The Fast Track to Real Time Clinical Insights

Stop Wasting $2.5B: How Drug Discovery Insights Slash R&D Timelines by 50%
Drug discovery insights are the critical findings — from target identification to clinical validation — that determine whether a new medicine makes it to patients or fails in development.
Here is what you need to know:
| Stage | Key Insight Needed | Why It Matters |
|---|---|---|
| Target identification | Which protein or gene drives disease? | Defines the entire research direction |
| Hit discovery | Which compounds bind the target? | Narrows millions of candidates to hundreds |
| Lead optimization | Which lead is safe and effective? | Prevents costly late-stage failures |
| Clinical translation | Does it work in humans? | Final gate before FDA approval |
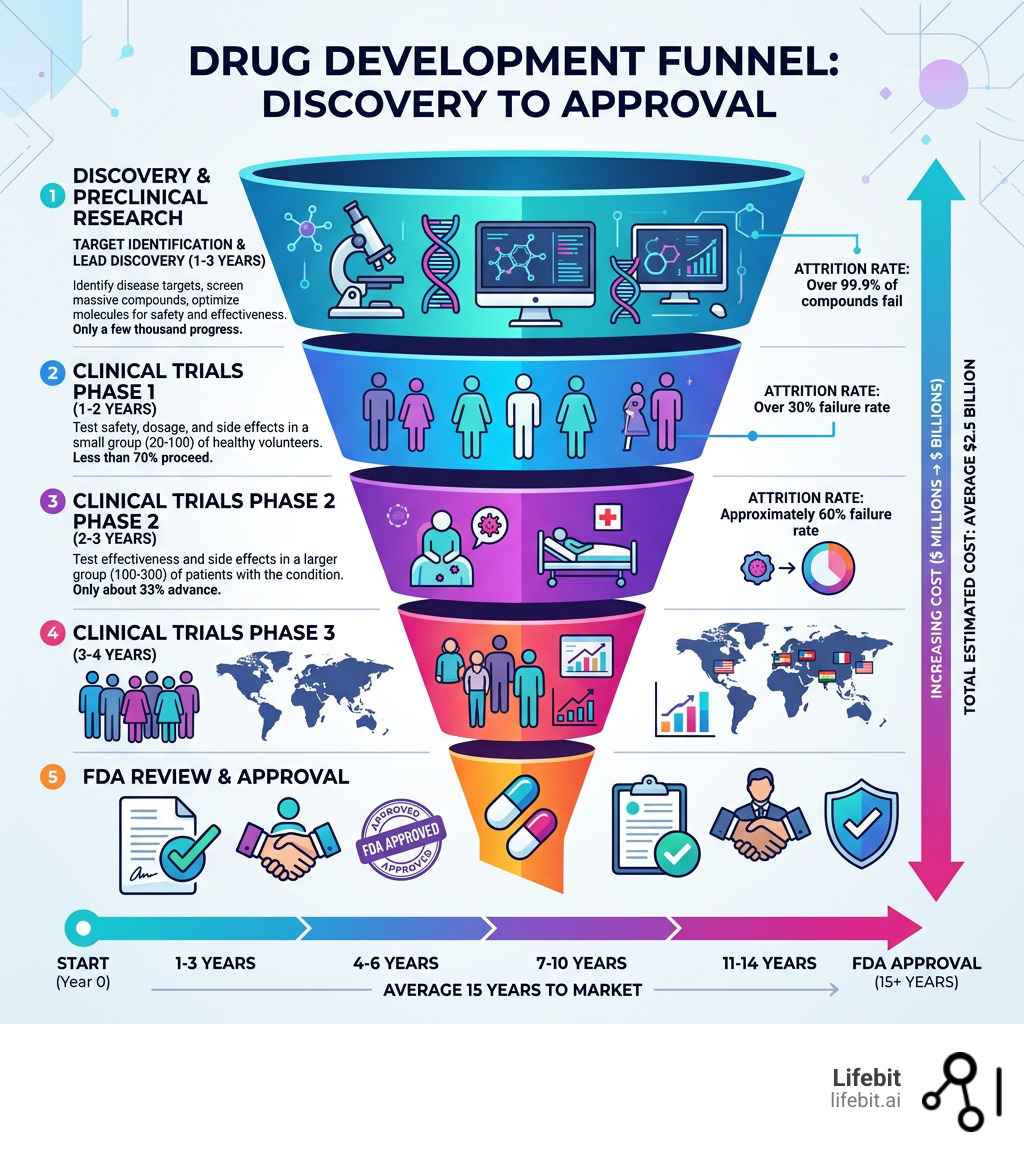
The numbers are stark. Bringing a single drug to market takes an average of 15 years and costs roughly $2.5 billion — and still, more than 90% of drug candidates fail before reaching patients.
That is not a science problem. It is an information problem.
Researchers are sitting on vast stores of genomic, clinical, and molecular data — but siloed systems, slow data pipelines, and regulatory barriers prevent them from acting on it in time. The labs that win are the ones that can turn raw data into actionable drug discovery insights faster than anyone else.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit, and with over 15 years in computational biology and federated data analytics, I have seen how the right data infrastructure transforms drug discovery insights from a bottleneck into a competitive advantage. In this expert roundup, I’ll bring together the most important advances reshaping how the industry finds, validates, and delivers new medicines.
Breaking the $2.5 Billion Barrier with Modern Drug Discovery Insights
We have all heard the “Eroom’s Law” joke—the observation that drug discovery is becoming slower and more expensive over time, despite improvements in technology. To break this cycle, we have to move away from trial-and-error and toward a data-first approach. The current paradigm is unsustainable; the pharmaceutical industry faces a “productivity gap” where R&D spending increases exponentially while the number of new molecular entities (NMEs) approved by the FDA remains relatively flat.
Traditional drug discovery often feels like looking for a needle in a haystack, while the haystack is on fire and the needle might not even exist. It starts with target identification, where we try to find a biological “switch” to flip. This involves understanding the complex signaling pathways of a disease. Then comes hit discovery, testing thousands of chemicals to see what sticks. By the time we reach lead optimization, we are refining those chemicals to ensure they don’t just work, but are safe, bioavailable, and metabolically stable. This phase is often where the “Rule of Five” (Lipinski’s Rule) comes into play, though modern AI is now helping us design molecules that break these traditional rules while remaining effective.
The shift toward drug discovery insights powered by AI is already changing the math. By using computational models for de novo design—creating new molecules from scratch rather than just screening existing libraries—we can explore a chemical space that is literally larger than the number of atoms in the known universe (estimated at 10^60 potential drug-like molecules). AI doesn’t just screen; it predicts properties like solubility, toxicity, and binding affinity before a single atom is synthesized in a lab.
| Feature | Traditional Development | AI-Driven Development |
|---|---|---|
| Average Cost | $2.5 Billion+ | Potential 30-50% reduction |
| Timeline | 12-15 Years | 5-7 Years (Targeted) |
| Success Rate | <10% | Improving via better target validation |
| Data Usage | Siloed, manual analysis | Integrated, real-time AI/ML |
According to scientific research on early drug discovery principles, the efficiency of the “hit-to-lead” phase is the biggest predictor of eventual clinical success. If we get the insights right early on, we don’t waste five years on a molecule that was never going to work. This “fail fast, fail cheap” philosophy is only possible when researchers have access to high-fidelity data early in the pipeline.
Accelerating Target Identification and De Novo Design
The “Holy Grail” of biology for the last 50 years was predicting how a protein folds just from its amino acid sequence. Enter AlphaFold. This technology has essentially indexed the shapes of nearly all known proteins, providing a massive shortcut for researchers. Before AlphaFold, determining a single protein structure could take a PhD student their entire degree using X-ray crystallography or Cryo-EM. Now, it takes seconds.
When we combine these structural insights with generative AI, we get a “search engine” for medicine. Instead of testing random compounds, we use AI-enabled molecular discovery to design molecules that fit perfectly into a target protein like a key in a lock. Advanced computational models and integrated omics platforms are now allowing us to identify disease-causing genes in weeks rather than years. This is particularly transformative for rare diseases, where the patient population is small and traditional research incentives are often lacking.
Leveraging Multi-Omics for Deeper Drug Discovery Insights
The human body doesn’t work in a vacuum. A single genetic mutation (genomics) might be influenced by which proteins are active (proteomics) or which metabolites are circulating (metabolomics). This is “multi-omics,” and it is the key to truly personalized medicine. By looking at the interaction between these layers, we can identify “biomarker signatures” that tell us exactly which patients will respond to a specific drug.
To make sense of this, we use tools like the Dimensions Knowledge Graph, which links millions of research papers, clinical trials, and patents. By harmonizing this data—making sure a “heart attack” in one database is coded the same way as “myocardial infarction” in another—we can run real-time analytics that reveal hidden patterns. At Lifebit, we see this as the “Real-time Evidence & Analytics Layer” (R.E.A.L.) that allows researchers to query global data ecosystems without the data ever having to move. This federated approach ensures that the insights are derived from the most diverse and comprehensive datasets available globally, reducing the risk of population-specific biases in drug efficacy.
From Lab to Clinic: AI Milestones and Real-World Success
We are no longer talking about “future potential”—AI-designed drugs are already in human arms. The transition from dry-lab predictions to wet-lab validation is the ultimate test of drug discovery insights.
A landmark moment occurred when the first AI-designed drug candidates for conditions like OCD entered Phase 1 trials in record time. Similarly, AI-driven discovery has identified novel targets and molecules for idiopathic pulmonary fibrosis (IPF), moving from concept to clinical trials in under 30 months. This is a staggering improvement over the traditional 5-to-7-year window for the same phase. These molecules weren’t just found; they were optimized for specific binding characteristics that traditional high-throughput screening (HTS) had missed for decades.

These successes prove that drug discovery insights generated by machine learning can significantly de-risk clinical trial design. One of the primary reasons drugs fail in Phase 2 or 3 is a lack of efficacy in the broader population. AI helps mitigate this by identifying the right patient subgroups through genomic screening. By selecting patients who are genetically predisposed to respond to a specific mechanism of action, we can ensure that trials are smaller, faster, and more likely to succeed. This is the essence of precision medicine: the right drug for the right patient at the right time.
Case Study: High-Throughput Screening at the NDDC
In Australia, the National Drug Discovery Centre (NDDC) at WEHI is a prime example of how high-tech infrastructure democratizes discovery. They provide researchers with access to ultra-high-throughput screening that was previously only available to “Big Pharma.” Their facility utilizes state-of-the-art robotics to screen hundreds of thousands of compounds against biological targets in a fraction of the time it would take a manual lab.
Through their subsidised screening programs, they have helped move projects forward that led to breakthroughs like Venetoclax, a life-changing cancer treatment that targets the BCL-2 protein. Venetoclax is a prime example of “structure-guided drug design,” where understanding the molecular interface allowed researchers to create a drug that triggers apoptosis (cell death) in cancer cells. You can see a video overview of NDDC screening to understand how they process hundreds of thousands of compounds with robotic precision. Their work shows that when you combine world-class robotics with smart data analysis, the “fast track” to insights becomes a reality for academic and biotech researchers alike, bridging the gap between basic research and clinical application.
Emerging Frontiers: CRISPR, PROTACs, and Next-Gen Therapies
The toolkit for treating disease is expanding beyond simple small-molecule pills. We are now entering the era of “living medicines,” precision editing, and targeted protein degradation. These modalities require a completely different set of drug discovery insights compared to traditional pharmacology.
CRISPR gene editing has moved from a lab curiosity to a clinical reality. We are now seeing personalized CRISPR therapy research that can fix genetic defects at the source. This isn’t just treating symptoms; it’s potentially curative. However, the challenge lies in “off-target effects”—ensuring the molecular scissors only cut where they are supposed to. AI models are now essential for predicting these off-target risks, allowing for safer therapeutic design.
Beyond gene editing, we are exploring the microbiome. The human gut contains trillions of bacteria that influence everything from immunity to mental health. The FDA has already approved the first orally administered fecal microbiota products to prevent recurrent infections, opening a whole new front in how we treat the human ecosystem. This represents a shift from “killing bugs” with antibiotics to “restoring balance” with microbial consortia.
In oncology, the focus has shifted to CAR-T cell therapies and PROTACs (Proteolysis Targeting Chimeras). While CAR-T has been successful in blood cancers, the new frontier is CAR-T for solid tumors, which requires even more sophisticated insights to overcome the “immunosuppressive microenvironment” of a tumor. PROTACs, on the other hand, are a revolutionary class of drugs that don’t just inhibit a protein; they hijack the cell’s own waste disposal system to completely degrade and remove the disease-causing protein from the body.
New Drug Discovery Insights for Complex Diseases
For decades, neurodegenerative diseases like Alzheimer’s and Parkinson’s were the “graveyard” of drug development. The failure rate was nearly 100% because we were intervening too late. That is changing thanks to biomarker discovery innovations. We can now detect signs of neuroinflammation and protein misfolding in the blood years before cognitive symptoms appear. This allows for “prevention trials,” where drugs are tested on at-risk individuals before irreversible brain damage occurs.
Similarly, in infectious diseases, we are moving away from “one bug, one drug” and toward broad-spectrum solutions. The PANVIPREP initiative is a great example, focusing on preparing for the next pandemic by identifying antiviral targets that work across entire families of viruses, such as Coronaviruses or Flaviviruses. By targeting conserved regions of the viral genome that don’t mutate easily, we can create “pandemic-proof” medicines.
Another emerging area is the use of “Digital Twins” in drug discovery. By creating a computational model of a patient’s biology, researchers can simulate how a drug will behave in a specific individual before they ever take a dose. This reduces the need for animal testing and allows for highly optimized dosing regimens, further pushing the boundaries of what is possible in clinical translation.
Overcoming Data Silos and Ethical Hurdles in AI Research
The biggest hurdle to better drug discovery insights isn’t the AI algorithms—it’s the data. Most medical data is locked away in “silos” within hospitals, national health services, or private pharmaceutical companies. Moving this data is risky, slow, and often legally impossible due to strict privacy laws like GDPR in Europe or HIPAA in the United States. This fragmentation means that AI models are often trained on limited, non-representative datasets.
Furthermore, if our data only comes from one population (for example, individuals of European descent), our AI will be biased. We need global, diverse data to ensure medicines work for everyone, regardless of their ethnic or geographic background. Without diversity in genomic data, we risk creating a new generation of “precision” medicines that only work for a fraction of the global population.
The solution we champion at Lifebit is federated learning. This is a paradigm shift in data science. Instead of moving the data to the researcher (the “centralized” model), we move the analysis to the data. In a federated system, the data stays safely where it was generated—in the hospital or the biobank. The AI model travels to each site, learns from the local data, and then sends only the “learned insights” (mathematical weights) back to a central hub. No patient-identifiable information ever leaves the secure environment.
By using Trusted Research Environments (TREs), we allow scientists to run complex AI models on sensitive genomic and clinical data while it stays safely behind the firewalls of the original institution. This secure collaboration is how we scale research without compromising patient trust. It allows for the creation of “virtual cohorts” of millions of patients, providing the statistical power needed to find the rarest genetic drivers of disease. This approach not only satisfies regulatory requirements but also encourages data sharing among competitors, as they can collaborate on insights without ever exposing their proprietary raw data. This “co-opetition” is essential for tackling the most complex challenges in human health, from cancer to rare genetic disorders.
Frequently Asked Questions about Drug Discovery
How much does it cost to develop a new drug?
The current industry average is approximately $2.5 billion. This includes the cost of all the failures that happen along the way. By using AI to improve “fail-fast” mechanisms, we aim to bring this number down significantly.
What is the role of AlphaFold in modern medicine?
AlphaFold provides the 3D structure of proteins. Knowing the shape of a protein is essential for designing drugs that can bind to it. It has essentially saved researchers millions of hours of manual lab work.
How does AI reduce clinical trial failure rates?
AI helps in two ways: first, by better predicting which drugs will be toxic before they ever reach humans, and second, by identifying the specific patients who are most likely to respond to a treatment based on their genetic profile.
Conclusion: The Future of Real-Time Biomedical Intelligence
The future of medicine isn’t just about better chemistry; it’s about better connectivity. We are moving toward a world of personalized medicine where drug discovery insights are generated in real-time from global, multi-omic datasets.
At Lifebit, we are proud to provide the federated AI platform that makes this possible. By enabling secure access to data across 5 continents—from London to Singapore—we are helping biopharma and governments turn data into cures. Whether it’s through our Trusted Data Lakehouse or our R.E.A.L. analytics layer, our goal is to ensure that the next life-saving insight is only a click away.
If you are a researcher looking to accelerate your pipeline, learn more about how to apply for NDDC screening or explore how federated AI can unlock your data’s potential. The fast track to clinical insights is already here—it’s time to start using it.

