolites) involved in cellular metabolism, reflecting the ultimate physiological state of the cell.
This integrated approach powers major initiatives like the All of Us Research Program and The Cancer Genome Atlas (TCGA), demonstrating that connecting the dots across omics layers transforms our understanding of disease.
Uncovering Deeper Molecular Mechanisms
By correlating changes across multiple omic layers, researchers can generate fundamentally better insights:
- Pinpoint robust biomarkers: A genetic variant (genomics) might only be pathogenic when it leads to altered gene expression (transcriptomics), abnormal protein function (proteomics), and a downstream metabolic shift (metabolomics). A multi-omics signature is far more reliable than a single-omics finding for early diagnosis, treatment response, and prognosis.
- Develop personalized treatment strategies: In oncology, for example, two patients with the same cancer type and even the same primary driver mutation may respond differently to therapy. Integrating proteomic and metabolomic data can reveal differences in signaling pathway activation or metabolic dependencies, allowing for therapies tailored to a patient’s complete molecular landscape.
- Improve patient outcome predictions: Models that incorporate clinical data with multi-omics profiles capture the complexity of disease progression with much higher accuracy, enabling proactive care and better clinical trial stratification.
- Uncover novel therapeutic targets: By revealing the complex molecular pathways underlying diseases, researchers can identify new intervention points. Techniques like spatial transcriptomics (which adds location data to gene expression) and liquid biopsies are already identifying therapeutic opportunities that were previously invisible.
The Statistical Proof of Superiority
The advantages of multi-omics are statistically proven. Research consistently shows that multi-omics classification models outperform single-omics models in accuracy and Area Under the Curve (AUC). For instance, a study on breast cancer subtyping might find that a model using only transcriptomic data achieves an AUC of 0.88 in predicting treatment response. By integrating genomic and proteomic data, the AUC could increase to 0.95. This is not a marginal gain; it represents a substantial reduction in patient misclassification, ensuring more patients receive the most effective treatment.
Furthermore, studies show that integrating three types of omics data produces significantly higher silhouette scores—a measure of clustering quality—than any single layer alone. This means multi-omics integration more accurately classifies cancer subtypes, creating distinct patient populations that can be matched with more effective therapies. The sum is truly greater than its parts, providing a more complete and actionable understanding of biology.
Navigating the Data Maze: Key Challenges Hindering Omics Innovation
The promise of omics data is extraordinary, but the reality is often frustrating. Biopharma and research institutions generate petabytes of data that remain scattered, siloed, and difficult to analyze. This isn’t just an inconvenience; it’s a fundamental barrier to how to drive innovation in omics data analysis.
The core problems are interconnected: massive data volume, the heterogeneity of different omics types, computational bottlenecks, and pervasive data silos. A lack of standardization in protocols and file formats further complicates efforts to combine datasets, while traditional workflows that funnel requests through small bioinformatics teams create crippling delays.
Technical Problems in Data Integration
Integrating omics data is like solving a puzzle with pieces from different boxes. Key technical problems include:
- The Curse of Dimensionality: This refers to the
p >> nproblem, where the number of features (p, e.g., 20,000 genes) vastly exceeds the number of samples (n, e.g., 200 patients). This high-dimensional space increases the risk of finding spurious correlations and building models that perform well on training data but fail to generalize to new patients. - Missing Data: No high-throughput platform is perfect. Mass spectrometry in proteomics may fail to detect low-abundance proteins, or low sequencing depth may create gaps in genomic data. This requires sophisticated imputation strategies (e.g., k-Nearest Neighbors, matrix factorization) to fill in the gaps without introducing bias.
- Collinearity: Biological variables are highly interconnected. The expression of one gene is often correlated with many others in the same pathway. This multicollinearity can make statistical models unstable and the interpretation of feature importance misleading.
- Batch Effects: These are systematic, non-biological variations introduced during data acquisition. Different sequencing machines, reagent lots, lab technicians, or even the time of day a sample is processed can create technical artifacts that obscure or mimic true biological signals. If not corrected using methods like ComBat, batch effects can lead to entirely false conclusions.
- Data Cleaning and Reduction: Before analysis can even begin, significant effort is required to normalize data, filter out low-quality reads, and align different data types. Research shows data reduction is the most frequent task in medical omics analysis, accounting for 55% of papers, highlighting the immense pre-processing burden.
The Clinico-Omics Conundrum
Integrating clinical data with omics data—clinico-omics—is essential for precision medicine but presents its own thorny challenges. Clinical data from electronic health records (EHRs) is notoriously complex. It exists in two forms: structured data (e.g., ICD-10 diagnosis codes, lab values, medication prescriptions) and unstructured data (e.g., physicians’ narrative notes, pathology reports, discharge summaries). The richest clinical context—nuances of disease progression, comorbidities, and lifestyle factors—is often locked away in this unstructured free text.
Unlocking this information requires advanced natural language processing (NLP). Techniques like Named Entity Recognition (NER) are used to automatically extract mentions of diseases, symptoms, drugs, and procedures. Relation Extraction then helps to link these entities, for example, by identifying that a specific drug was administered to treat a particular disease. This process is computationally intensive and requires careful validation.
Furthermore, combining identifiable health information with detailed molecular profiles raises significant data privacy concerns, demanding robust security and strict adherence to regulations like GDPR and HIPAA. Analyzing this data over time (longitudinal analysis) adds another layer of complexity, requiring methods that can manage evolving data structures, irregularly spaced clinical visits, and complex temporal correlations to truly understand a patient’s journey.
How to Drive Innovation in Omics Data Analysis with Advanced Technology
The challenges of omics data—silos, complexity, and dimensionality—are solvable with advanced technology. Artificial intelligence and machine learning are fundamental enablers that transform how we interact with biological data. The breakthrough happens when these sophisticated methods are combined with democratized access, putting powerful capabilities directly into the hands of domain experts.
Opening up Data for Everyone: The Role of AI and NLP
A common bottleneck in research is the gap between domain experts (clinicians, researchers) and the data. They have the critical biological questions but are blocked by technical barriers, forced to wait weeks for answers from overburdened bioinformatics teams.
Natural language processing and AI-powered interfaces eliminate this barrier. Imagine a clinician asking: \”Show me the metabolomic profiles for patients who responded poorly to this treatment, stratified by their BRCA mutation status.\” No coding, no tickets—just a plain-language question that triggers a sophisticated analysis. This is what empowering non-technical stakeholders looks like. GenAI-powered data assistants translate natural language into complex pipelines, accelerating insight generation and allowing researchers to test hypotheses in real time.
Mastering Multi-Omics Integration: A Breakdown of Key Approaches
Integrating multiple omic layers requires different analytical strategies depending on the biological question. Understanding these approaches is central to how to drive innovation in omics data analysis.
- Statistical Methods: Approaches like correlation analysis and Weighted Gene Co-expression Network Analysis (WGCNA) are simple and interpretable, excellent for identifying modules of co-expressed genes and relating them to clinical traits.
- Multivariate Methods: Techniques like Principal Component Analysis (PCA), Partial Least Squares Discriminant Analysis (PLS-DA), and Multi-Omics Factor Analysis (MOFA+) reduce dimensionality. They are valuable for identifying shared axes of variation across multiple omic datasets, helping to separate strong biological signals from technical noise.
- Machine Learning and AI: Algorithms like Random Forests and Gradient Boosting excel at capturing complex, non-linear relationships. Random Forests build hundreds of decision trees on different subsets of the data and average their outputs, making them robust to overfitting. Gradient Boosting builds models sequentially, where each new model corrects the errors of its predecessor, often yielding state-of-the-art predictive performance. They are robust to missing data and deliver superior predictive power.
For more detailed guidance, the Integration strategies for machine learning analysis resource provides comprehensive technical approaches.
Pushing the Boundaries with Deep Learning
Deep learning represents the cutting edge of omics analysis, using neural networks to find patterns that traditional algorithms cannot.
- Autoencoders perform unsupervised dimensionality reduction. Their encoder-decoder architecture learns a compressed, low-dimensional representation (latent space) of the data that captures its most salient features.
- Convolutional Neural Networks (CNNs), originally for image analysis, use learnable filters to scan for spatial patterns. In genomics, they can identify regulatory motifs in DNA sequences; in spatial transcriptomics, they can find cellular neighborhood patterns that correlate with disease.
- Variational Autoencoders (VAEs) are generative models that learn the underlying probability distribution of the data. This allows them to generate new, synthetic data points, which is useful for augmenting small datasets and improving model robustness.
- Graph Neural Networks (GNNs) are designed to work with data structured as a network, such as protein-protein interaction networks or gene regulatory networks. GNNs can model how information propagates through these biological systems, enabling predictions about gene function or drug target effects that are impossible with other methods.
The Imperative of Interpretability: Explainable AI (XAI)
In a clinical context, a correct prediction is not enough; we need to understand why the model made it. A “black box” model that predicts cancer survival without explaining its reasoning will not be trusted by clinicians. Explainable AI (XAI) addresses this by providing insights into model behavior. Techniques like SHAP (SHapley Additive exPlanations) and LIME (Local Interpretable Model-agnostic Explanations) can identify which specific features—such as the expression of certain genes or the presence of a metabolite—were most influential in a particular prediction. XAI is crucial for validating biological findings, building trust with clinicians, and moving AI models from research to clinical practice.
The Collaborative Framework: Secure Environments and Ethical Governance
Sophisticated analysis is meaningless if data remains locked in silos. How to drive innovation in omics data analysis requires a framework for sharing, protecting, and governing sensitive biomedical data. The future of precision medicine is collaborative, but this creates a tension between the need to share data and the absolute requirement to protect patient privacy.
Federated platforms resolve this tension. Lifebit’s federated AI platform, for example, enables secure, real-time analysis of global biomedical data without the data ever leaving its secure location. This approach breaks down silos while maintaining the highest security standards, powering compliant research in regions like the UK, USA, Europe, and beyond.

Building a Fortress: The ‘Five Safes’ of Trusted Research Environments (TREs)
A Trusted Research Environment (TRE) is a highly controlled computing environment that balances data protection with cutting-edge analysis. Its governance is often guided by the ‘Five Safes’ framework:
- Safe People: Researchers must be vetted, trained, and authorized to access the data, ensuring they are trustworthy and understand their responsibilities.
- Safe Projects: All research projects must be reviewed and approved by an ethics committee to ensure they are in the public interest and scientifically sound.
- Safe Settings: The TRE itself provides the secure technical infrastructure. It prevents unauthorized access, blocks data downloads, and logs all activity to ensure a complete audit trail.
- Safe Data: Data is de-identified to the greatest extent possible before being placed in the TRE. This involves removing direct identifiers and pseudonymizing records.
- Safe Outputs: All results, such as tables or graphs, are automatically or manually checked for disclosure risk before they can be exported from the environment. This prevents the accidental release of potentially re-identifiable information.
This framework ensures that TREs foster collaboration while maintaining robust security and governance.
Fostering Team Science Through Collaboration
Breakthroughs in omics research happen when biologists, clinicians, and data scientists work together. Secure collaborative environments replace clunky, risky data-sharing practices like emailing files or shipping hard drives. By allowing researchers to access and analyze data from a common, secure platform, they can break down data silos and correlate findings across institutions in real-time. This collaborative velocity transforms an overwhelming data deluge into actionable insights and accelerates the translation of research into clinical applications.
Upholding Trust: Navigating Ethical and Privacy Considerations
Working with omics data comes with great responsibility. Upholding trust is paramount.
- Patient Consent and Privacy: Individuals must give informed consent, understanding how their data will be used. Many large-scale projects rely on \”broad consent,\” which allows data to be used for future, unspecified research, a model that requires strong governance and transparency to maintain public trust.
- Data Anonymization and Its Limits: Genomic data is inherently identifiable. Even after removing names and addresses, a genomic sequence can potentially be used to re-identify an individual. This makes true anonymization nearly impossible. Therefore, the focus must shift from perfect anonymization to robust, access-controlled environments like TREs where data use is strictly monitored.
- Ethical Oversight: All research must be subject to strict review by Institutional Review Boards (IRBs) or Research Ethics Committees (RECs) to ensure it is conducted responsibly and that the potential benefits outweigh the risks.
- Responsible AI and Algorithmic Fairness: AI models trained on data from one population group (e.g., individuals of European ancestry) may perform poorly and generate inequitable outcomes for other groups. It is an ethical imperative to use diverse datasets, audit models for bias, and ensure that the benefits of AI in medicine are accessible to all.
- Regulatory Compliance (GDPR & HIPAA): Regulations like GDPR in Europe and HIPAA in the US provide legal frameworks for data protection. Navigating them with omics data is complex. For example, GDPR’s \”right to be forgotten\” is difficult to implement for genetic data that is shared among family members. Similarly, HIPAA’s \”Safe Harbor\” de-identification method is often insufficient for genomic data, necessitating the use of an \”Expert Determination\” to certify that the risk of re-identification is very small.
From Lab to Insight: Selecting an End-to-End Omics Analysis Partner
The path from raw omics data to actionable insight shouldn’t be an obstacle course of vendor handoffs and data silos. Fragmented workflows slow down research. This is why end-to-end solutions, which unify data generation, analysis, and interpretation, have become essential for how to drive innovation in omics data analysis.
The Value of a Unified Solution
A unified framework eliminates the friction points of a typical omics workflow. Instead of transferring data between incompatible systems, streamlined workflows allow your team to focus on answering biological questions.
The time savings are dramatic. While traditional multi-omics projects can take 6-18 months, end-to-end solutions can compress this timeline to just 4-8 weeks. This shift fundamentally changes how quickly you can validate findings and advance drug candidates. A unified approach also ensures consistent data quality through standardized kits and validated pipelines, and provides integrated expertise from a single partner who understands the project from start to finish. This comprehensive model has been shown to accelerate drug findy workflows significantly.
Key Considerations for Choosing a Provider
When evaluating end-to-end partners, ask these critical questions:
- Scalability: Can the platform handle your future data volumes without breaking?
- Security: Does the provider have industry-standard certifications and a proven track record of protecting sensitive biomedical data?
- Scientific Support: Do they offer expert guidance from experimental design through data interpretation?
- Flexibility: Can the platform accommodate various omics data types and integrate with your existing workflows?
- Track Record: Can they provide concrete examples of how their solutions have accelerated drug findy projects?
At Lifebit, our federated AI platform is built to address these needs, delivering the speed, security, and scientific rigor that modern omics research demands through our integrated TRE, TDL, and R.E.A.L. components. The right partner doesn’t just provide tools—they become an extension of your research team.
Frequently Asked Questions about Omics Data Innovation
What is the single biggest benefit of multi-omics integration?
The biggest benefit is gaining a complete biological picture. Single-omics is like looking at a car’s blueprint; multi-omics shows you the running engine, the fuel being used, and how all the parts work together. This holistic view leads to more accurate disease subtyping and more robust biomarker findy. It connects the dots between a gene mutation, its effect on protein levels, and the resulting metabolic changes, turning isolated data points into actionable insights.
How does AI help researchers who are not data scientists?
AI, particularly through Natural Language Processing (NLP), democratizes data analysis. It breaks down the technical barrier between domain experts and complex data. A clinician or researcher can ask a question in plain English, like \”Show me patients with high expression of gene X,\” and an AI copilot runs the sophisticated analysis behind the scenes. This empowers non-coders to explore data, test hypotheses in real-time, and accelerate findy without waiting for a bioinformatician.
What is a Trusted Research Environment (TRE) and why is it important?
A Trusted Research Environment (TRE) is a highly secure digital workspace for analyzing sensitive biomedical data. Its importance lies in solving the core tension of modern research: the need for collaboration versus the need for privacy. A TRE allows researchers from different institutions to work together on the same dataset without the data ever leaving the secure environment. This enables large-scale, multi-institutional studies while enforcing strict governance and compliance with regulations like GDPR and HIPAA, protecting patient data and building public trust.
What is the difference between federated analysis and centralizing data in a cloud TRE?
Centralizing data means physically moving and copying datasets from various sources into a single cloud location, such as one TRE. This can be slow, expensive, and often prohibited by institutional or national data governance laws that forbid data from leaving its jurisdiction. Federated analysis is a different paradigm that brings the analysis to the data. The data remains in its original, secure location (which could be its own local TRE), and only the computational query is sent to the data. The analysis runs locally, and only the aggregated, non-sensitive results are returned to the researcher. This model respects data sovereignty, enhances security by minimizing data movement, and enables collaboration across legal and geographical boundaries where data cannot be centralized.
Conclusion: Your Blueprint for the Future of Omics Research
The future of biomedical research is here, and it moves beyond analyzing single omics layers in isolation. How to drive innovation in omics data analysis requires a new blueprint built on three pillars:
- Multi-Omics Integration: Accept the complexity of biology to achieve superior accuracy, better disease classification, and more robust biomarkers.
- AI-Powered Democratization: Use AI and natural language processing to empower clinicians and domain experts to explore data directly, removing bottlenecks and accelerating findy.
- Secure Collaboration: Implement Trusted Research Environments (TREs) to break down data silos and foster team science while maintaining the highest standards of privacy and compliance.
Organizations that master these pillars will lead the next decade of precision medicine, reducing project timelines from months to weeks. At Lifebit, our federated AI platform is built on this blueprint. Our integrated TRE, Trusted Data Lakehouse (TDL), and R.E.A.L. (Real-time Evidence & Analytics Layer) deliver the secure, real-time, and AI-driven capabilities needed for large-scale research. We are already powering compliant research and pharmacovigilance for leading biopharma, governments, and public health agencies across the globe.
The future is about extracting meaningful insights faster and more collaboratively than ever before to transform patient lives.
Discover how to accelerate your research with a next-generation federated AI platform

