Safe Haven: Why Your Data Needs a Secure Research Environment

TRE Secure Data Environment: Stop Data Breaches Without Slowing Research
TRE secure data environment solutions provide the answer to a critical challenge facing modern biomedical research: how do you open up the insights hidden in sensitive patient data while ensuring absolute privacy and regulatory compliance? A Trusted Research Environment (TRE)—also known as a Secure Data Environment (SDE) or Data Safe Haven—is a highly secure computing environment that allows approved researchers to analyze de-identified health, genomic, and clinical data without that data ever leaving its protective walls.
Key features of a TRE secure data environment:
- Centralized secure access – Data remains in a protected environment; researchers bring their questions to the data, not the other way around
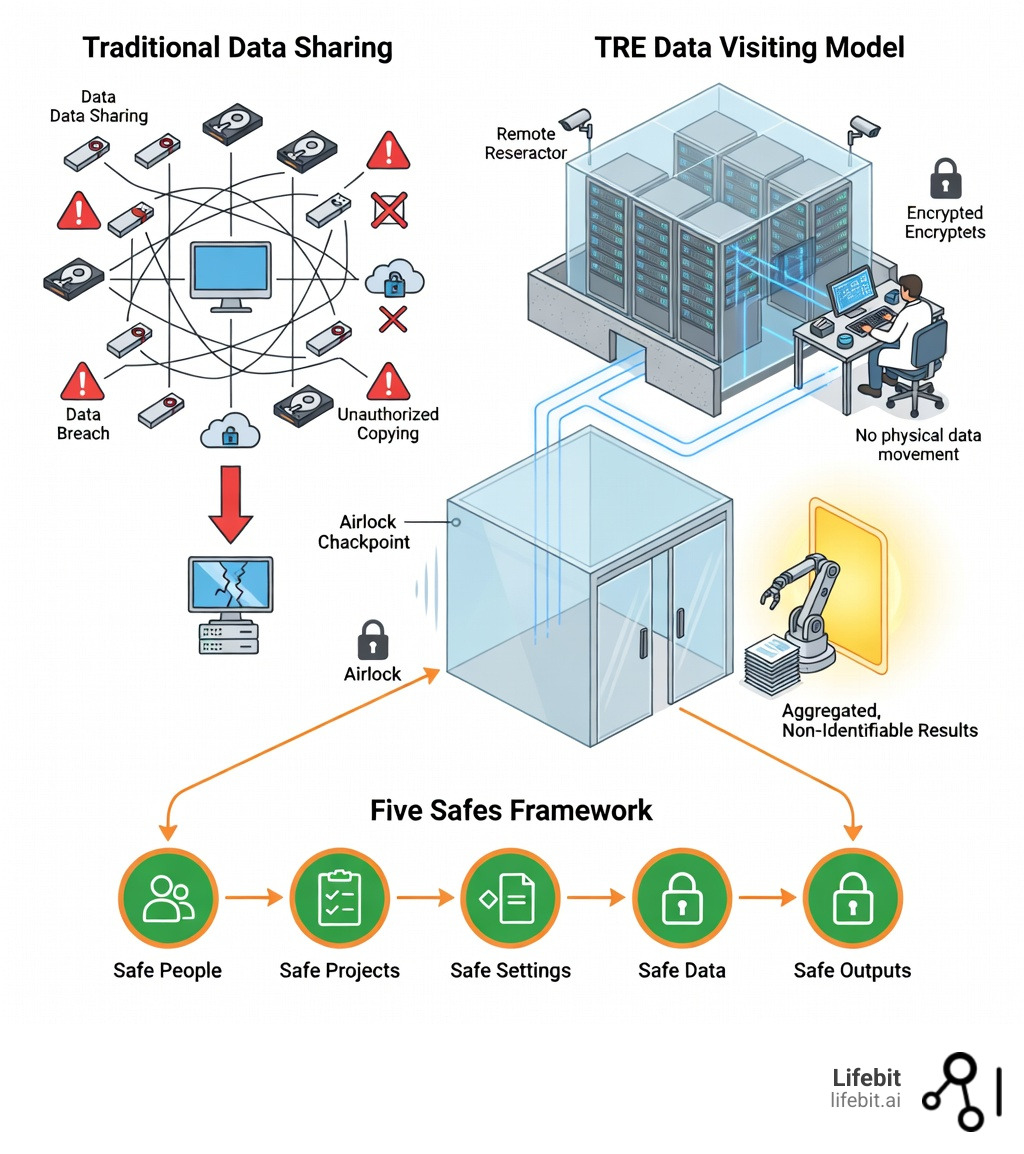
- Five Safes framework – Multi-layered controls covering Safe People (vetting), Safe Projects (approval), Safe Settings (encryption, isolation), Safe Data (de-identification), and Safe Outputs (disclosure review)
- No data egress – Raw patient data never leaves the environment; only aggregated, non-identifiable results pass through strict “airlock” controls
- Compliance-ready – Built to meet GDPR, HIPAA, ISO 27001, and emerging standards like the European Health Data Space (EHDS)
The shift from traditional data sharing—where copies of datasets are distributed on USB drives or via file transfer—to the data visiting model of TREs represents a fundamental change in how we balance innovation with privacy. Instead of moving sensitive data to researchers, TREs bring researchers to the data in a controlled, auditable space.
This matters because the scale of biomedical data is exploding. A single whole genome requires 750MB of storage. The UK Biobank alone exceeds 20 petabytes. Genomics England houses over 135,000 whole genomes. Traditional approaches to data sharing simply cannot scale to meet the computational, security, and sovereignty demands of modern genomic and real-world evidence research.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve built a federated biomedical data platform that powers TRE secure data environment solutions for pharmaceutical companies, public health institutions, and regulatory agencies worldwide. Over the past 15 years working across computational biology, AI, and health-tech entrepreneurship, I’ve seen how secure, federated environments open up life-saving insights without compromising patient trust.
What is a TRE Secure Data Environment?
At its simplest, a TRE secure data environment is a “walled garden” for sensitive information. Think of it like a high-security reference library. In a traditional library, you might check out a book and take it home. In a TRE, the “books” (your sensitive data) never leave the building. You are allowed into a secure room, you can take notes on what you find, but a security guard checks your notes at the door to make sure you haven’t hidden any original pages in your pockets.
This environment is often referred to by various names, including What is a Secure Data Environment (SDE)? or a Data Safe Haven. Regardless of the name, the goal is the same: to provide a space where approved users can access non-identifiable health data for research under strict controls. This approach is governed by the Five Safes framework, which ensures that every aspect of the data access—from the person to the final result—is vetted for safety.
The Evolution of the Data Visiting Model
The shift toward TREs was accelerated by the realization that traditional data sharing—where datasets are copied and sent to researchers—is inherently risky and unscalable. In the past, once a dataset left a hospital’s server, the hospital lost all control over how that data was stored, who accessed it, or whether it was properly deleted after the project ended. The “data visiting” model pioneered by TREs flips this script. Instead of the data moving to the researcher, the researcher moves to the data. This ensures that the data custodian maintains 100% oversight at all times.
Core Components of a TRE Secure Data Environment
To function effectively, these environments rely on several Key Features of a Trusted Research Environments:
- Virtual Research Environment (VRE): A digital workspace where researchers log in to find the tools they need, such as RStudio, Python, or Jupyter Notebooks. These are often hosted on high-performance cloud infrastructure to handle massive genomic files.
- Data Ingress: A secure pathway to bring new datasets or research code into the environment. This process includes scanning for malware and ensuring the code doesn’t contain hidden scripts designed to exfiltrate data.
- Egress Control (The Airlock): A rigorous process where every file a researcher wants to take out of the environment is reviewed by a human to prevent accidental data leaks. This is the final line of defense.
- Network Isolation: The environment is typically disconnected from the open internet to prevent hackers from reaching the data and to stop researchers from accidentally uploading sensitive files to public clouds or social media.
Why Your Research Needs a TRE Secure Data Environment
The old way of sharing data is broken. When 66% of medical researchers cite data sensitivity as their top concern, we know that privacy is the biggest hurdle to innovation. Furthermore, the sheer volume of data makes physical sharing impossible. With a single whole genome taking up 750MB, moving thousands of them across borders is a logistical nightmare.
The independent review by Prof Ben Goldacre for the UK government emphasized that SDEs are the only way to earn and maintain “social license”—the public’s trust that their health data is being used safely. The Goldacre Review specifically recommended that SDEs become the default for all NHS data access, moving away from the “trust and pray” model of data dissemination. By adopting a “data visiting” model, we ensure data sovereignty (the data stays where it belongs) while still allowing global experts to collaborate on solving diseases like cancer or Alzheimer’s.

The Five Safes: A Blueprint for Secure Data Access
How do we actually define “safe”? We don’t just guess. We use the Five Safes, an internationally recognized gold standard for data governance. This framework allows us to enjoy the Advantages of Trusted Research Environments by breaking security down into five manageable layers.
Implementing Safe People and Safe Projects
Trust starts with the human element. Safe People means that only vetted researchers gain entry. Research shows that 85% of TREs mandate specialized training, and 79% require researchers to sign legally binding agreements with clear penalties for misuse. This vetting often includes identity verification and institutional sponsorship, ensuring that the researcher is who they say they are and is affiliated with a reputable organization.
Safe Projects ensures the research is ethical and serves the public good. Data custodians use “data minimization” principles, meaning researchers only get access to the specific data points they need for their approved question—nothing more. If you’re studying heart disease, you don’t need access to dental records. This vetting process often involves an ethics committee or a Data Access Committee (DAC) that reviews the scientific merit of the proposal. You can find more info about secure research environments and how these vetting processes work on our blog.
Ensuring Safe Settings, Data, and Outputs
The technical side of the framework is just as rigorous:
- Safe Settings: This is the “walled garden” itself. We use network isolation, Multi-Factor Authentication (MFA), and Role-Based Access Control (RBAC) to ensure only the right people reach the right data. Modern TREs often utilize “air-gapped” virtual machines that have no connection to the external internet, preventing any unauthorized data transfer.
- Safe Data: Before a researcher even sees a file, it is pseudonymised or de-identified. Names, addresses, and ID numbers are removed or masked. Advanced techniques like k-anonymity or differential privacy may be applied to ensure that even if a researcher tries to cross-reference the data with other public datasets, they cannot re-identify an individual.
- Safe Outputs: This is the most critical step. Before results are published, they go through an Airlock Data Export Trusted Research Environments process. Interestingly, 0% of TRE operators believe software can fully replace human checks here. A human must verify that the “aggregate” data (like a chart showing trends) doesn’t accidentally reveal a specific individual’s identity, particularly in studies involving rare diseases where a single data point could be identifying.
How a TRE Secure Data Environment Works: From Ingestion to Insight
The journey of data through a TRE secure data environment is a carefully choreographed lifecycle designed to maximize utility while minimizing risk. This lifecycle ensures that data is not only secure but also usable for high-quality scientific inquiry.
- Ingestion: Raw, sensitive data is brought into the secure environment via encrypted channels. This often involves data from multiple sources, such as Electronic Health Records (EHR), genomic sequencing labs, and clinical trial databases.
- Harmonization: Data from different sources is cleaned and standardized. This is a critical step; without harmonization (using standards like OMOP CDM or FHIR), researchers cannot easily compare data from different hospitals or countries. This process turns “messy” real-world data into a research-ready asset.
- Secure Analysis: Researchers use advanced tools like RStudio, Jupyter, and SAS to run their models. The TRE provides the high-performance computing (HPC) power necessary to process petabytes of data, which would be impossible on a standard laptop.
- Review: The research outputs are placed in the “Airlock” for disclosure control. This involves checking for “small cell counts”—for example, if a table shows only one patient in a specific category, that table might be blocked from export because it could identify that person.
- Insight: Once cleared, the results are used to inform policy, clinical practice, or drug development. The raw data remains safely behind the firewall.
A great example of this in action is a population-based cohort study of 46 million adults in England. This massive study, which analyzed COVID-19 vaccine safety, was only possible because a TRE allowed researchers to query nearly the entire adult population of a country without ever compromising a single patient’s privacy. The study provided real-time evidence that saved lives during the pandemic.
The Role of Encryption and Auditing
Security isn’t a “set it and forget it” feature. We use end-to-end encryption for data both “at rest” (stored on disks) and “in transit” (moving between servers). Furthermore, Secure Clinical Data requires continuous auditing. Every click, every query, and every file access is logged in an immutable audit trail. If something unusual happens—such as a researcher attempting to run a query that targets a specific individual—an incident response plan kicks in immediately. This level of oversight is far superior to traditional data-sharing methods where, once a file is sent, the sender has no idea what happens to it.
Managing the Airlock and Disclosure Control
The “Airlock” is where the most sophisticated security happens. While 100% of TREs allow the export of aggregate-level data (like “30% of patients responded well”), exporting AI models is much riskier. Only 23% of TREs currently allow AI model exports because of “membership inference attacks”—where a clever hacker could potentially reverse-engineer the model to find out if a specific person’s data was used to train it. We tackle these Airlock Challenges and AI Solution TRE issues by combining human expertise with advanced privacy-enhancing technologies like synthetic data generation for model testing.
Centralized vs. Federated: The Evolution of Secure Architectures
As research grows, the way we build these environments is changing. Historically, TREs were centralized—you moved all your data into one big bucket. But for global research, this creates massive “data silos” and sovereignty issues. If a researcher in the US wants to analyze data from 10 different European hospitals, moving all that data to a single central server is legally and logistically impossible under GDPR.
The Rise of Federated TREs
The Federated Trusted Research Environment model is the future. In a federated setup, the data stays exactly where it was created (e.g., in a hospital in Germany or a lab in Singapore). The researcher sends their analysis to the data, the computation happens locally within each site’s own TRE, and only the non-identifiable results come back to the researcher to be aggregated. This “compute-to-data” approach is the vision driving the European Health Data Space (EHDS), allowing for secure, cross-border collaboration without moving a single byte of raw patient data.
| Feature | Centralized TRE | Federated TRE |
|---|---|---|
| Data Location | Moved to a single central repository | Stays with the original custodian |
| Sovereignty | Challenging for cross-border data | High (data never leaves the country) |
| Scalability | Limited by central storage costs | High (uses distributed computing) |
| Latency | Low (data is all in one place) | Can be higher (network dependent) |
| Security | Single point of failure | Distributed risk profile |
Scaling to Petabyte-Scale Genomic Research
Size matters. 45% of TRE operators admit they struggle to scale their computing and storage for petabyte-scale datasets. When you are dealing with the UK Biobank (20+ petabytes) or Genomics England (135,000+ genomes), you need a platform that doesn’t buckle under the pressure. Our Lifebit Trusted Research Environment Ultimate Guide explores how we use cloud-native architectures and containerization (like Docker and Nextflow) to provide the high compute capacity needed for these massive genomic studies. This allows researchers to run complex GWAS (Genome-Wide Association Studies) across millions of variants in hours rather than weeks.
Future-Proofing with Privacy-Enhancing Technologies (PETs)
The next generation of TREs will be even smarter. We are already integrating Privacy-Enhancing Technologies (PETs) like:
- Confidential Computing: Encrypting data even while it is being processed in the computer’s memory (RAM), protecting it from even the cloud provider’s administrators.
- Federated Learning: Training AI models across multiple locations without sharing data, allowing for more robust algorithms that aren’t biased by a single hospital’s patient population.
- Synthetic Data: Creating “fake” data that looks and acts like real patient data for testing and code development, without any privacy risk. This allows researchers to write and test their code before ever touching the real sensitive data.
These innovations, detailed in our Lifebit TRE Guide 2026, will help automate the airlock process and reduce the time it takes for researchers to get from “question” to “insight.”
Real-World Impact: Success Stories in Healthcare and Genomics
TREs aren’t just a theoretical concept; they are saving lives today. By providing a secure bridge between data owners and data users, these environments have unlocked discoveries that were previously hidden in inaccessible silos.
Genomics England and the 100,000 Genomes Project
One of the most prominent examples is Genomics England. By housing over 135,000 whole genomes within a secure TRE, they have enabled researchers to identify new genetic causes for rare diseases and cancers. For many patients, this has meant finally receiving a diagnosis after years of uncertainty. The TRE ensures that this incredibly sensitive genetic blueprint is never exposed, yet remains accessible to the world’s best scientists.
The UK Biobank and Global Collaboration
The UK Biobank has enabled over 10,000 publications, providing insights into everything from heart disease to rare genetic disorders. By providing a secure cloud-based TRE, the UK Biobank allows researchers from around the world to analyze its 20+ petabytes of data without the data ever leaving the UK. This has democratized access to high-quality data, allowing researchers in smaller institutions to compete with those at major universities.
National Safe Havens and Public Health
These national Safe Haven models prove that when you build a secure environment, researchers can accelerate disease research at an unprecedented pace. For example, the SAIL Databank in Wales has been securely housing 30 years of population health data. This longitudinal data has led to breakthroughs in how we manage chronic conditions like asthma and diabetes by allowing researchers to see how treatments work over decades in the real world.
Governance and Public Trust
The most successful TREs are the ones that talk to the public. Earning a “social license” requires transparency. This means embedding Patient and Public Involvement and Engagement (PPIE) into governance. There are many examples demonstrating how public contributors help decide who gets access to data. By publishing data use registers, TREs show the public exactly who is using their data and why. This transparency is the foundation of the trust required to keep these vital research resources running.
Frequently Asked Questions about TREs
What is the difference between a TRE and an SDE?
Essentially, they are the same thing. “Trusted Research Environment” (TRE) is the term traditionally used by academia and research councils, while “Secure Data Environment” (SDE) is the term increasingly used by the NHS and UK government. Because these systems developed organically, UK Statistics Authority has set out standards to help harmonize the definitions.
How do TREs comply with GDPR and HIPAA?
TREs are designed to meet the highest legal standards. They act as a “Data Processor” under GDPR, providing the technical and organizational measures required to protect data. By using pseudonymisation, strict access controls, and ISO 27001-certified infrastructure, TREs ensure that the “Data Controller” (the hospital or biobank) remains in compliance. You can even deploy a Trusted Research Environment Azure instance that is pre-configured for these regulations.
Can researchers export raw data from a TRE?
No. This is the fundamental rule of the “walled garden.” You can export your code, your charts, and your summary statistics, but the raw, patient-level data stays inside. This is managed through the airlock process we described in our TRES UK Complete Guide.
Conclusion
The future of medicine depends on our ability to analyze vast amounts of sensitive data securely. At Lifebit, we believe that you shouldn’t have to choose between privacy and progress. Our federated AI platform provides a next-generation TRE secure data environment that enables real-time access to global biomedical and multi-omic data.
By bringing the analysis to the data, we empower researchers to collaborate across 5 continents, delivering real-time insights and AI-driven safety surveillance while keeping patient trust at the heart of everything we do. Whether you are in biopharma, government, or public health, we invite you to explore the Lifebit Federated Biomedical Data Platform and see how we can help you turn data into knowledge, safely and responsibly.

