Mapping the Future: Top Genomic Research Hubs in New York

New York Genomic Research: Map 148K Genomes Faster with 5 Top Hubs
New york genomic research is anchored by some of the most advanced institutions on the planet — and if you’re trying to locate them quickly, here’s the short answer:
Top genomic research institutions in New York:
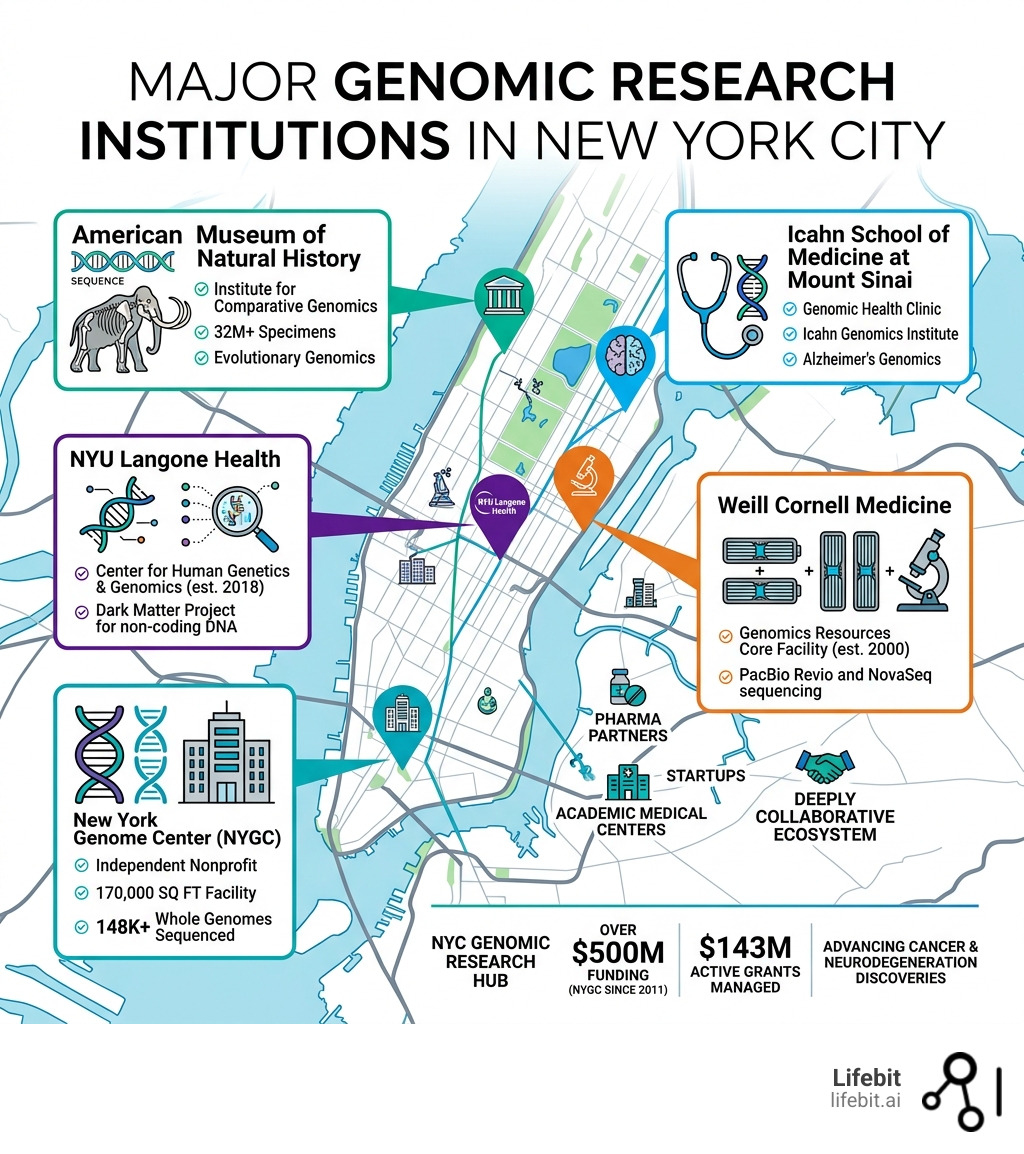
- New York Genome Center (NYGC) — Independent nonprofit; 170,000 SF facility; 148K+ whole genomes sequenced
- Icahn School of Medicine at Mount Sinai — Genomic Health Clinic; Icahn Genomics Institute; Alzheimer’s genomics
- NYU Langone Health — Center for Human Genetics & Genomics (est. 2018); Dark Matter Project for non-coding DNA
- Weill Cornell Medicine — Genomics Resources Core Facility (est. 2000); PacBio Revio and NovaSeq sequencing
- American Museum of Natural History — Institute for Comparative Genomics; 32M+ specimens; evolutionary genomics
New York’s genomic ecosystem is not just large — it’s deeply collaborative. Academic medical centers, nonprofits, pharma partners, and life science startups all operate in close proximity, sharing data, infrastructure, and talent. The result is a city-wide engine driving discoveries in cancer, neurodegeneration, rare disease, and preventive health.
Since its founding in 2011, the New York Genome Center alone has raised over $500 million in funding and sequenced more than 148,000 whole genome equivalents. That scale matters — both for the science and for the systems needed to manage, analyze, and share that data securely.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit and a computational biologist with over 15 years of experience building genomic data infrastructure used by institutions operating at the scale of new york genomic research. My work on federated AI platforms and tools like Nextflow was built precisely to solve the data access and compliance challenges these institutions face every day.
New York Genomic Research: Inside the $500M Hub Powering Breakthroughs
If you are looking for the heart of new york genomic research, look no further than 101 Avenue of the Americas. This is where the New York Genome Center (NYGC) makes its home. Spanning a massive 170,000 square feet, the NYGC isn’t just a lab; it’s a collaborative hub that connects the city’s leading academic and medical institutions. The facility serves as a beacon for high-throughput sequencing, providing the physical and intellectual infrastructure necessary to handle the most complex biological questions of our time.
The primary mission of the NYGC is to advance genomic discovery to improve healthcare. They do this by acting as a central nexus for whole genome sequencing, analysis, and tool development. Since its inception, the center has raised over $500 million from federal grants, New York State, and generous philanthropy. As of late 2023, they manage roughly $143 million in active grants, fueling high-impact studies across the board. This financial backing allows the center to maintain a level of technological sophistication that few other places on Earth can match, ensuring that New York remains at the absolute forefront of the genomic revolution.
Under the leadership of Dr. Bing Ren, the current Scientific Director and CEO, the NYGC has become a “Manhattan Project” for genomics. They operate a state-of-the-art facility that includes 30,000 square feet of dedicated sequencing lab space. To understand the scale, consider this: they can sequence more than 60 whole human genomes per day. That is a staggering amount of data, and it is exactly why the field of genomics requires such robust computational infrastructure. The center’s high-performance computing (HPC) cluster is one of the largest dedicated to genomics in the Northeast, capable of processing petabytes of raw sequencing data into actionable clinical insights in record time.
One of their standout features is their CLIA-certified and CLEP-accredited clinical laboratory. This allows them to perform cutting-edge oncology and genetics tests that are approved in all 50 U.S. states. When we talk about bringing research to the bedside, this is what it looks like. They aren’t just finding variants; they are helping doctors treat patients by identifying the specific genetic drivers of a patient’s cancer or rare disease. This clinical integration is vital for the “bench-to-bedside” pipeline, ensuring that a discovery made in the lab on Monday can potentially influence a treatment plan in the clinic by Friday.
Leading the Charge in Cancer and Neurodegenerative Disease
The NYGC doesn’t just sequence DNA; they dive deep into the specific mechanisms of disease. Their core faculty and labs are world-renowned for their work in three major areas: cancer, neurodegenerative diseases (like ALS and Alzheimer’s), and neuropsychiatric disorders (like autism and schizophrenia). The center’s approach is inherently multi-disciplinary, bringing together biologists, clinicians, and computer scientists to tackle these problems from every possible angle.
- The Ren Lab: Led by Dr. Bing Ren, this lab is a pioneer in epigenomics. They focus on how the “switches” in our DNA turn genes on and off. Their work mapping millions of regulatory elements is critical for understanding why certain cells become cancerous or why neurons begin to fail in aging brains. By utilizing technologies like Hi-C and ATAC-seq, they are creating a 3D map of the genome that shows how distant pieces of DNA interact to regulate gene expression.
- The Zhu Lab: Dr. Chenxu Zhu and his team use single-cell multiomics to study brain development and aging. By looking at individual cells, they can identify the “vulnerability code” that makes certain parts of the brain susceptible to disease. This high-resolution view is essential because traditional “bulk” sequencing can mask the rare cell types that are often the first to succumb to neurodegeneration.
- The Lappalainen Lab: Directed by Tuuli Lappalainen, this lab studies functional genetic variation. They want to know how differences in our DNA affect the transcriptome—the set of all RNA molecules in a cell—and how those changes lead to disease. Her work is foundational for interpreting the thousands of genetic variants discovered in large-scale population studies, helping to distinguish between harmless mutations and those that cause illness.
These labs don’t work in a vacuum. For example, their neurodegenerative disease research involves massive collaborations with groups like Target ALS, studying gene expression changes across different stages of the disease to find new therapeutic targets. The NYGC also hosts the Center for Genomics of Neurodegenerative Disease (CGND), which acts as a global repository for ALS genomic data, providing researchers worldwide with the resources they need to accelerate the search for a cure.
Scaling New York Genomic Research with Advanced Sequencing Capacity
The sheer technical capacity of the NYGC is what allows new york genomic research to compete on a global scale. Their lab houses an impressive fleet of machines, including Illumina HiSeq 2500s, HiSeq Xs, and the newer NovaSeq 6000 and NovaSeq X Plus models. The addition of the NovaSeq X Plus has been a game-changer, significantly reducing the cost and time required for whole genome sequencing, which in turn allows for much larger study cohorts.
To date, they have sequenced over 148,000 whole genome equivalents. This level of output generates petabytes of data—specifically, their collaboration on the Illumina Billion Cell Atlas has generated 3.1 petabytes of data from over 150 million single cells. This is exactly the kind of “big data” that powers use case population genomics, where researchers look at tens of thousands of people to find the rare genetic needles in the haystack. Managing this data requires not just storage, but sophisticated AI and machine learning algorithms to identify patterns that would be invisible to the human eye. The NYGC’s bioinformatics team is constantly developing new software tools to handle this deluge, many of which are released as open-source resources for the global scientific community.

New York Genomic Research: How NYU and Mount Sinai Stop Reactive Medicine
While the NYGC acts as a central hub, individual institutions like NYU Langone and Mount Sinai are pushing the boundaries of what we can do with that data. These institutions are not just participants in the genomic revolution; they are its architects, building the tools and clinical frameworks that will define the next century of medicine.
At NYU Langone, The Dark Matter Project is tackling one of the biggest mysteries in biology. Did you know that less than 2% of your genome actually codes for proteins? The other 98% is often called “dark matter.” For decades, this portion of the genome was dismissed as “junk DNA,” but we now know it contains the complex regulatory instructions that govern when and where genes are turned on. This project uses “Big DNA” technology—synthetic genomics—to build large DNA constructs from scratch. By “humanizing” mice (replacing mouse genes with human ones to create GREAT-GEMMs, or Genomically-Rewritten Endogenous Animals for Therapeutics), they can study how this dark matter regulates our genes and contributes to disease risk in a living organism. This is a massive leap forward from traditional methods, providing “Google Maps” style precision compared to the “dime-store compass” of older techniques. Researchers can now systematically swap out segments of the non-coding genome to see exactly which changes lead to developmental defects or disease susceptibility.
This kind of advanced research is exactly why we believe in the importance of genomic medicine accessibility. If we can decode the dark matter, we can build better gene therapies and drugs for everyone, not just those with mutations in the well-understood protein-coding regions. NYU’s Institute for Systems Genetics is a leader in this space, merging engineering principles with biological research to “write” DNA as easily as we currently “read” it.
Advancing New York Genomic Research Through Preventive Health Clinics
Mount Sinai is taking a different, more clinical approach through its Genomic Health Clinic and the Institute for Genomic Health. They are moving from reactive medicine (treating you when you’re sick) to proactive medicine (preventing you from getting sick in the first place). This shift is powered by the BioMe Biobank, one of the largest and most diverse biorepositories in the world, which contains genetic and clinical data from over 60,000 Mount Sinai patients.
Statistics show that 1 in 75 people may be at high genomic risk for conditions like hereditary breast and ovarian cancer (HBOC), Lynch syndrome, or familial hypercholesterolemia. The clinic offers elective genetic testing and one-on-one counseling to identify these risks early. When a high-risk variant is found, patients are integrated into specialized surveillance programs, often catching diseases in their earliest, most treatable stages. They also specialize in pharmacogenomics—using your genetic profile to determine which medications will work best for you and which ones might cause adverse reactions. For example, certain common blood thinners or antidepressants can be ineffective or even dangerous depending on a patient’s CYP450 gene variants. By testing for these variants before a prescription is written, Mount Sinai is eliminating the “trial and error” approach to prescribing. It’s personalized medicine in its purest form, ensuring that every patient receives the right dose of the right drug at the right time.
New York Genomic Research: How Weill Cornell and AMNH Scale Discovery
Not all new york genomic research happens in a hospital. Some of it happens in the “core facilities” that support thousands of researchers, and some happens in museums! These institutions provide the breadth and depth that make the New York ecosystem so unique, covering everything from the molecular details of a single patient’s tumor to the broad evolutionary history of life on Earth.
Weill Cornell Medicine’s Genomics Resources Core Facility (GRCF) has been a staple of the NYC research community since 2000. They provide a comprehensive suite of services, from sample quality control and library preparation to advanced long-read sequencing using the PacBio Revio. Long-read sequencing is particularly important for resolving complex structural variants and repetitive regions of the genome that short-read technologies often miss. This facility is essential for the basic and translational research communities at Weill Cornell, NewYork-Presbyterian, and the Hospital for Special Surgery. They also offer cutting-edge spatial transcriptomics services, such as 10x Genomics Visium, which allows researchers to see exactly where specific genes are being expressed within a tissue sample. This “geography of the cell” is crucial for understanding how the microenvironment of a tumor influences its growth and resistance to treatment.
On the other side of Central Park, the American Museum of Natural History (AMNH) operates the Institute for Comparative Genomics. While most labs focus on human health, AMNH looks at the 3.8 billion-year history of life. They use their collection of over 32 million specimens and cultural artifacts to map evolutionary relationships across the entire Tree of Life. Their Ambrose Monell Cryo Collection (AMCC) is a frozen tissue library that serves as a vital resource for conservation and disease ecology. By sequencing the genomes of endangered species, AMNH researchers can identify populations with low genetic diversity and develop strategies to prevent extinction. Furthermore, their work in comparative genomics helps us understand the origins of human traits and diseases by looking at how similar genes function in other organisms. For instance, studying the regenerative abilities of certain amphibians or the extreme longevity of some whale species can provide clues for human regenerative medicine and aging research.
| Feature | Clinical Sequencing (e.g., Mount Sinai) | Evolutionary Genomics (e.g., AMNH) |
|---|---|---|
| Primary Goal | Patient diagnosis and treatment | Understanding species evolution |
| Sample Type | Patient blood, saliva, or tumor tissue | Frozen tissues, museum specimens, ancient DNA |
| Key Technology | Whole Genome/Exome Sequencing (WGS/WES) | Comparative phylogenomics |
| Outcome | Personalized medicine and drug discovery | Conservation and biodiversity insights |
Driving Innovation in New York Genomic Research via Institutional Collaboration
The secret sauce of New York’s success is collaboration. The NYGC was founded by 12 institutions, including Rockefeller University, NYU, Columbia, and Memorial Sloan Kettering. This “federated” model allows these giants to share the high costs of sequencing infrastructure while maintaining their independent research programs. It prevents the duplication of expensive resources and fosters an environment where data and ideas flow freely across institutional boundaries. This is exemplified by the “New York City Virus Hunter” program, a collaborative effort that monitors local wildlife and environment for emerging pathogens, using the same genomic tools developed for human health.
We also see this in the startup ecosystem. JLABS@NYC, located within the NYGC facility, hosts 26 life science startups. These companies get access to world-class equipment and expertise, helping them turn genomic discoveries into viable businesses. This collaborative spirit is very similar to how we view the global research community—much like the efforts seen in the UK Biobank, where shared data leads to exponential breakthroughs. By creating a dense network of academic, clinical, and commercial partners, New York has built a self-sustaining engine for innovation that attracts the best talent from around the world.
New York Genomic Research: 3 Critical Questions Answered Now
What is the primary mission of the New York Genome Center?
The NYGC is an independent, nonprofit research institution dedicated to advancing genomic discovery. Its mission is to transform healthcare by leveraging whole genome sequencing and analytics to understand the genetic basis of diseases like cancer and neurodegenerative disorders. It serves as a collaborative hub for its member institutions.
Which institutions are founding members of the NYGC?
The NYGC was established through a collaboration of several leading institutions, including Cold Spring Harbor Laboratory, Columbia University, Albert Einstein College of Medicine, Icahn School of Medicine at Mount Sinai, Memorial Sloan Kettering Cancer Center, NYU, NewYork-Presbyterian Hospital, Northwell Health, Rockefeller University, Stony Brook University, and Weill Cornell Medicine.
How does New York genomic research contribute to clinical cancer testing?
New York is a leader in clinical genomics. The NYGC, for instance, operates a CLIA-certified and CLEP-accredited lab that offers the first New York State-approved clinical whole genome and transcriptome test for cancer. This allows for precise variant detection in tumors, helping oncologists choose the most effective treatments for their patients.
New York Genomic Research: Stop Wasting Time and Start Solving Disease
The landscape of new york genomic research is breathtakingly complex and incredibly productive. From the high-throughput pipelines of the NYGC to the “Big DNA” synthetic genomics at NYU Langone, the city is a literal engine for human health. But all this sequencing—over 148,000 genomes and counting—creates a massive data challenge. The sheer volume of information is overwhelming traditional data management systems, leading to “data silos” where valuable information is trapped within a single institution’s servers.
At Lifebit, we know that the biggest bottleneck in genomics isn’t the sequencing itself; it’s the ability to access and analyze that data securely without moving it. Moving petabytes of genomic data is not only slow and expensive but also raises significant security and privacy concerns, especially under strict regulations like HIPAA and New York’s own health data laws. Our next-generation federated AI platform is built to solve this. We provide Trusted Research Environments (TREs) and Trusted Data Lakehouses (TDLs) that allow biopharma companies, governments, and researchers to run advanced AI/ML analytics on multi-omic data while keeping that data compliant and secure at its source. This federated approach means the data never leaves the institution’s secure environment; only the analysis results are shared.
Our platform is designed to handle the diversity of data types generated in New York, from the whole genome sequences at the NYGC to the clinical records at Mount Sinai and the spatial transcriptomics at Weill Cornell. By providing a unified interface for this data, we enable researchers to perform cross-institutional studies that were previously impossible. Whether you are looking for a definition of sequencing or you are ready to scale a global population genomics project, we are here to provide the infrastructure that makes “impossible” research possible. The future of medicine is being mapped right now in New York—and we are proud to provide the tools that help researchers navigate it, ensuring that every byte of data contributes to a faster cure and a healthier future for all.

