In Depth Guide to Multi-Omics Data

Multi-Omics Data: Stop Missing Disease Signals — Get the Full Picture Fast
Multi-omics data refers to the combined measurement and analysis of multiple biological layers — including genomics, transcriptomics, proteomics, metabolomics, and epigenomics — to build a more complete picture of how living systems work.
Here is a quick overview of what multi-omics data includes:
| Omics Layer | What It Measures |
|---|---|
| Genomics | DNA sequence and structural variation |
| Transcriptomics | Gene expression via RNA |
| Proteomics | Protein abundance and function |
| Metabolomics | Small molecules and metabolic activity |
| Epigenomics | Chemical modifications that regulate gene activity |
| Microbiomics | Microbial communities and their interactions |
Single-omics studies — like looking at genomics alone — give you one piece of the puzzle. But biology doesn’t work in isolation. A gene variant means little without knowing whether it’s expressed, what proteins it produces, or how those proteins behave in a metabolic context.
That gap between genotype and phenotype is exactly where multi-omics comes in.
The research community recognized this fast. Publications mentioning multiomics grew from zero in 2000 to more than 1,400 per year by 2021 — an exponential rise driven by falling sequencing costs, better computational tools, and the clear limits of studying one biological layer at a time.
The result? Researchers can now ask — and answer — questions that were simply out of reach a decade ago. From cancer subtyping to vaccine response to infectious disease, multi-omics is reshaping how we understand and treat complex conditions.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, and with over 15 years in computational biology and biomedical data integration, I’ve seen how multi-omics data unlocks insights that no single dataset can provide. In this guide, I’ll walk you through everything you need to know — from the core data types and analysis tools to real-world clinical applications and the infrastructure needed to do it at scale.
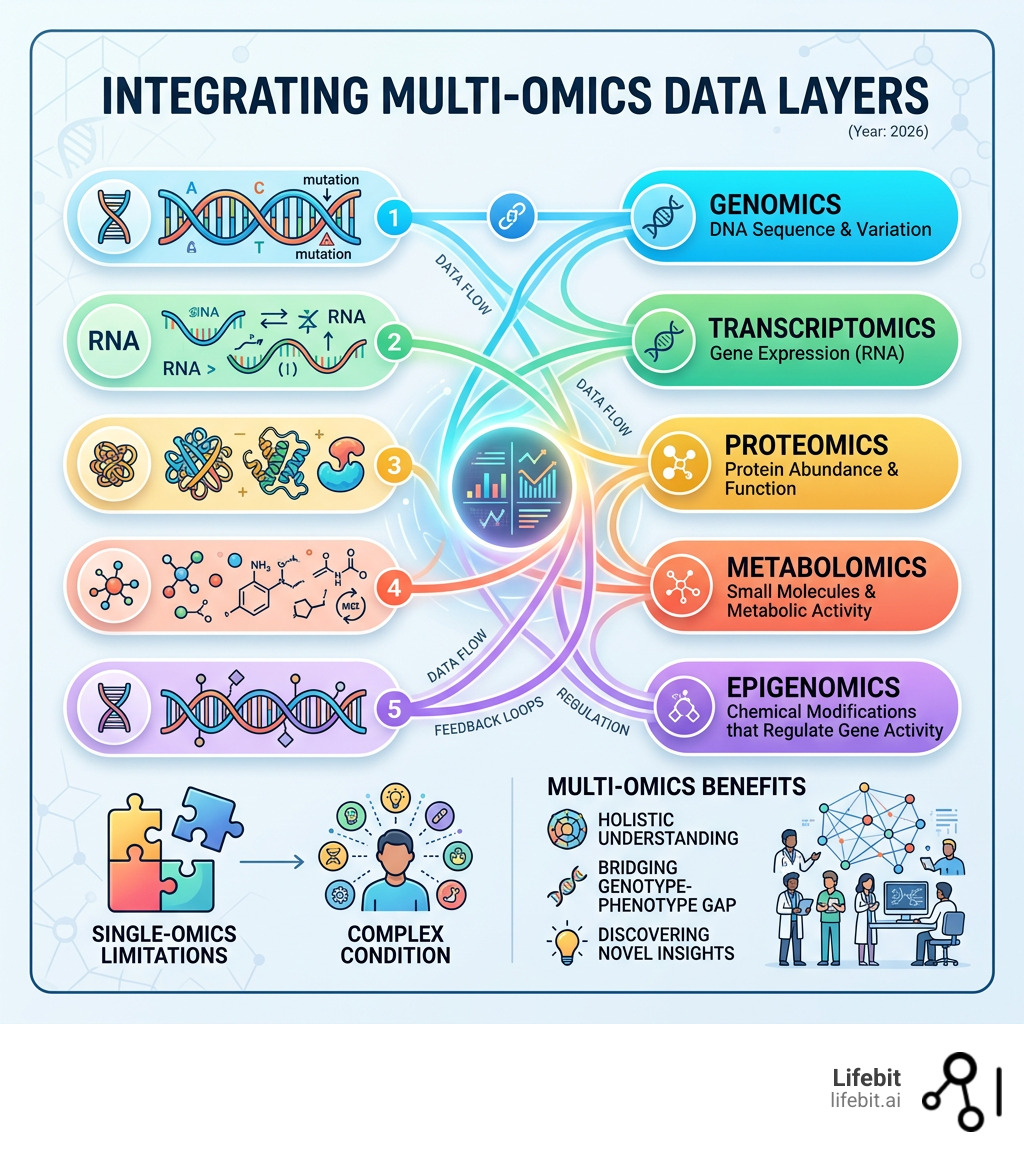
Multi-Omics Data: What You Miss With Genomics Alone (And How to Fix It)
For years, genomics was the “main character” of precision medicine. We thought if we could just map the DNA, we’d have the manual for human health. But as it turns out, the genome is more like a library of blueprints; whether a house actually gets built—and how it looks—depends on the builders (proteins), the materials available (metabolites), and the environment.
Integrating multi-omics allows us to bridge the “genotype-phenotype gap.” By assessing information flow across these layers, we can see not just what might happen (DNA), but what is happening (RNA and proteins) and what has happened (metabolites). This holistic approach is essential because biological systems are characterized by non-linear relationships and feedback loops.
The Biological Cascade: Beyond the Central Dogma
The traditional “Central Dogma” of molecular biology suggests a linear flow: DNA → RNA → Protein. However, multi-omics data reveals a far more complex reality. Feedback loops exist at every level. For instance, metabolites can act as signaling molecules that modify the epigenome, which in turn regulates gene expression. Without a multi-omic view, these regulatory circuits remain invisible.
The primary omics data types we integrate include:
- Genomics: The foundation. It identifies mutations, SNPs, and copy number variations (CNVs). It tells us about the inherited risk and the potential for certain biological processes to occur.
- Transcriptomics: Measures the “active” part of the genome. Using techniques like RNA-Seq, it tells us which genes are turned on or off in response to a stimulus, providing a snapshot of cellular intent.
- Proteomics: Proteins are the functional workhorses. Proteomics captures the actual physical state of the cell, including post-translational modifications that DNA sequences cannot predict.
- Metabolomics: The end products of cellular processes. This layer is closest to the actual phenotype and clinical symptoms, reflecting the sum of genetic activity and environmental influences like diet and medication.
- Epigenomics: The “volume knobs” of the cell, such as DNA methylation and histone modification, which control gene accessibility without changing the underlying sequence.
According to scientific research on multi-omics integration, the mathematical challenge is significant. We aren’t just adding datasets together; we are looking for correlations across different scales and noise patterns. However, the reward is a massive increase in phenotype prediction accuracy.
Single-Cell vs. Bulk Multi-Omics Data
In the past, we relied on “bulk” analysis—essentially taking a tissue sample, putting it in a blender, and measuring the average signal. The problem? If you have a tumor with a few highly aggressive cells hidden among healthy ones, the bulk average might miss them entirely. This “averaging effect” has historically led to the failure of many clinical trials because the underlying cellular heterogeneity was ignored.
Single-cell multi-omics data solves this by measuring multiple modalities within a single cell. This allows for:
- Resolution of Heterogeneity: Identifying rare cell types, such as cancer stem cells or specific immune subsets, that drive disease progression.
- Lineage Tracing: Understanding the developmental trajectories of cells, seeing how a stem cell differentiates into a specialized cell or how a healthy cell transforms into a malignant one.
- Cellular State Transitions: Seeing exactly when a cell starts becoming “exhausted” or cancerous, allowing for earlier intervention.
Key methods like G&T-seq (Genome and Transcriptome sequencing) and CITE-seq (Cellular Indexing of Transcriptomes and Epitopes) allow us to see DNA/RNA or RNA/Protein levels simultaneously in the same cell. Furthermore, the burgeoning field of Spatial Omics preserves the physical orientation of cells within a tissue, showing us how cells “talk” to their neighbors and how the microenvironment influences disease. You can explore more about these Single-cell and Spatial methods to see how they are catching up to traditional single-cell transcriptomics.
Read more on research on single-cell multi-omics strategies to understand how these strategies are being applied to map human development.
The Evolution of Multi-Omics Data Research
The journey to multi-omics data didn’t happen overnight. It started with the discovery of the DNA double helix in 1953, followed by the development of transcriptome analysis in the 1980s. However, the real “boom” occurred after the completion of the Human Genome Project, which provided the reference map needed to anchor other omics layers.
A major milestone was the historical method for sequential separation published in 1967, which first proposed separating proteins, RNA, and DNA from a single tissue homogenate. Fast forward to today, and we have sophisticated protocols like MPLEx and MOST that do this with incredible precision, ensuring that the data from different layers is truly matched from the same biological source.
We’ve also seen the rise of massive consortiums. The The Cancer Genome Atlas (TCGA) has profiled over 20,000 samples across 33 cancer types, while the Human Microbiome Project (HMP) invested $170 million to understand how our microbial “second genome” interacts with our own. This explosion in data volume has turned biology into a data science problem, where the bottleneck is no longer sequencing, but the computational power to integrate these disparate streams.
Multi-Omics Data Integration: Stop Drowning in Data — Start Getting Answers
The biggest hurdle in multi-omics data isn’t collecting the data—it’s making sense of it. Each “ome” has its own language, scale, and noise. Genomics data is often binary (mutation present or not), while metabolomics data is continuous and highly variable. Furthermore, the “curse of dimensionality” is a constant threat; we often have thousands of features (genes, proteins) but only a few hundred samples, making it easy for models to find false correlations.
Machine learning (ML) is the glue that holds these layers together. We generally categorize integration into two strategies:
| Strategy | Approach | Best For… |
|---|---|---|
| Early Integration | Concatenating all data into one giant matrix before analysis. | Simple datasets where features are highly correlated and scales are similar. |
| Late Integration | Analyzing each omic layer separately and then merging the results. | Handling datasets with very different noise levels, missing values, or different sample sizes. |
| Intermediate Integration | Using joint dimensionality reduction to find shared latent spaces. | Complex biological systems where we want to find the underlying drivers across all layers. |
Advanced ML techniques like Dimensionality Reduction (e.g., PCA, t-SNE, or UMAP) help us find the “hidden factors” that explain the most variation across all layers. Deep Learning and Latent Variable Path Modeling are now being used to predict how a change in the epigenome will ripple down to the metabolome. For example, Variational Autoencoders (VAEs) can learn a compressed representation of multi-omic data, allowing researchers to visualize complex disease states in a 2D or 3D space.
Overcoming the Computational Bottleneck
To successfully integrate multi-omics data, researchers must address several technical hurdles:
- Data Normalization: Ensuring that the high signal of transcriptomics doesn’t drown out the subtle signals of metabolomics.
- Batch Effect Correction: Removing technical noise introduced when samples are processed at different times or in different labs.
- Missing Value Imputation: Multi-omics studies often have “holes” in the data (e.g., a sample has genomics but not proteomics). Advanced algorithms now use the available layers to predict the missing ones.
Tools like iOmicsPASS use network analysis to discover “predictive subnetworks”—groups of genes, proteins, and metabolites that work together to drive a specific disease phenotype. This is far more powerful than looking for a single “smoking gun” biomarker because it captures the functional modules of the cell.
Top Software Tools for Multi-Omics Data Analysis
If you are a researcher looking to dive into the data, the R/Bioconductor ecosystem is your best friend. There are now over 99 software packages dedicated to this field. Some of our favorites include:
- mixOmics: A powerhouse for feature selection and multiple data integration using multivariate methods like PLS (Partial Least Squares).
- MultiAssayExperiment: Provides a standard Bioconductor interface for managing overlapping samples across different assays, ensuring data integrity.
- omicade4: Excellent for multiple co-inertia analysis to find common structures in multi-omic datasets.
- bioCancer: A package specifically designed for the visualization of multi-omic cancer data, making complex results accessible to clinicians.
- IMAS: Focused on using multi-omics to evaluate alternative splicing, a key driver of protein diversity.
- Omics Pipe: A Python-based framework that automates multi-omics analysis pipelines for reproducibility, essential for large-scale pharma research.
Using these tools, we can move from raw sequences to biological insights in a fraction of the time it used to take, turning raw data into actionable clinical hypotheses.
Multi-Omics Data in Healthcare: Find Better Targets Faster (Or Fall Behind)
The true value of multi-omics data is seen in the clinic. In Precision Oncology, for example, we no longer treat “lung cancer” as a single disease. By using resources like LinkedOmics, which connects TCGA data across 32 cancer types, we can identify specific molecular subtypes that respond to different targeted therapies. This has led to the discovery of “basket trials,” where patients with different types of cancer but the same multi-omic signature are treated with the same drug.
Beyond Oncology: Expanding the Reach of Multi-Omics
While cancer has been the early adopter, multi-omics is now transforming other fields:
- Rare Disease Diagnostics: For many patients with rare diseases, genomics alone provides a diagnosis in only 25-30% of cases. By adding transcriptomics and metabolomics, researchers can identify functional abnormalities that DNA sequencing missed, raising the diagnostic yield significantly.
- The Integrative Human Microbiome Project: This study showed how the gut microbiome interacts with host genetics to influence conditions like pre-diabetes and Inflammatory Bowel Disease (IBD). You can read more about the Integrative Human Microbiome Project and its findings on host-microbe dynamics. It revealed that the function of the microbiome (what it produces) is often more important than which specific species are present.
- Systems Immunology: By integrating multi-modal genomic and multi-omics data for precision medicine, researchers are developing “systems vaccinology.” This helps predict who will have a strong immune response to a vaccine and who might suffer side effects, allowing for personalized dosing schedules.
- Cardiovascular Disease: The cardiovascular disease database (C/VDdb) provides a multi-omics expression profiling resource to identify non-obvious therapeutic targets for heart health, such as specific lipid metabolites that drive plaque formation.
- Pharmacogenomics 2.0: We are moving beyond simple “slow vs. fast metabolizer” genetic tests. Multi-omics allows us to see how a patient’s current proteomic and metabolomic state influences drug efficacy and toxicity in real-time.
- COVID-19 Response: Multi-omics was instrumental in identifying the “cytokine storm” and lipid dysregulation associated with severe cases. By analyzing the blood of patients across all omic layers, researchers identified specific metabolic signatures that could predict which patients would require intensive care weeks before their symptoms worsened.
Through these efforts, we are moving toward a roadmap for personalized immunology where treatments are tailored to your unique molecular fingerprint, reducing the “trial and error” approach that currently dominates medicine.
Multi-Omics Data Access Is Broken: How Federated Analysis Fixes It
Despite its promise, multi-omics data faces a major “Access Crisis.” Most of this data is locked in “silos”—different hospitals, universities, and countries have strict rules about sharing sensitive genetic information. For the pharma industry, multi-omic health data access is the number one bottleneck to drug discovery. Moving petabytes of multi-omic data is not only slow and expensive but often legally impossible due to data sovereignty laws.
At Lifebit, we believe the solution isn’t to move the data, but to move the analysis to the data. This is called Federated Analysis.
The Pillars of a Modern Multi-Omics Infrastructure
Our platform provides a secure federated ecosystem where researchers can access global biomedical data in real-time without violating privacy laws (like GDPR or HIPAA). By using Trusted Research Environments (TREs) and a Trusted Data Lakehouse (TDL), we enable a new paradigm of research:
- Data Harmonization: Making sure “Genomics A” from a cohort in Japan can talk to “Proteomics B” from a cohort in the UK. We use standardized ontologies and metadata schemas to ensure that disparate datasets are interoperable.
- Secure Collaboration: Allowing scientists in London and New York to work on the same dataset without the data ever leaving its original jurisdiction. The raw data stays behind the provider’s firewall; only the aggregated, non-identifiable results are shared.
- Scalable Infrastructure: Running complex ML models on petabytes of data using cloud-native power. Multi-omics integration requires massive compute resources; our federated approach allows researchers to tap into the power of the cloud where the data lives.
- Zero-Trust Security: Implementing advanced encryption and strict access controls. Every action taken on the data is logged and auditable, ensuring that patient privacy is never compromised.
This approach is how we drive innovation in omics data analysis, ensuring that the next big breakthrough isn’t delayed by a paperwork nightmare. By breaking down silos, we allow for the creation of “virtual cohorts” of millions of individuals, providing the statistical power needed to find the rarest disease signals.
The Future: AI-Driven Discovery at Scale
As we move forward, the integration of Generative AI and Large Language Models (LLMs) with multi-omics data will allow researchers to query biological databases using natural language. Imagine asking, “Find all metabolic pathways upregulated in patients with this specific genetic mutation who also failed to respond to anti-PD1 therapy,” and getting an answer in seconds. This is the future Lifebit is building—a world where data access is no longer a barrier to saving lives.
Multi-Omics Data FAQs: Biggest Problems, Best Databases, Fast Answers
What are the primary challenges in collecting multi-omics data?
The biggest challenges are data heterogeneity (different formats) and high costs. Additionally, samples must be handled carefully to avoid degradation; for example, RNA is much more fragile than DNA. Batch effects—where data looks different just because it was processed on a different day—can also lead to false conclusions if not corrected by software.
How does single-cell multi-omics provide an advantage?
It provides the ultimate resolution. While bulk analysis gives you the “average” of a tissue, single-cell analysis lets you see rare cell types, such as the specific immune cells attacking a tumor. It also allows for Spatial Omics, which tells you where those cells are located. You can see more on Spatial Omics methods and how they preserve tissue architecture.
Which databases are best for multi-omics research?
There are several world-class resources available:
- ProteomicsDB: A multi-organism resource for life science research.
- LinkedOmics: Ideal for cancer-specific multi-omics.
- PaintOmics: A web resource for pathway analysis and visualization.
- TCGA (The Cancer Genome Atlas): The gold standard for cancer datasets.
- MOPED: A multi-omics profiling expression database.
- ZikaVR: An integrated resource for Zika virus genomics and proteomics.
Multi-Omics Data: The Old One-Layer Approach Is Costing You Discoveries
The era of looking at biology through a single lens is over. Multi-omics data has proven that the most profound insights live in the intersections between our DNA, our proteins, and our environment. The integration of AI and machine learning will only get deeper, allowing us to predict disease before it even manifests.
However, the future of medicine depends on our ability to access this data securely. At Lifebit, we are building the “operating system” for this new era—a platform for every insight that connects the world’s most valuable biomedical data through federated AI.
Whether you are a researcher in a lab or a clinical lead at a major pharma company, the tools to transform human health are now within reach. It’s time to stop looking at the pieces and start seeing the whole picture.
For more information on how we can help you unlock your data’s potential, visit Lifebit.

