Bridging Systems with Modern Data Interoperability Solutions

Why Healthcare Systems Are Racing to Connect Their Data
Data interoperability solutions enable different healthcare and research systems to exchange, interpret, and use information seamlessly—without costly custom integrations or data loss. Organizations implementing these solutions gain:
- Unified patient views across EHRs, labs, imaging, and genomics, allowing for a longitudinal record that tracks a patient’s journey from birth through specialized care.
- Faster time-to-insight through automated data harmonization, reducing the time researchers spend on data cleaning from months to minutes.
- Reduced integration costs by adopting open standards like FHIR and GA4GH Beacon, which eliminate the need for proprietary “black box” connectors.
- Improved care coordination via real-time clinical data exchange, ensuring that a specialist in one city has the same information as a primary care physician in another.
- Scalable research infrastructure supporting federated analysis without moving sensitive data, which is critical for complying with strict residency laws like GDPR.
The healthcare data landscape is fundamentally broken. Patients, quality of care, the healthcare ecosystem, research and development, and government policies are all reliant on the full availability of healthcare data across disparate systems—yet critical information remains trapped in siloed databases. A Bristol, Connecticut hospital discovered this when their previous data-sharing solution proved costly, failed to meet regulatory requirements, and lacked technical control over patient data protection. They found themselves unable to share basic imaging results with neighboring clinics without manual faxing or physical CD-ROMs, leading to delayed diagnoses and redundant testing.
Without interoperability, doctors can’t access MRI results without custom frameworks, researchers can’t discover genomic variants across institutions, and manufacturers can’t scale production with incompatible machines. This fragmentation creates a “data tax” on every clinical decision. For example, when a patient moves between health systems, the lack of interoperability often results in a 20% duplication rate for laboratory and imaging tests, as providers cannot verify previous results.
The cost of inaction is staggering. Traditional integration approaches lead to permanent technical debt, with major version upgrades costing $500,000–$5 million each and a 20-year data lifecycle totaling $5–25 million across 4–6 forced migrations. These costs stem from the need to constantly rewrite mapping logic every time a source system (like an EHR) updates its schema. Meanwhile, one of the main bottlenecks in human genomics research remains lack of accessible data, despite over 90 Beacons serving 200+ datasets and resources like Progenetix curating 113,322 genomic copy number profiles from 1,600 studies. The inability to query these datasets simultaneously prevents the discovery of rare disease markers that only appear in large, diverse cohorts.
Modern data interoperability solutions address these challenges through standardized protocols like HL7 FHIR for clinical data exchange, USCDI for nationwide data harmonization, TEFCA for trusted network agreements, and GA4GH Beacon for federated genomic discovery. These standards enable organizations to consolidate fragmented data pipelines, minimize transformation errors, simplify compliance, and promote scalability—all while reducing middleware costs and development expenses. By moving away from point-to-point integrations toward a “hub-and-spoke” or federated model, healthcare entities can ensure that their data remains an asset rather than a liability.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent over 15 years building data interoperability solutions that power secure, federated analysis across genomic and biomedical datasets for pharmaceutical organizations and public sector institutions. My background in computational biology, AI, and health-tech entrepreneurship—including contributions to Nextflow and research at the Centre for Genomic Regulation—has shown me how the right interoperability architecture unlocks precision medicine at scale. We have seen firsthand how breaking down these silos allows for the identification of novel drug targets in a fraction of the time previously required.
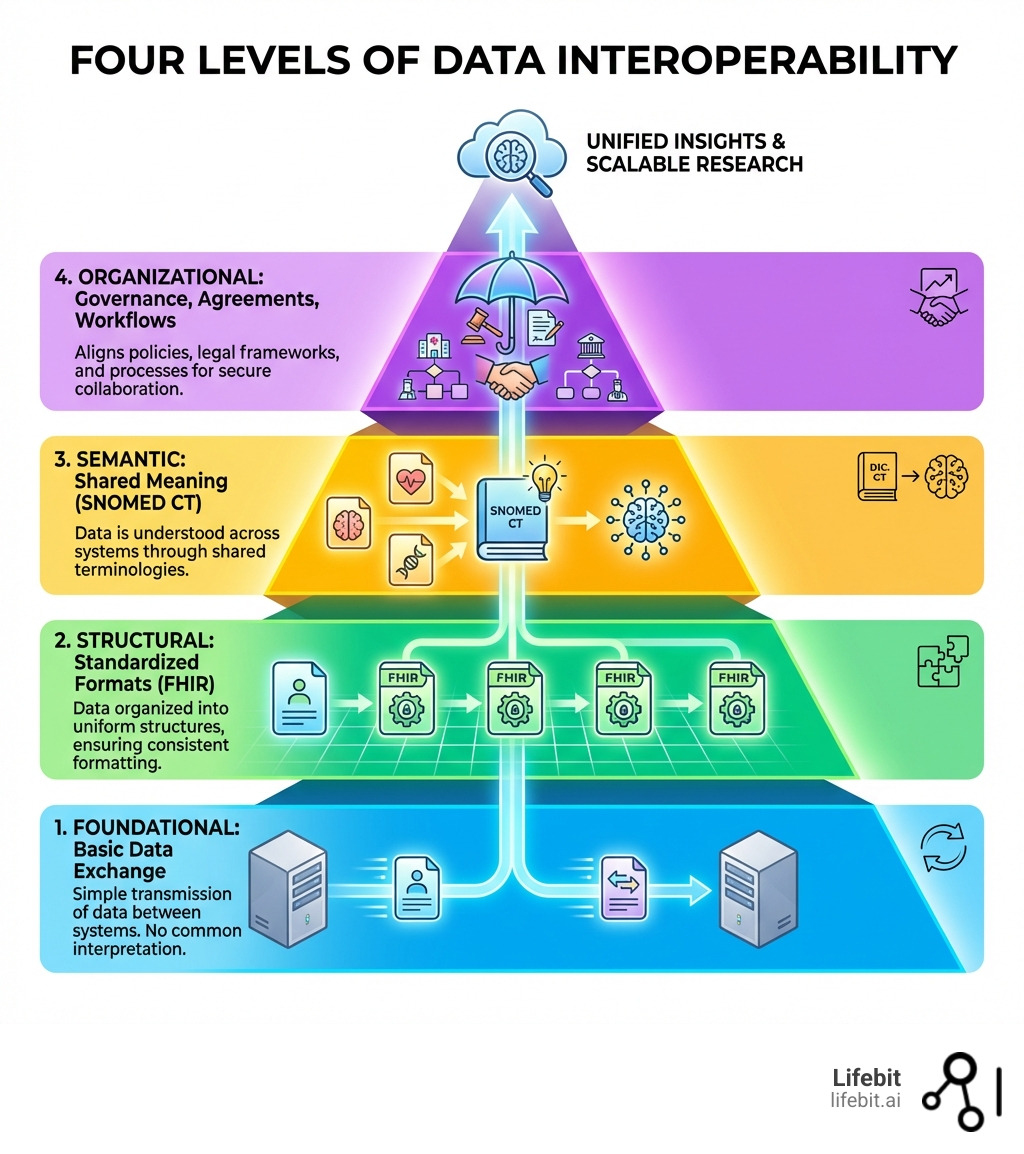
Data interoperability solutions terms to remember:
Why Healthcare and Genomics Need Data Interoperability Solutions Now
In the modern medical landscape, data is the lifeblood of discovery. However, About Public Health Data Interoperability remains a significant hurdle. For biopharma and public health agencies, health data interoperability is no longer a luxury—it is a requirement for survival. When systems “speak the same language,” we see a dramatic shift in organizational efficiency, moving from reactive data management to proactive clinical intelligence.

Interoperability allows diverse systems to develop an overlapping understanding of domain-specific data. This is essential for Data Exchange and Data System Interoperability, as it:
- Streamlines Data Management: It consolidates access into a single platform, ensuring accuracy with minimal transformation. Instead of maintaining fifty different data pipelines for fifty different hospitals, a single interoperable gateway can ingest data in a standardized format.
- Boosts Productivity: Real-time access removes redundant processing and manual data entry errors. Clinicians spend less time acting as “data clerks” and more time with patients, while researchers can automate the ingestion of new trial data.
- Promotes Scalability: Organizations can expand operations and adapt to market trends without rebuilding their entire IT stack. When a hospital acquires a new clinic, interoperability ensures the new facility’s data integrates into the main system in days, not years.
- Reduces Costs: By avoiding expensive middleware and custom “point-to-point” integrations, agencies can save millions in long-term maintenance. Point-to-point integrations are fragile; if one system changes its API, the entire connection breaks. Interoperable standards provide a buffer against this volatility.
For genomics, the stakes are even higher. Large-scale approaches are needed because of the vast diversity of genomic variations. A single human genome contains roughly 3 billion base pairs; multiplying this by thousands of patients creates a data volume that is impossible to manage without strict data interoperability solutions. Without these solutions, the “Internet of Genomics” remains a dream, and life-saving research is delayed by data silos that prevent scientists from comparing genetic markers across different ethnic and geographic populations.
Solving the Semantic Gap in Data Interoperability Solutions
The most difficult hurdle isn’t moving the data; it’s understanding it. This is where data harmonization meaning complete guide becomes vital. Semantic interoperability ensures that when two systems exchange a “blood pressure” reading, they both understand the units (mmHg), the context (systolic vs. diastolic), and the clinical significance. Without semantic standards, one system might record “High BP” while another records “140/90,” making automated analysis impossible.
Traditional ETL (Extract, Transform, Load) processes are often described as “ETL Hell” because they create permanent technical debt. Every time a vendor updates their software, the mappings break, requiring expensive consultants to fix the pipes. To solve this, we look toward data harmonization services bridging the gap between disparate datasets that utilize “permanence architecture.” By using CUID2 identifiers—which decouple structure from semantics—data remains stable over decades, even as underlying systems evolve. This ensures that a record created in 2024 remains readable and useful in 2044, regardless of what EHR software is in use.
The Role of Open-Source Infrastructure in Modern Systems
We believe the foundation of data trust must be open and community-driven. Open-source clinical data integration software offers several “punch-to-the-face” advantages over proprietary “black box” systems:
- Superior Cybersecurity: With more eyes on the code, vulnerabilities are found and patched faster. In healthcare, where ransomware is a constant threat, the transparency of open-source code is a security feature, not a bug.
- Easier Customization: Agencies can tweak the software to fit specific jurisdictional needs. A public health department in Singapore may have different reporting requirements than one in New York; open-source tools allow for this flexibility without waiting for a vendor’s product roadmap.
- Lower Total Cost of Ownership: No “vendor lock-in” means you aren’t held hostage by rising license fees. Organizations can switch service providers while keeping their core infrastructure intact.
- Better Interoperability: Tools like Match*Pro Software allow for probabilistic record linkage using open, validated models rather than secret algorithms. This allows for the accurate merging of patient records from different sources without compromising privacy.
Core Standards Driving Global Health Data Exchange
To achieve nationwide and global exchange, we rely on a “stack” of recognized standards. These aren’t just suggestions; they are the framework for the future of medicine. They provide the rules of the road that allow data to travel safely and meaningfully across borders.
| Standard | Primary Focus | Key Benefit |
|---|---|---|
| FHIR® | Clinical Data Exchange | RESTful APIs for real-time EHR access and mobile health integration |
| USCDI | Data Harmonization | Standardized data classes (e.g., medications, allergies) for US health exchange |
| GA4GH Beacon | Genomic Discovery | Privacy-preserving “discovery” of variants across federated networks |
| TEFCA | Network Governance | Pre-negotiated legal and technical agreements for nationwide connectivity |
| OMOP CDM | Research Analytics | Standardized format for observational data to enable large-scale studies |
Fast Healthcare Interoperability Resources (FHIR®) has become the global gold standard. Since the release of FHIR R4 in 2018, it has been widely adopted by certified developers. FHIR uses modern web technologies (JSON and REST) that make it easy for developers to build apps that sit on top of EHRs, much like apps sit on a smartphone OS. Meanwhile, the United States Core Data for Interoperability (USCDI) sets the “floor” for what data must be shareable. USCDI v4 and v5 have expanded to include social determinants of health (SDOH), which are critical for understanding health equity.
For those looking for the latest recognized specifications, the Interoperability Standards Advisory (ISA) provides a coordinated catalog of “recognized” standards to ensure you aren’t building on dead-end tech. The ISA is updated annually to reflect the maturity of emerging standards like those for public health reporting and laboratory results.
The Evolution of GA4GH Beacon for Genomic Discovery
The Beacon protocol is a brilliant solution to the “privacy vs. access” dilemma. It allows a researcher to ask a simple question: “Do you have any samples with this specific genetic mutation?” and receive a “Yes” or “No” without ever seeing sensitive patient identifiers. This “discovery” phase is essential for identifying which biobanks have the data needed for a specific study.
The Beacon Project Timeline shows a massive leap from v1 (simple presence/absence) to Beacon v2. The new version supports data interoperability solutions by adding:
- Complex Filters: Query by phenotypes, disease codes (using ICD-10 or SNOMED), sex, or age ranges.
- System Jumps: The ability to “jump” from a discovery result directly into a secure EHR or research environment (like a Trusted Research Environment) once proper permissions are granted.
- Standardized Models: Using the GA4GH Model to ensure genomic data is context-rich, including information about how the sequencing was performed.
Implementations like Progenetix have already curated over 113,000 CNV profiles, proving that federated discovery isn’t just a theory—it’s happening now. This allows a researcher in London to find a rare genetic variant in a dataset in Tokyo without the data ever leaving its home jurisdiction.
Streamlining Clinical Workflows with FHIR R4
The real magic happens when interoperability hits the clinic. By moving legacy ehr warehouses to fhir, hospitals can unlock real-time analytics. However, FHIR “bundles” (the way data is packaged) can be technically dense and difficult for standard SQL-based analytics tools to process.
A major breakthrough in clinical efficiency is “flattening FHIR bundles.” This process turns complex, nested JSON data into a format that can be queried instantly by data scientists. This allows for the creation of longitudinal records—a 360-degree view of the patient—that follows them from the GP to the specialist to the hospital. For a deep dive into how these records are structured, see our ehrs complete guide. When data is flattened and harmonized, AI models can be trained to predict patient deterioration or suggest personalized treatment plans based on the patient’s entire history, not just their most recent visit.
Federal Initiatives and the Future of Interoperable Systems
Governments are no longer asking for interoperability; they are mandating it. In the US, the 21st Century Cures Act has made “information blocking” illegal, forcing vendors to open up their systems or face significant fines. This shift in policy is designed to ensure that patients have easy access to their own health data via the apps of their choice.
Key drivers include:
- HHS Health IT Alignment Policy: A 2023 initiative to ensure all federal agencies—including the VA, CDC, and CMS—are pulling in the same direction regarding data standards. This prevents the “siloing” of data within different branches of the government.
- USCDI+: An extension of the core standard that adds mission-critical data elements for specific use cases like public health, behavioral health, or cancer research. USCDI+ allows for more granular data collection than the base standard.
- ehr claims integration: Linking clinical data with insurance claims to provide a full picture of patient outcomes and costs. This is vital for “value-based care” models where providers are paid based on patient health outcomes rather than the number of tests performed.
- TEFCA (Trusted Exchange Framework and Common Agreement): This creates a “network of networks,” allowing different health information exchanges (HIEs) to talk to each other. Under TEFCA, a hospital in California can securely request records from a clinic in Florida through a Qualified Health Information Network (QHIN).
Looking forward, FHIR R6 and the integration of AI will allow for automated “safety surveillance,” where algorithms monitor interoperable data streams to catch adverse drug reactions in real-time across the entire population. This would allow the FDA to identify issues with a new medication in days rather than months or years.
Reducing Clinician Burden with Advanced Data Interoperability Solutions
One of the biggest complaints from doctors is “death by a thousand clicks.” The AMA has been vocal in its feedback on USCDI v7, pushing for semantic clarity to ensure that health data integration complete guide strategies actually reduce documentation burden rather than increasing it. Interoperability should mean that data is entered once and used everywhere, rather than being re-typed into multiple systems.
By harmonizing disparate electronic health records, we can ensure that a “Referral Note” in one system automatically populates the “Appointment” field in another. This requires strict adherence to data privacy laws like HIPAA and GDPR, ensuring that while data flows freely between authorized systems, it remains locked away from unauthorized eyes. Advanced data interoperability solutions use automated consent management to ensure that patient preferences regarding data sharing are respected at every step of the process.
Emerging Approaches: The Semantic Data Charter (SDC)
We are seeing a shift toward “Vibe Coding”—where formal data models and AI generate applications that are natively interoperable. The Semantic Data Charter (SDC) leverages W3C standards like RDF (Resource Description Framework) and SPARQL to create “self-describing” data. This means the data itself contains the instructions on how it should be interpreted, reducing the need for external documentation.
Instead of custom code for every integration, SDC-compliant data carries its own “blueprint.” This is particularly powerful when using the omop complete guide for clinical research. By following a niaid data harmonization omop guide, researchers can export data in multiple formats (JSON, XML, RDF) without losing the underlying meaning or lineage. This “polyglot” approach to data ensures that researchers can use the tools they are most comfortable with—whether that’s R, Python, or SQL—without worrying about data compatibility issues. As we move toward a future of federated AI, these semantic standards will be the foundation upon which we build the next generation of medical breakthroughs.
Frequently Asked Questions about Data Interoperability
What is the difference between technical and semantic interoperability?
Technical interoperability is the “plumbing”—the ability of two systems to send and receive bits and bytes (e.g., via a REST API or secure file transfer). It ensures the connection is established. Semantic interoperability is the “language”—the ability of those systems to understand the meaning of the data exchanged. It ensures that when a system receives a code like “C0005823,” it knows this refers specifically to “Essential Hypertension” and not just a generic blood pressure reading. Without semantic interoperability, the data is received but cannot be acted upon automatically.
How does the GA4GH Beacon protocol protect patient privacy during discovery?
Beacon uses a federated query model. Instead of moving sensitive patient data to a central location, the query travels to the data. It only returns aggregate information (like “Yes, this variant exists in our cohort”) or uses “noise” (differential privacy) to ensure that individuals cannot be re-identified through their genetic markers. This allows researchers to find the data they need without ever seeing a patient’s name, address, or full genetic sequence until they have gone through a formal ethics approval process.
Why is FHIR R4 considered the foundational version for US health IT?
Released in 2018, FHIR R4 was the first version to include “normative” content—meaning the core parts of the standard are stable and won’t change in ways that break existing systems. This stability gave federal agencies like the ONC and private developers the confidence to invest in it as a long-term national standard. While newer versions like R5 and R6 introduce more features, R4 remains the baseline for regulatory compliance in the United States.
What is the role of a Trusted Research Environment (TRE) in interoperability?
A TRE is a secure space where researchers can access and analyze sensitive data without downloading it. Interoperability solutions allow different TREs to be “federated,” meaning a researcher can run the same analysis script across multiple TREs simultaneously. This is the gold standard for privacy-preserving research, as it ensures that data stays behind the firewall of the institution that collected it while still contributing to global scientific knowledge.
Conclusion
At Lifebit, we believe that the future of medicine is federated. The era of moving massive datasets into central silos is ending, replaced by a more secure, efficient, and scalable model where the analysis goes to the data. Our next-generation platform provides the data interoperability solutions needed to bridge the gap between global data silos, enabling researchers to query across borders while maintaining total data sovereignty.
By utilizing our Trusted Research Environment (TRE) and Trusted Data Lakehouse (TDL), organizations can achieve secure, real-time access to multi-omic data without the risks and costs of traditional data movement. Our architecture is built on the open standards discussed here—FHIR, GA4GH, and OMOP—ensuring that your infrastructure is future-proof and ready for the AI revolution.
Whether you are a biopharma giant looking to accelerate your drug discovery pipeline or a public health agency aiming to improve population health outcomes, our lifebit-federated-biomedical-data-platform delivers the federated governance and AI-driven insights required for modern research. Don’t let your data stay trapped in the past, limited by incompatible systems and manual processes. Explore our data interoperability solutions today and start building the connected healthcare ecosystem of tomorrow.

