The Ultimate Guide to Integrating Multi-Omic Data Without Losing Your Mind

Why Multi-Omics Integration Is the Biggest Data Challenge in Precision Medicine
A roadmap for multi omics data integration using deep learning starts with understanding that you’re combining fundamentally different data types—genomics with 500,000 features, transcriptomics with 20,000, proteomics with 3,000, and epigenomics with 450,000—all from different sample sizes and measurement platforms. Here’s the essential roadmap:
Core Integration Steps:
- Choose your fusion strategy – Early (concatenate raw features), intermediate (merge learned representations), or late (combine predictions)
- Select your architecture – Non-generative methods (FNNs, GCNs, autoencoders) for complete data, or generative methods (VAEs, GANs, GPTs) for missing data
- Handle missing values – Use product-of-experts frameworks or cross-encoders when modalities are incomplete
- Incorporate biological knowledge – Add PPI networks or pathway constraints to improve interpretability
- Validate rigorously – Test on benchmarks like TCGA or GTEx with proper cross-validation
The reason this matters? Deep learning-based integration achieves AUC-ROC scores of 0.85, outperforming traditional methods like SNF (0.71) and MOFA (0.74). But here’s the catch: most FNN methods require complete data across all modalities for every sample, and only methods like GLUER handle incomplete datasets effectively.
The convergence of systems biology and artificial intelligence has opened new frontiers, but the path forward is littered with challenges—high dimensionality, batch effects, computational intensity, and the dreaded problem of missing data that plagues real-world datasets.
As Maria Chatzou Dunford, CEO and Co-founder of Lifebit with 15 years in computational biology and AI, I’ve built tools that empower precision medicine through better data integration—and I’ve seen how a roadmap for multi omics data integration using deep learning can transform research when implemented correctly. This guide breaks down the technical complexity into practical steps you can actually use.
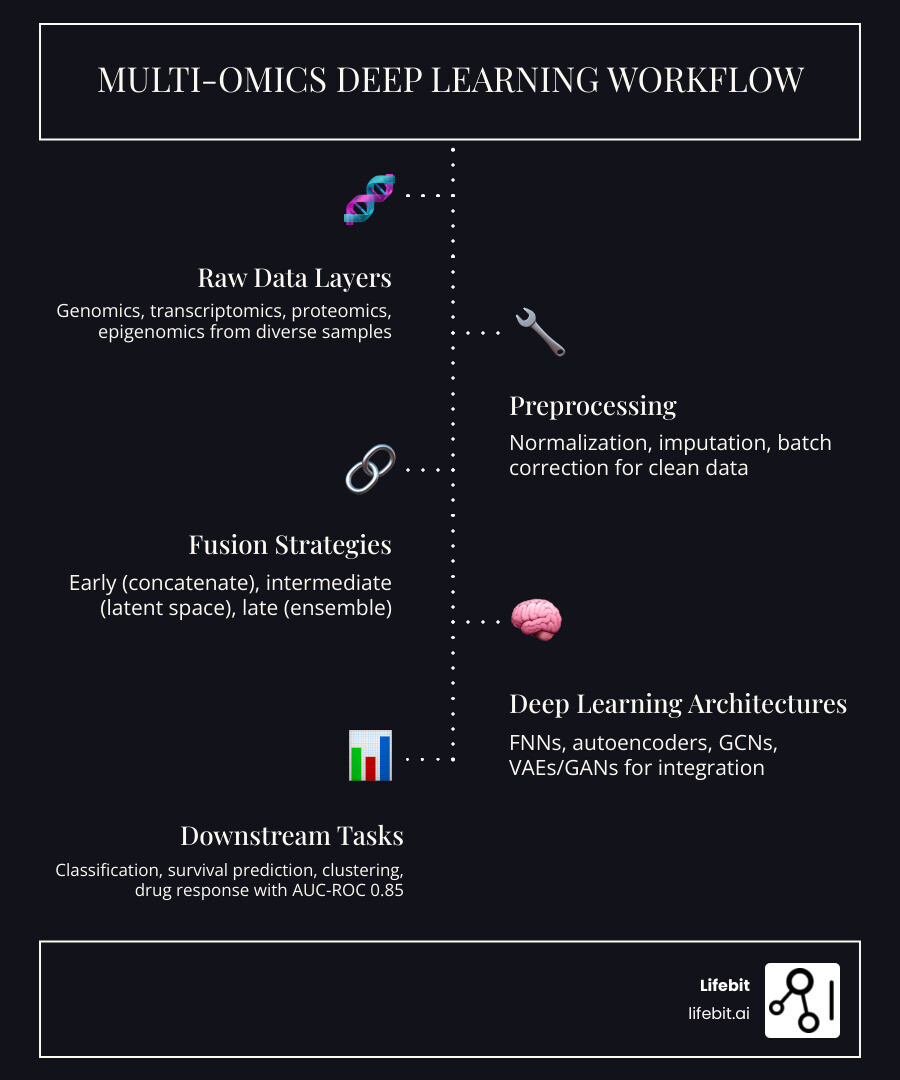
Why Multi-Omics Integration Fails: Overcoming the Data Complexity Barrier
Let’s be honest: if multi-omics integration were easy, we’d have cured everything by now. The reality is that biological data is messy. When we try to combine different “omes,” we aren’t just adding numbers; we are trying to harmonize entirely different languages of life.
The “Curse” of Heterogeneity and High Dimensionality
The first wall we hit is heterogeneity. Genomics data might consist of 500,000 binary SNPs, while transcriptomics gives us 20,000 continuous gene expression values. Mixing these is like trying to make a smoothie out of rocks and water—the textures just don’t match. Furthermore, the statistical distributions vary wildly: RNA-seq data often follows a negative binomial distribution, while proteomics data might be closer to log-normal. Forcing these into a single model without proper normalization leads to “feature dominance,” where the modality with the largest numerical range or highest feature count (usually genomics or epigenomics) drowns out the subtle signals in smaller datasets like metabolomics.
Then there’s the “High-Dimensional, Low-Sample Size” (HDLSS) problem, often referred to as the $p \gg n$ problem. We often have hundreds of thousands of features ($p$) but only a few hundred samples ($n$). In a standard neural network, a simple architecture (1000 input nodes → 100 → 2) requires over 100,000 parameters. Without careful management, the model will simply memorize the noise in your data rather than learning actual biology. This is why data cleaning and feature selection are not just “pre-steps”—they are the foundation of the entire roadmap. Techniques like LASSO (Least Absolute Shrinkage and Selection Operator) or Elastic Net are frequently used to prune the feature space before it ever reaches the deep learning layers.
Batch Effects and Missing Data
Batch effects are the silent killers of reproducibility. If one set of samples was processed in London and another in New York, a deep learning model might “learn” the difference between the lab equipment rather than the difference between healthy and diseased tissue. This is particularly dangerous in deep learning because these models are exceptionally good at finding patterns, even if those patterns are artifacts of the sequencing machine’s calibration. We use techniques like ComBat for batch correction and KNN imputation for missing values to mitigate this. More recently, adversarial training has been used to create “batch-invariant” representations, where a sub-network is specifically trained to fail at identifying which batch a sample came from, ensuring the main model focuses only on biological signal.
Comparison of Fusion Strategies
To steer this, we must choose a fusion strategy. Each has its own strengths and weaknesses:
| Strategy | Approach | Strengths | Weaknesses |
|---|---|---|---|
| Early Fusion | Concatenate all raw data into one giant matrix. | Simple; captures correlations between raw features. | High risk of overfitting; ignores different data distributions. |
| Intermediate Fusion | Map each modality to a shared latent space first. | Best for inter-modality relationships; handles different data types well. | More complex to design; requires careful architecture tuning. |
| Late Fusion | Train separate models and aggregate their predictions. | Preserves modality-specific info; easy to debug. | Misses complex cross-talk between different omics layers. |
Intermediate fusion is currently the most popular in high-impact research because it allows for the creation of a “joint embedding.” This embedding acts as a universal biological signature that captures the flow of information from DNA to phenotype, something that early and late fusion often fail to do effectively.
A Roadmap for Multi Omics Data Integration Using Deep Learning: Architectures and Strategies
When we talk about a roadmap for multi omics data integration using deep learning, we are moving beyond simple linear correlations. Deep learning excels here because it can model the non-linear, hierarchical nature of biology (DNA → RNA → Protein). Traditional methods like Principal Component Analysis (PCA) or Multiple Co-inertia Analysis (MCIA) assume linear relationships, but biological systems are governed by feedback loops and threshold effects that only deep architectures can capture.
The gold standard for these models is achieving an AUC-ROC of 0.85, significantly outperforming traditional statistical methods. But choosing the right “engine” for your roadmap is critical.
Non-Generative vs. Generative Deep Learning Roadmap
We generally categorize deep learning methods into two buckets: non-generative and generative.
Non-Generative Methods (The “Discriminators”):
These methods focus on learning the mapping from input $X$ to outcome $Y$ (calculating $P(Y|X)$).
- Feedforward Neural Networks (FNNs): Great for tabular data. Methods like MOLI use modality-specific encoders to predict drug responses. However, FNNs can be “brittle” when faced with noisy clinical data.
- Graph Convolutional Networks (GCNs): These are the “social butterflies” of the DL world. They use Patient Similarity Networks (PSNs) or Protein-Protein Interaction (PPI) networks to exploit relationships between samples or genes. By treating patients as nodes in a graph, GCNs can leverage the fact that patients with similar molecular profiles should have similar clinical outcomes.
- Autoencoders (AEs): These are essential for dimensionality reduction. They compress high-dimensional omics into a “latent space” while ensuring the most important information is preserved. A variant, the Denoising Autoencoder, is particularly useful for multi-omics as it learns to reconstruct the original data from a corrupted version, making the model robust to experimental noise.
Generative Methods (The “Creators”):
Generative methods like Variational Autoencoders (VAEs) and GANs model the underlying data distribution ($P(X, Y)$). These are the powerhouses for handling missing data because they can “imagine” or impute what the missing modality should look like based on the others. For instance, if a patient has transcriptomic data but lacks proteomics, a VAE trained on a large cohort can estimate the protein levels with surprising accuracy.
The Rise of Transformers in Multi-Omics
While not mentioned as often in older roadmaps, Transformers and self-attention mechanisms are the new frontier. Originally designed for Natural Language Processing, Transformers can treat different omics features as “tokens” in a sentence. This allows the model to learn which specific genes or SNPs are “attending” to each other across different modalities. Models like OmiEmbed and Cross-Omics Transformers are now setting new benchmarks in pan-cancer classification by identifying long-range dependencies between distal enhancers and gene promoters.
Selecting Architectures for a Roadmap for Multi Omics Data Integration Using Deep Learning
The “best” architecture depends entirely on your goal:
- For Classification: GCN-based methods like MOGONET are excellent because they use similarity networks to group similar patients together.
- For Survival Prediction: Models like DeepProg or SALMON are designed to handle censored data, using eigengene modules to reduce noise. These models often employ a Cox Proportional Hazards loss function integrated directly into the neural network.
- For Drug Response: MOLI (Multi-omics Late Integration) is a go-to for predicting how a specific cancer cell line will react to a compound, using triplet loss to ensure that responders and non-responders are clearly separated in the latent space.
Solving the “Missing Data” Problem in Multi-Omics Workflows

In the real world, data is like Swiss cheese—full of holes. Most FNN-based methods (14 out of 15 reviewed in recent studies) require a complete set of modalities for every single patient. If you’re missing even one miRNA expression profile, the patient is out. This is a massive waste of clinical data, especially in rare disease research where every sample is gold.
Handling Incomplete Data with a Roadmap for Multi Omics Data Integration Using Deep Learning
This is where advanced methods like GLUER and generative approaches come in. GLUER uses mutual nearest neighbors to match cells across different modalities, even when the data is unpaired or incomplete. This is particularly useful for single-cell multi-omics, where you might sequence the RNA of one cell and the ATAC-seq of another, but never both from the same physical cell.
Generative models like VAEs use a “Product-of-Experts” (PoE) framework. Imagine each omic modality is an “expert.” If one expert is missing, the model can still make a high-quality joint representation using the experts that are present. This is mathematically superior to simple mean imputation, which tends to “wash out” the variance in the data and lead to overly conservative predictions.
Trans-omics block missing data imputation (ToBMI) is another breakthrough. It uses a weighted approach to fill in missing blocks of data, allowing us to use the full power of datasets like GTEx or TCGA without discarding half our samples. ToBMI leverages the correlation structure between modalities; for example, it uses the methylation state of a promoter to predict the expression level of the downstream gene when that gene’s expression data is missing.
The Role of GANs in Data Augmentation
Beyond imputation, Generative Adversarial Networks (GANs) are being used for data augmentation. In many clinical trials, the sample size is too small for deep learning. Researchers are now using GANs to generate “synthetic patients” that maintain the complex correlations of the original multi-omics data. These synthetic samples can be used to pre-train models, which are then fine-tuned on the real, limited clinical data—a strategy known as transfer learning that significantly boosts model stability.
Beyond Molecular Data: Integrating Imaging and Prior Biological Knowledge
Biology doesn’t happen in a vacuum, and it certainly doesn’t happen only at the molecular level. Emerging methods are now integrating “non-molecular” modalities like radiomics (from MRI/CT scans) and pathomics (from whole-slide tissue images). This is the “macro-to-micro” integration that represents the next evolution of the roadmap.
The Power of Pathomic Fusion
Pathomic Fusion is a framework that uses Kronecker products to model pairwise interactions between histopathology features and genomic markers. This allows us to see how a specific mutation actually changes the physical structure of a tumor. For example, a TP53 mutation might correlate with specific nuclear pleomorphism patterns visible on a slide. By fusing these, the model can predict survival more accurately than if it looked at the mutation or the image in isolation. The Kronecker product is key here because it captures the interaction between every pixel-level feature and every molecular feature, creating a high-dimensional tensor of biological insights.
Single-Cell Multi-Omics: The Resolution Revolution
We cannot discuss a modern roadmap without mentioning single-cell technologies (scRNA-seq, scATAC-seq, CITE-seq). Integrating these is a unique challenge because the data is extremely sparse (many zeros). Deep learning models like scVI (single-cell Variational Inference) use zero-inflated negative binomial (ZINB) distributions to model this sparsity. Integrating single-cell data allows researchers to see which specific cell types (e.g., T-cells vs. B-cells) are driving the multi-omic signals observed in bulk tissue samples, providing a much higher resolution of the disease microenvironment.
Enhancing Model Performance with a Biological Roadmap
One of the biggest criticisms of deep learning is that it’s a “black box.” We can fix this by baking prior biological knowledge directly into the model’s architecture. This is often called “Informed Machine Learning.”
Instead of a fully connected layer where every gene talks to every other gene (which is biologically impossible), we can use sparse neural networks. For example, PASNet uses a layer where nodes represent biological pathways from databases like KEGG or Reactome. Connections only exist if a gene actually belongs to that pathway. This reduces the parameter count from millions to just a few thousand, making the model more interpretable and less likely to overfit.
When we applied these “knowledge-based” constraints, we saw significant enrichment in critical pathways:
- AMPK Signaling: Enrichment score 3.2 (p=0.001) – critical for cellular energy homeostasis.
- mTOR Pathway: Enrichment score 2.9 (p=0.004) – a central regulator of cell growth and proliferation.
- Apoptosis Pathway: Enrichment score 2.8 (p=0.006) – the programmed cell death mechanism often evaded by cancer.
This tells us the model isn’t just guessing; it’s finding the actual biological “circuitry” of the disease. By constraining the AI with what we already know about biology, we ensure that the patterns it finds are clinically relevant and not just statistical noise.
Benchmarking and Future Directions for Clinical Translation
To know if a roadmap for multi omics data integration using deep learning is actually working, we need standard benchmarks. The Cancer Genome Atlas (TCGA) and the Genotype-Tissue Expression (GTEx) project are our primary proving grounds. However, many papers report high accuracy on TCGA but fail when applied to independent hospital cohorts. This “generalization gap” is the biggest hurdle to clinical adoption.
Computational Demands and Optimization
Training these models often requires high-end NVIDIA V100 or A100 GPUs and sophisticated hyperparameter optimization tools like Optuna or Ray Tune. Because the search space for multi-omics (learning rates, dropout rates, layer sizes for 4+ modalities) is so vast, automated machine learning (AutoML) is becoming a standard part of the workflow. Researchers must balance model depth with the risk of “vanishing gradients,” a common problem when trying to backpropagate through multiple modality-specific encoders.
The Future: Exposomics and EHR Integration
The focus is now shifting toward:
- Multi-modal expansion: Integrating Electronic Health Records (EHR), lifestyle factors (diet, exercise), and environmental “exposomics” (pollution, chemical exposure). A patient’s genomic risk for lung cancer is only half the story; their smoking history and zip code complete the picture.
- Interpretability (XAI): Moving from “it works” to “here is why it works.” Using SHAP (SHapley Additive exPlanations) or LIME (Local Interpretable Model-agnostic Explanations) values allows clinicians to see which specific biomarkers led to a high-risk score, fostering trust in the AI’s recommendations.
- Clinical Translation: Moving models out of the lab and into the clinic. This requires “locked” models that can handle the messy, real-time data of a hospital environment without needing constant retraining.
Federated AI: The Game-Changer for Privacy
This is where federated AI becomes the game-changer. In the past, to train a model on 10,000 patients, you had to move all that sensitive data to one central server—a privacy nightmare. At Lifebit, we believe the future of precision medicine isn’t about moving data to the model; it’s about moving the model to the data. Federated learning allows a model to learn from data at a hospital in Germany, then a clinic in Japan, and finally a research center in the US, without the raw data ever leaving its original jurisdiction. This ensures GDPR and HIPAA compliance while still allowing the AI to benefit from a massive, diverse global dataset.
Frequently Asked Questions
What is the best deep learning architecture for multi-omics?
There is no single “best” model, but Autoencoders are generally the most versatile for initial dimensionality reduction, while GCNs are superior for incorporating biological networks like PPIs. For drug response, Late Integration FNNs (like MOLI) are highly effective.
How do you handle batch effects in deep learning models?
We recommend a multi-pronged approach: use ComBat for initial normalization, and then incorporate Batch Normalization layers within your neural network. Additionally, using adversarial training can help the model learn features that are “invariant” to the batch they came from.
Can deep learning integrate imaging with genomic data?
Yes! Architectures like Pathomic Fusion and Deep-Bilinear Networks are specifically designed to fuse the spatial information from whole-slide images with molecular profiles. This “multi-modal” approach often provides a much more accurate prognosis than either data type alone.
Conclusion
Building a roadmap for multi omics data integration using deep learning is a journey from data silos to biological insights. By choosing the right fusion strategy, leveraging generative models for missing data, and grounding our AI in biological reality, we can finally open up the full potential of precision medicine.
At Lifebit, we provide the infrastructure to make this roadmap a reality. Our Trusted Research Environment (TRE) and Federated AI platform enable you to securely access and integrate global multi-omic datasets without the headache of data migration or privacy risks. We help you harmonize heterogeneous data, run advanced ML analytics, and collaborate across 5 continents—all while keeping your data “in place.”
Ready to stop losing your mind over data integration and start finding answers?

