How to Integrate Multi-omics Data Without Losing Your Mind

Why Multi-omics Integration Is Your Key to Deeper Biological Insights
If you’re looking to truly understand complex biological systems, multi omics integration methods are essential. They combine different types of biological data to give you a full picture.
- What are multi-omics integration methods? They are computational strategies that combine different types of biological data (like DNA, RNA, proteins, and metabolites) from the same or related samples.
- Why use them? To get a full, holistic view of biological processes, uncover hidden relationships between molecular layers, identify new biomarkers, and advance precision medicine.
- How they work: By linking diverse data types, these methods help researchers understand how genes, proteins, and other molecules interact to influence health and disease.
Imagine trying to solve a complex puzzle with only a few pieces. That’s often how it feels when researchers try to understand biology using just one type of data. This is where multi omics integration methods come in.
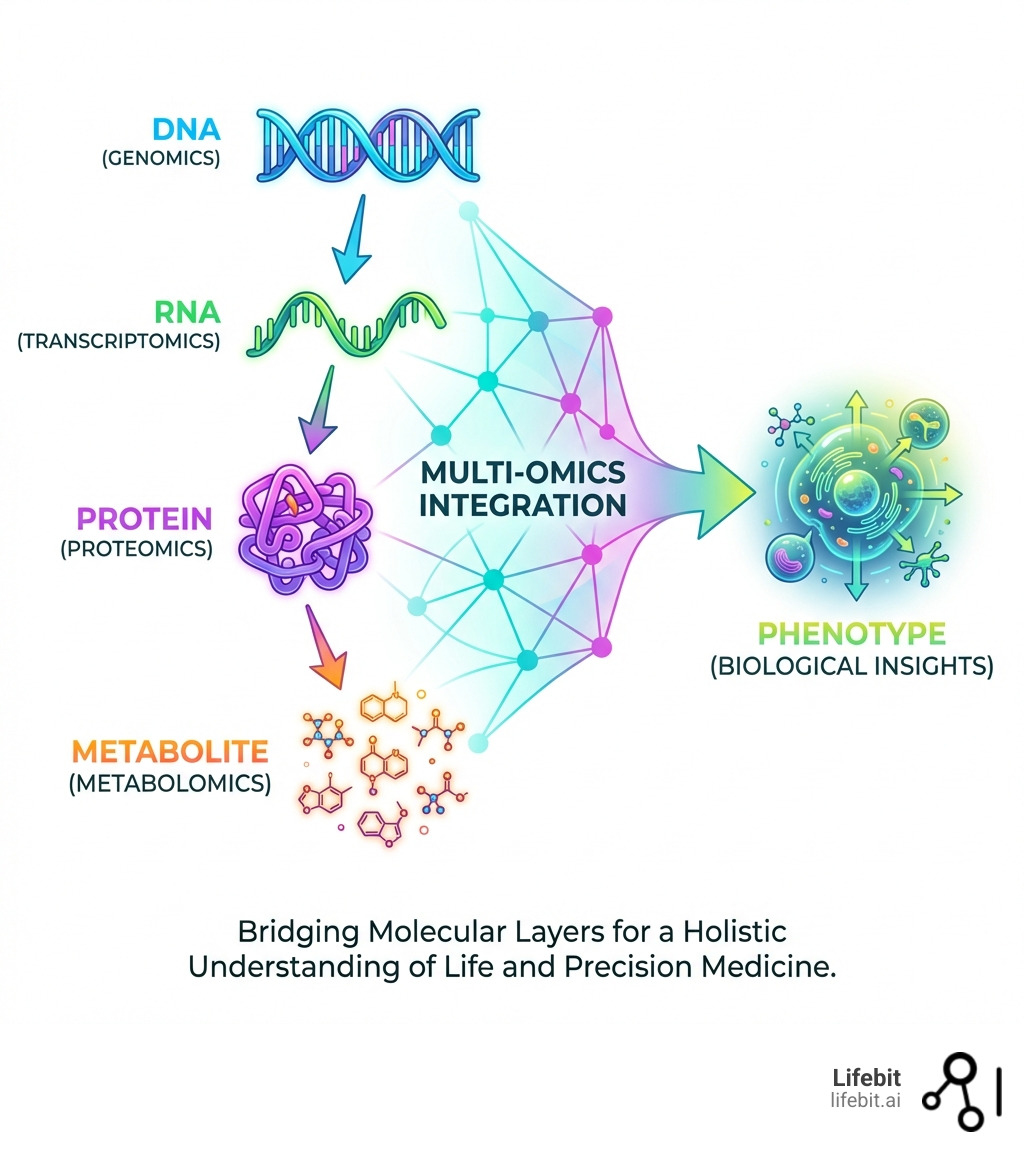
These powerful techniques bring together different ‘omics’ datasets. Think of genomics (DNA), transcriptomics (RNA), proteomics (proteins), and metabolomics (metabolites). Each one offers a unique angle. DNA is the blueprint, RNA shows active genes, proteins are the cellular workers, and metabolites are the chemical fingerprints.
By combining these insights, we can build a much clearer, more complete picture of biological systems. This integrated view is vital for understanding diseases, discovering new treatment targets, and developing truly personalized medicine.
My name is Maria Chatzou Dunford. My work in computational biology, AI, and high-performance computing has provided me with deep insights into multi omics integration methods, especially for biomedical data. I’ve seen how these techniques transform healthcare, empowering data-driven drug discovery through secure, federated platforms.
Why Multi-omics Integration Methods are the Backbone of Precision Medicine
In the old days (which, in biotech, was about ten minutes ago), we relied heavily on single-modality studies. We would look at a genome and try to guess what happened next. But biology doesn’t work in a vacuum. It’s a messy, beautiful conversation between different molecular layers. Multi-omics integration is the “universal translator” that allows us to listen to that conversation.
The Shift from Reductionism to Holism
Historically, biological research followed a reductionist approach, focusing on individual genes or proteins in isolation. However, the emergence of multi omics integration methods has catalyzed a shift toward systems biology. This holistic view recognizes that phenotype is not the result of a single linear pathway but an emergent property of complex, multi-layered networks. By integrating genomics, epigenomics, transcriptomics, proteomics, and metabolomics, we can map the flow of information from the static blueprint (DNA) to the dynamic functional state (metabolites).
Precision medicine demands this holistic view. When we use Integrating Multi-Modal Genomic and Multi-Omics Data for Precision Medicine, we aren’t just looking for a single mutation; we are looking for how that mutation ripples through the transcriptome and proteome. According to Scientific research on multi-omics approaches to disease, these methods are critical for:
- Biomarker Discovery: Identifying “molecular footprints” that predict disease or drug response. For instance, integrating metabolomics and transcriptomics helped identify sphingosine as a specific marker for prostate cancer. In cardiovascular research, combining lipidomics with genomic risk scores has significantly improved the prediction of major adverse cardiac events compared to traditional cholesterol tests.
- Disease Subtyping: Cancer isn’t one disease; it’s hundreds. Methods like iClusterPlus have identified 12 distinct clusters across cancer cell lines, revealing subgroups driven by shared genetic alterations rather than just the tissue of origin. This allows for “basket trials” where patients are treated based on their molecular profile rather than their tumor location.
- Pharmacogenomics: Understanding why a drug works for Patient A but causes a reaction in Patient B. Multi-omics allows us to see if a lack of response is due to a genetic variant (genomics), a silenced promoter (epigenomics), or a specific metabolic breakdown pathway (metabolomics).
By bridging the gap from genotype to phenotype, these methods provide the roadmap for AI for Precision Medicine.
Overcoming the Complexity of Diverse Datasets
If integrating data was easy, everyone would do it. The reality is that Omics data is inherently “noisy” and high-dimensional. You might have thousands of genes from scRNA-seq but only 100 proteins from a limited proteomic assay. This imbalance creates a massive computational headache known as the “p >> n” problem, where the number of features (p) vastly exceeds the number of samples (n).
One of the biggest hurdles is the “batch effect”—technical variations introduced when samples are processed at different times, by different technicians, or in different labs. Research on mitigating batch effects in large-scale studies highlights that without proper correction, your “breakthrough” might just be an artifact of the sequencing machine’s mood that day. Advanced algorithms like ComBat-seq or MNN (Mutual Nearest Neighbors) are now standard in the pipeline to ensure biological signal outweighs technical noise.
To handle this, we use rigorous Beyond Integration: Understanding Data Harmonization techniques. Common standards include:
- Feature Retention: Keeping features only if they are detected in more than 90% of samples to avoid zero-inflation bias.
- Cross-batch QC: For RNA, we might keep features present in 70% of batches, while for the more “fickle” metabolites, that threshold might drop to 30%.
- Normalization: Using methods like TMM (Trimmed Mean of M-values) or quantile normalization to make different datasets comparable.
The Impact of Multi-omics on Clinical Outcomes
The ultimate goal is better patient care. Research on multi-omics for cancer prognosis shows that integrated models consistently outperform single-omic models in predicting survival.
For example, in glioblastoma, combining DNA methylation and copy number alterations (CNAs) revealed synergistic effects on gene expression. Patients with specific hypomethylated markers had significantly different prognoses than those with the same markers amplified. This level of detail is only possible through robust multi omics integration methods.
We are also seeing the rise of “liquid biopsies,” which combine circulating tumor DNA (ctDNA), fragmentomics, and protein biomarkers for early cancer detection. This multi-modal approach increases sensitivity, allowing for the detection of stage I cancers that single-modality tests often miss. This is the future of Genomics in the clinic: faster, less invasive, and incredibly precise.
Categorizing Integration Strategies: Matched, Unmatched, and Mosaic
To choose the right tool, you first need to understand the “geometry” of your data. We generally categorize integration into four main strategies based on how the samples and features overlap:
| Strategy | Description | Best Use Case |
|---|---|---|
| Horizontal | Same omics type across different sample groups or batches. | Correcting batch effects in a large multi-center study. |
| Vertical | Different omics types measured on the same set of samples. | Linking DNA mutations to protein expression in a cohort. |
| Diagonal | Different omics types measured on different sets of samples. | Integrating scRNA-seq from one patient with scATAC-seq from another. |
| Mosaic | A mix of overlapping and non-overlapping modalities across samples. | Large consortia data where some samples lack certain measurements. |
The Three Paradigms of Integration: Early, Late, and Intermediate
Beyond the geometry, researchers must decide when the integration happens in the analytical pipeline:
- Early Integration (Concatenation-based): All data types are simply joined into one giant matrix before analysis. While simple, this often allows high-dimensional layers (like transcriptomics) to overwhelm smaller layers (like metabolomics).
- Late Integration (Ensemble-based): Each omic layer is analyzed separately to produce a model or prediction, and these results are then combined (e.g., through weighted averaging). This preserves the unique properties of each layer but may miss cross-layer correlations.
- Intermediate Integration (Joint Modeling): This is the current state-of-the-art. It uses mathematical frameworks to transform all data types into a shared “latent space” simultaneously. This allows the model to learn the relationships between layers while maintaining the integrity of the individual datasets.
Vertical Multi-omics Integration Methods for Matched Samples
Vertical integration is the “gold standard” because it uses matched samples. When you have the RNA and the protein from the exact same cell, the correlations you find are much more likely to be biologically meaningful.
Key multi omics integration methods for vertical data include:
- Similarity Network Fusion (SNF): This method creates a network of samples for each omic type and then “fuses” them into a single representative network. It’s excellent for identifying patient subtypes because it emphasizes the commonalities across layers while dampening noise.
- Multi-Omics Factor Analysis (MOFA+): MOFA+ is a statistical framework that disentangles the sources of variation in your data. It uses Bayesian group factor analysis to identify “factors” that explain variance across multiple omics layers. For example, Factor 1 might represent “Treatment Response” (shared across RNA and Protein), while Factor 2 might represent “Technical Noise” (specific to RNA). You can find More info about MOFA software on GitHub.
- iClusterBayes: A joint latent variable model that is particularly good at clustering samples when you have a mix of continuous (expression) and categorical (mutation) data. It uses a penalty-based approach to select the most informative features across all layers.
Solving the Unmatched Data Puzzle with Diagonal and Mosaic Integration
What if you don’t have matched samples? This is common in single-cell research, where the process of measuring the transcriptome often destroys the cell, making it impossible to measure the proteome of that exact same cell.
This is where diagonal integration comes in. These methods project different cells into a shared “co-embedded” latent space using anchor points. If two cells from different datasets end up in the same spot in this virtual space, we assume they are the same cell type. Algorithms like Seurat v4 use “Canonical Correlation Analysis” (CCA) to find these shared dimensions.
Mosaic integration takes this a step further. Research on mosaic integration and knowledge transfer explains how tools like MIDAS or UINMF leverage partial data overlap. If Sample A has RNA+Protein and Sample B has RNA+ATAC, we use the “RNA bridge” to link the Protein and ATAC data. It’s like a molecular game of “Six Degrees of Separation,” allowing us to infer relationships between modalities that were never measured together.
From Statistics to Deep Learning: The Computational Toolkit
The tools we use to perform these integrations have evolved from simple correlations to complex neural networks capable of modeling the non-linearities of life.

Classical Statistical and Machine Learning Approaches
Before AI was the talk of the town, we relied on solid multivariate statistics. These methods are still incredibly powerful because they are interpretable—you can actually see why the model made a decision, which is vital for clinical validation.
- Correlation-based methods: Sparse and Regularized Generalized Canonical Correlation Analysis (sGCCA/rGCCA) are the workhorses here. They look for the highest correlation between linear combinations of features from different omics layers. This is particularly useful for identifying “co-expression modules” that span across DNA and RNA.
- Network-based methods: WGCNA (Weighted Gene Co-expression Network Analysis) helps identify modules of highly correlated genes that might be involved in the same biological pathway. When extended to multi-omics, it can link these gene modules to specific metabolite clusters.
- Kernel-based methods: Multiple Kernel Learning (MKL) allows us to use different “kernels” to represent different data types. A kernel is essentially a similarity matrix; by combining kernels from genomics and proteomics, we can build more robust classifiers for disease state.
Most of these are implemented in R statistical packages, which remain the primary language for How to Drive Innovation in Omics Data Analysis.
The Rise of Deep Learning and Multi-omics Integration Methods
Since 2020, Deep Learning (DL) has taken over, specifically for large-scale single-cell datasets. Variational Autoencoders (VAEs) have become the dominant paradigm. According to Research on deep learning for multi-omics integration, VAEs are preferred because they can:
- Handle High Dimensionality: They compress thousands of features into a small, manageable “latent space” (bottleneck layer) that captures the essential biological signal.
- Impute Missing Data: If a sample is missing its proteomic data, a VAE can “guess” what it should look like based on the transcriptome by sampling from the learned distribution.
- Denoise: By forcing the data through a bottleneck, the model naturally filters out stochastic noise, leaving behind the robust biological patterns.
Graph Neural Networks (GNNs) and Transformers
The latest evolution involves Graph Neural Networks (GNNs). Unlike standard neural networks, GNNs can incorporate prior biological knowledge, such as protein-protein interaction (PPI) networks or metabolic pathways. By treating genes and proteins as “nodes” and their interactions as “edges,” GNNs can integrate multi-omics data in a way that respects the known architecture of the cell.
Furthermore, Transformer models—the technology behind ChatGPT—are being adapted for biology. These models use “attention mechanisms” to learn which genes are most relevant to one another across different omics layers, even if they are not directly connected in a known pathway. This allows for the discovery of entirely new regulatory mechanisms.
Advanced Training Strategies for High-Dimensional Data
To make these deep learning models work effectively without overfitting to the noise, we use some pretty clever training tricks.
- Adversarial Training: We use a “discriminator” to challenge the model. If the model can’t tell which batch or lab a piece of data came from, we know we’ve successfully removed batch effects. This is the core principle behind Generative Adversarial Networks (GANs) used in data harmonization.
- Disentanglement: This involves forcing the model to separate different biological factors—like separating “cell type” from “disease state” or “cell cycle phase” in the latent space. This makes the model’s output much more interpretable for biologists.
- Contrastive Learning: The model learns by comparing similar and dissimilar samples. By maximizing the similarity between matched multi-omics profiles from the same patient and minimizing it between different patients, the model learns the core features that define a specific biological state.
- Foundation Models: We are now seeing the birth of “biological LLMs.” Models like scGPT or Geneformer are pre-trained on millions of single-cell profiles across various tissues. This allows them to understand the “grammar” of biology—how genes typically interact—before they even see your specific, smaller dataset. This “transfer learning” approach is a game-changer for rare disease research where data is scarce.
Specialized Strategies for Spatial Multi-omics
The newest frontier is spatial omics—measuring where things are in a tissue slice. This adds a “where” to the “what.” Research on computational principles in single-cell integration suggests that spatial context is the key to understanding cell-cell communication.
In a tumor, the cells at the core behave differently than those at the invasive edge. Tools like SpatialGlue and MEFISTO are adapting single-cell methods to account for the physical distance between cells. This involves “deconvolution” algorithms that can take a spatial spot (which might contain 10-50 cells) and use single-cell reference data to figure out exactly which cell types are present and how they are interacting. This is crucial for understanding the “tumor microenvironment” and why some immune cells are “excluded” from attacking cancer cells.
Quality Control and Feature Selection Standards
You can have the most advanced VAE in the world, but if you put “garbage in,” you will get “garbage out.” Rigorous QC is non-negotiable.
We typically follow these standards:
- 90% Sample Retention: A feature must be present in nearly all samples to be considered reliable for downstream integration.
- Consistent Regulatory Directionality: If you’re looking at DEFs (Differentially Expressed Features), they should show the same direction of change (e.g., both up-regulated) across at least 70% of batches to be considered a robust biomarker.
- High-Confidence Relationships: Cross-omics relationships (like a gene and its corresponding protein) are only kept if they are observed consistently in over 70% of the data. This filters out spurious correlations that arise by chance in high-dimensional space.
To calculate these complex relationships, many researchers use the Hmisc package for correlations, which provides robust tools for handling large-scale data and calculating p-values that are adjusted for multiple testing (e.g., using the Benjamini-Hochberg procedure).
Frequently Asked Questions about Multi-omics Integration
What is the difference between horizontal and vertical integration?
Horizontal integration combines the same type of data (e.g., all RNA-seq) across different batches or studies to increase sample size. Vertical integration combines different types of data (e.g., RNA-seq + Proteomics) for the same set of samples to increase biological depth. Think of horizontal as “more of the same” and vertical as “more layers of detail.”
Why are Variational Autoencoders (VAEs) preferred for multi-omics tasks?
VAEs are highly flexible. They can handle the non-linear relationships common in biology, they are great at dimensionality reduction, and they have built-in mechanisms for handling missing data and noise. Since 2020, they have become the go-to for creating joint embeddings of multi-omics data because they provide a probabilistic framework rather than just a static projection.
How do you handle missing data in large-scale omics studies?
Missing data is inevitable, especially in proteomics and metabolomics. We handle it through “imputation” (predicting missing values based on available data using K-Nearest Neighbors or Deep Learning) or by using integration methods like MOFA+ or MultiVI that are designed to be “missing-data aware,” allowing them to use all available information without discarding samples.
What are the FAIR principles in multi-omics?
FAIR stands for Findable, Accessible, Interoperable, and Reusable. In multi-omics, this means using standardized file formats (like H5AD or Loom), providing clear metadata, and using ontologies to describe biological entities so that different datasets can be integrated seamlessly by researchers worldwide.
Conclusion
Integrating multi-omics data is no longer a “nice-to-have”—it is the foundation of modern biomedical research. Whether you are using classical statistics or the latest VAEs and GNNs, the goal remains the same: to turn fragmented, noisy data into actionable knowledge that can save lives.
As we move toward a future of “digital twins” and real-time health monitoring, the ability to synthesize information across the molecular spectrum will be the defining factor in medical breakthroughs. We are moving away from a world of “one-size-fits-all” medicine toward a future where every treatment is as unique as the patient’s own multi-omic profile.
At Lifebit, we understand that the biggest challenge isn’t just the math—it’s the access. Data is often siloed in different institutions or countries, making integration nearly impossible. Our Lifebit Federated Biomedical Data Platform solves this by bringing the analysis to the data.
Through our Federated Data Platform Ultimate Guide, we enable researchers to perform large-scale multi omics integration methods across global datasets without moving sensitive information. With a Trusted Research Environment (TRE) and a Biomedical Knowledge Graph, we provide the infrastructure needed to power the next generation of precision medicine.
If you’re ready to stop losing your mind and start finding insights, it’s time to look at Who Provides Advanced Analytics Solutions for Multi-Omic Data? and see how a Bioinformatics Platform can transform your research.
For more information on how we can help you secure and analyze your data, visit Lifebit.

