How Generative AI is Revolutionizing Bioinformatics Platforms

How a Bioinformatics AI Platform Cuts Genomic Analysis from Weeks to Minutes
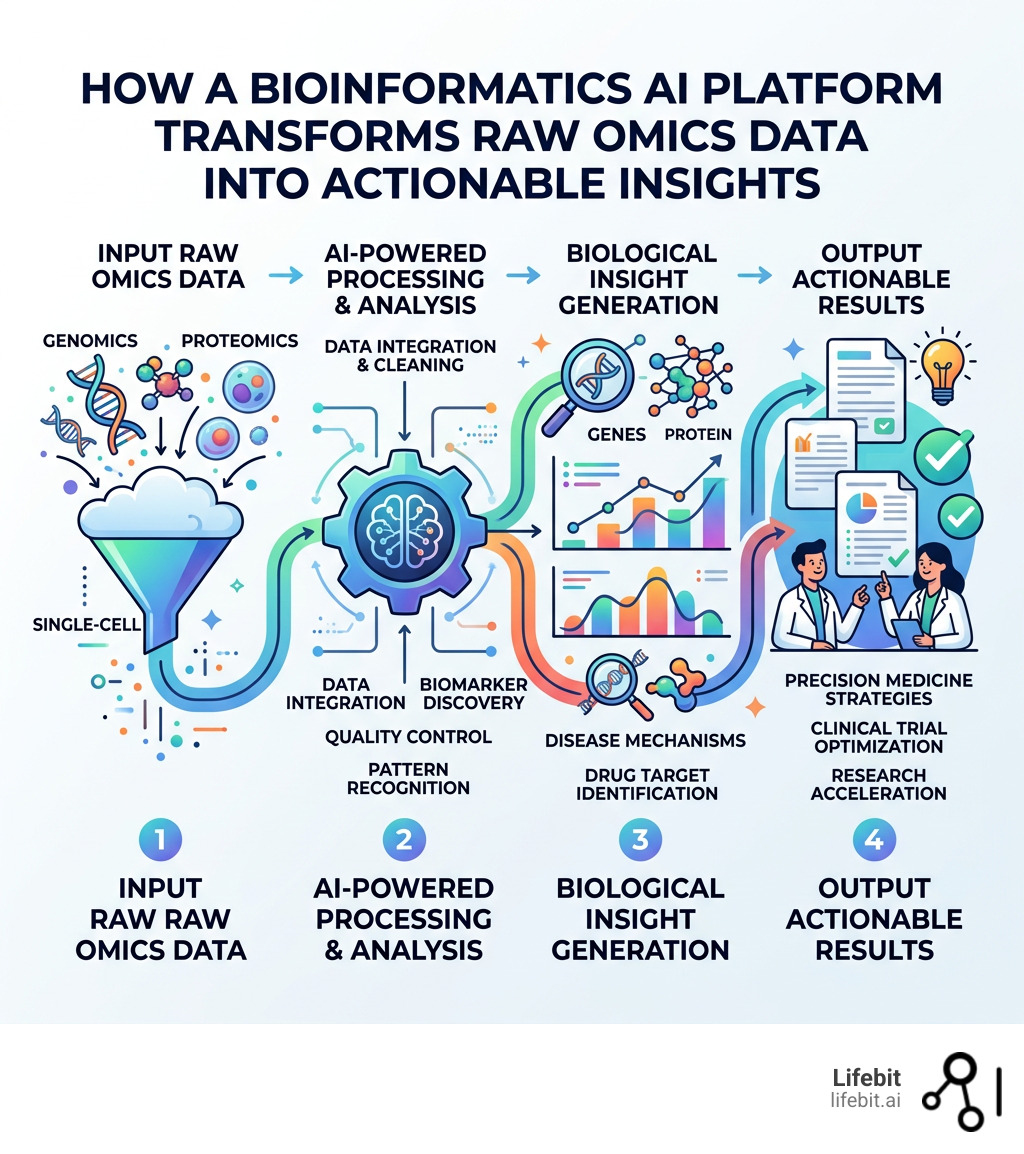
A Bioinformatics AI platform is software that combines artificial intelligence with biological data analysis — helping researchers process genomic, proteomic, and multi-omics data faster, cheaper, and more accurately than traditional methods. Leading platforms like Lifebit are transforming research through:
- Federated Data Access: Run analysis in situ across global datasets without moving sensitive data.
- Cost-Efficient Orchestration: Automate multi-omics pipelines with up to 81% reduction in compute costs.
- AI-Powered Insights: Use natural language to query complex cohorts and genomic data in seconds.
- End-to-End Security: Maintain full compliance with GDPR and HIPAA while scaling research.
- Multi-Omics Integration: Harmonize WGS, RNA-seq, and clinical data for a 360-degree patient view.
Not long ago, analyzing a genomic dataset meant days of manual coding, fragmented tools, and waiting on overloaded bioinformatics teams. Today, AI is compressing that timeline — from weeks down to minutes in many documented cases.
The shift is significant. Biological datasets are growing faster than any team can manually process. Whole-genome sequencing, single-cell RNA-seq, proteomics, and clinical EHR data are all converging — and traditional workflows simply weren’t built for this scale. Researchers are increasingly turning to AI-powered platforms to close that gap.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit and a key contributor to Nextflow — the global standard workflow framework used in genomic data analysis worldwide. With over 15 years building bioinformatics AI platform infrastructure across pharma, public sector, and precision medicine, I’ve seen how the right platform transforms what research teams can actually accomplish.
Bioinformatics AI platform further reading:
Why Traditional Workflows are Failing Modern Life Sciences

For years, bioinformatics data analysis felt like trying to assemble a 10,000-piece puzzle where half the pieces are missing and there’s no picture on the box. Traditional workflows are brittle, fragmented, and increasingly unable to keep pace with the “data deluge” of modern omics. The cost of sequencing a human genome has plummeted to under $200, yet the cost of analyzing that same data has remained stubbornly high, often exceeding the sequencing cost by a factor of ten.
We often see research teams hitting a wall because of these specific bottlenecks:
- Data Silos and Data Gravity: Biological data is often scattered across different clouds, on-premise servers, and geographic regions. Moving this data is slow, expensive, and often legally impossible due to privacy regulations. This is the concept of “Data Gravity”—as datasets grow into the petabyte scale, the cost of moving them (egress fees) and the time required for transfer become insurmountable barriers to collaboration.
- The “Mental Breakdown” Factor: A typical manual workflow involves over five distinct steps—finding papers, downloading data, QC, clustering, and reporting—often requiring 200+ lines of custom code. It’s no wonder researchers report “mental breakdowns” when a pipeline fails after 48 hours of runtime due to a minor configuration error or a version mismatch in a software dependency.
- Infrastructure Barriers and Technical Debt: Setting up research tools can take upwards of 10 hours of configuration. For a bench scientist without deep coding skills, this creates a dependency on IT teams that can delay insights by weeks or months. Furthermore, legacy systems often rely on “zombie scripts”—code written by post-docs who have long since left the lab, leaving no documentation behind.
- Redundant Tasks and Low Value Work: High-value scientists spend 80% of their time on data cleaning, format conversion, and pipeline debugging rather than actual discovery. This “data janitor” work stifles innovation and leads to high turnover in bioinformatics departments.
By moving toward a unified Bioinformatics AI platform, we can eliminate these friction points, transforming raw DNA files into refined insights with the push of a button. This transition allows organizations to shift from a reactive stance—struggling to manage data—to a proactive stance, where data becomes a strategic asset for drug discovery and precision medicine.
Core Capabilities of a Leading Bioinformatics AI Platform
What makes a platform “AI-powered” rather than just a standard cloud folder? It comes down to intelligence and integration. A modern bioinformatics platform acts as an intelligent layer that sits on top of your data, understanding the biological context of the files it processes. It doesn’t just store files; it understands that a .vcf file contains genetic variants and can automatically link those variants to known clinical phenotypes.
Key capabilities include:
- Multi-omics Integration: The ability to look at genomics, proteomics, and transcriptomics simultaneously to find correlations that single-modality tools miss. For example, an AI platform can correlate a specific SNP (genomics) with decreased protein expression (proteomics) and a specific clinical outcome (EHR data) in a single, unified query.
- Agentic Workflows: These are AI “agents” that don’t just follow a script; they can troubleshoot errors, optimize resource allocation, and even suggest the next logical step in an experiment. If a pipeline fails due to a memory error, an agentic platform can automatically restart the process with a larger instance type without human intervention.
- Knowledge Graphs: By connecting millions of data points—from gene variants to clinical outcomes—advanced platforms use AI-powered knowledge graphs to restructure data for reimagined insights. This allows researchers to perform “semantic searches,” asking questions like “Which genes in this cohort are associated with both inflammation and drug resistance?”
| Feature | Traditional Bioinformatics | AI-Powered Platform |
|---|---|---|
| Setup Time | 10+ Hours | < 5 Minutes |
| Interface | Command Line / Scripting | Natural Language / No-Code |
| Data Handling | Manual Downloads/Moves | Federated Access / In Situ |
| Error Handling | Manual Debugging | AI-Driven Troubleshooting |
| Analysis Speed | Days to Weeks | Minutes to Hours |
| Scalability | Limited by Local Hardware | Elastic Cloud Orchestration |
| Reproducibility | Low (Script-dependent) | High (Versioned Pipelines) |
For a deeper dive, check out our essential guide to ai platforms for biomedical data.
Automated Multi-Omics Integration in a Bioinformatics AI Platform
The “holy grail” of modern biology is understanding how different layers of biological information interact. This requires a bioinformatics AI platform that can ingest and harmonize diverse data types at scale. The challenge is that different “omics” data types use different formats, coordinate systems, and naming conventions. AI platforms use Large Language Models (LLMs) and specialized encoders to map these disparate data points into a common latent space.
Whether you are working with ai for genomics complete guide principles or performing advanced protein docking with AlphaFold, automation is key. Modern platforms provide day-0 access to pre-built pipelines for:
- Single-cell sequencing: Revealing cellular heterogeneity in cancer research by analyzing thousands of individual cells simultaneously to identify rare sub-populations that drive metastasis.
- Epigenomics: Decoding cytosine methylation patterns to understand how environmental factors influence gene expression, providing insights into aging and chronic diseases.
- Proteomics: Identifying protein dynamics and post-translational modifications for faster biomarker discovery, which is critical for developing targeted therapies.
By leveraging Nextflow documentation, platforms ensure these multi-omic workflows are reproducible and scalable across any cloud environment (AWS, GCP, Azure). This containerized approach means that a pipeline run in London will produce the exact same results as one run in New York, eliminating the “it works on my machine” problem.
Natural Language Processing and Agentic Bioinformatics AI Platform
Perhaps the most visible revolution is the death of the command line for everyday tasks. We are seeing the rise of “agentic bioinformatics,” where scientists can interact with their data using conversational AI. This democratizes access to complex data, allowing principal investigators and clinicians to explore data without waiting for a bioinformatician to write a script.
Imagine asking a platform: “Find papers about tumor-infiltrating lymphocytes, download the related scRNA-seq datasets, perform QC, and integrate them with our internal proteomics data.”
Advanced platforms use specialized agents—such as an “RNA Expert” or a “Literature Analysis Agent”—to execute these complex strings of commands. This best ai for genomics approach allows non-experts to perform high-level analysis without writing a single line of Python or R. These agents are trained on vast corpuses of biological literature, enabling them to provide context-aware suggestions, such as identifying potential off-target effects in a CRISPR experiment or suggesting a more appropriate normalization method for a specific dataset.
Quantifiable Benefits: Speed, Cost, and Scalability
The move to an AI-driven approach isn’t just about convenience; it’s about the bottom line and the speed of scientific discovery. In the competitive landscape of drug development, a three-month delay in clinical trials can cost a pharmaceutical company millions in potential revenue and, more importantly, delay life-saving treatments for patients.
The statistics from current users of a Bioinformatics AI platform are staggering:
- Cost Efficiency: Organizations have reported an 81% reduction in compute costs for RNA-Seq and a 40%+ reduction in cloud costs per run. This is achieved through intelligent resource orchestration, where the AI selects the most cost-effective cloud instances (such as Spot instances) and automatically scales them down the moment a task is completed.
- Speed: Time-to-results has been slashed by 24%, with some genomic surveillance capacities increasing by 10x. In public health crises, such as tracking viral mutations, this speed is the difference between a contained outbreak and a global pandemic.
- Accuracy: AI models like DeepVariant boast a 99.9% accuracy rate in sequence alignment and variant calling. Unlike traditional heuristic methods that rely on manual thresholds, AI models learn from millions of examples, allowing them to catch subtle patterns and rare variants that human-designed algorithms often miss.
- Throughput: Some teams have seen a 35x increase in sample throughput, allowing them to process thousands of pipelines weekly. This scalability is essential for population-scale genomics projects, such as the UK Biobank or All of Us Research Program, where researchers must analyze hundreds of thousands of genomes.
By following bioinformatics platform best practices 2025, labs can maximize their ROI. Collaborative setups allow for faster model development and easier adoption of machine learning among lab scientists, reducing the long-term maintenance costs of proprietary code. Furthermore, the ability to automate documentation and reporting ensures that all findings are audit-ready, which is a critical requirement for regulatory submissions to the FDA or EMA.
Solving the Compliance and Security Bottleneck
When dealing with 800,000+ patient genomic records or 3 million clinical diagnoses, security isn’t optional—it’s the foundation. A world-class Bioinformatics AI platform must address the “compliance bottleneck” that often stops research in its tracks. Traditional data sharing models, which involve copying and moving data, are increasingly viewed as too risky and legally complex.
We focus on several critical security layers to ensure data integrity and patient privacy:
- Data Residency and Sovereignty: Platforms must allow for “in situ” analysis, meaning the AI goes to the data, rather than the data moving to the AI. This is vital for complying with GDPR in Europe, HIPAA in the US, and national data sovereignty laws that strictly forbid sensitive health data from leaving its country of origin.
- Federated Governance and the “Five Safes”: Our platform enables secure, real-time access to global data via a Trusted Research Environment (TRE). We adhere to the “Five Safes” framework: Safe People (authorized researchers), Safe Projects (approved research goals), Safe Settings (secure environments), Safe Data (de-identified records), and Safe Outputs (non-disclosing results).
- Audit Readiness and Provenance: Every run must be 100% traceable. AI platforms now provide full audit trails and versioned analyses, ensuring that if a breakthrough is made, it can be defended during regulatory review. This includes tracking every software version, parameter setting, and data transformation used in the analysis.
- IP Ownership and Zero-Trust Architecture: Leading platforms ensure that users retain 100% ownership of any intellectual property produced. By utilizing a zero-trust architecture, the platform ensures that even the service provider cannot access the raw data or the proprietary algorithms without explicit, time-limited permission from the data owner.
For more on how to manage these requirements, read our saas platform for biomedical data guide. This approach not only protects patients but also builds the trust necessary for large-scale data sharing partnerships between academia, hospitals, and the pharmaceutical industry.
Frequently Asked Questions about AI in Bioinformatics
How do AI platforms handle sensitive patient data?
Modern platforms use federated learning and Trusted Research Environments (TREs). This means the data stays in its original, secure location (like a hospital or government database) while the AI performs analysis locally. Only the “insights” (the results) are shared, never the sensitive raw data. We also adhere to global standards like HIPAA, GDPR, SOC2 Type 2, and NIST 800-171.
Can non-experts use these platforms without coding?
Yes. The latest generation of Bioinformatics AI platform tools features natural language interfaces and no-code visual builders. This allows wet-lab scientists to run complex pipelines, like AlphaFold or variant calling, simply by asking questions in plain English or using drag-and-drop interfaces.
What types of omics data are supported?
Leading platforms are “multi-omic,” meaning they support Genomics (WGS, WES), Transcriptomics (RNA-Seq), Proteomics, Epigenomics (Methyl-Seq), and even Spatial Transcriptomics. They also integrate these with clinical EHR data and phenotypic records for a 360-degree view of patient health.
Conclusion: The Future of Biological Discovery
The era of the “lone bioinformatician” struggling with legacy scripts is ending. We are entering the age of the Bioinformatics AI platform, where the barrier between a biological question and a data-driven answer is thinner than ever.
At Lifebit, we believe the future of discovery lies in federated AI. By enabling secure, real-time access to global biomedical data through our Trusted Data Lakehouse and R.E.A.L. (Real-time Evidence & Analytics Layer), we are helping biopharma and public health agencies turn months of waiting into minutes of insight.
Whether it’s reducing query times for 28,000+ patient records from weeks to seconds or scaling genomic surveillance to meet a global crisis, AI is the engine. The question is no longer if you should adopt an AI platform, but how fast you can integrate one to stay at the forefront of science.
Ready to see the future of research? Explore the Lifebit platform and join the revolution in precision medicine.

