Unlocking the Power of OMOP: A Deep Dive into Data Transformation

OMOP Data Transformation: Stop Manual Wrangling and Scale Research Now
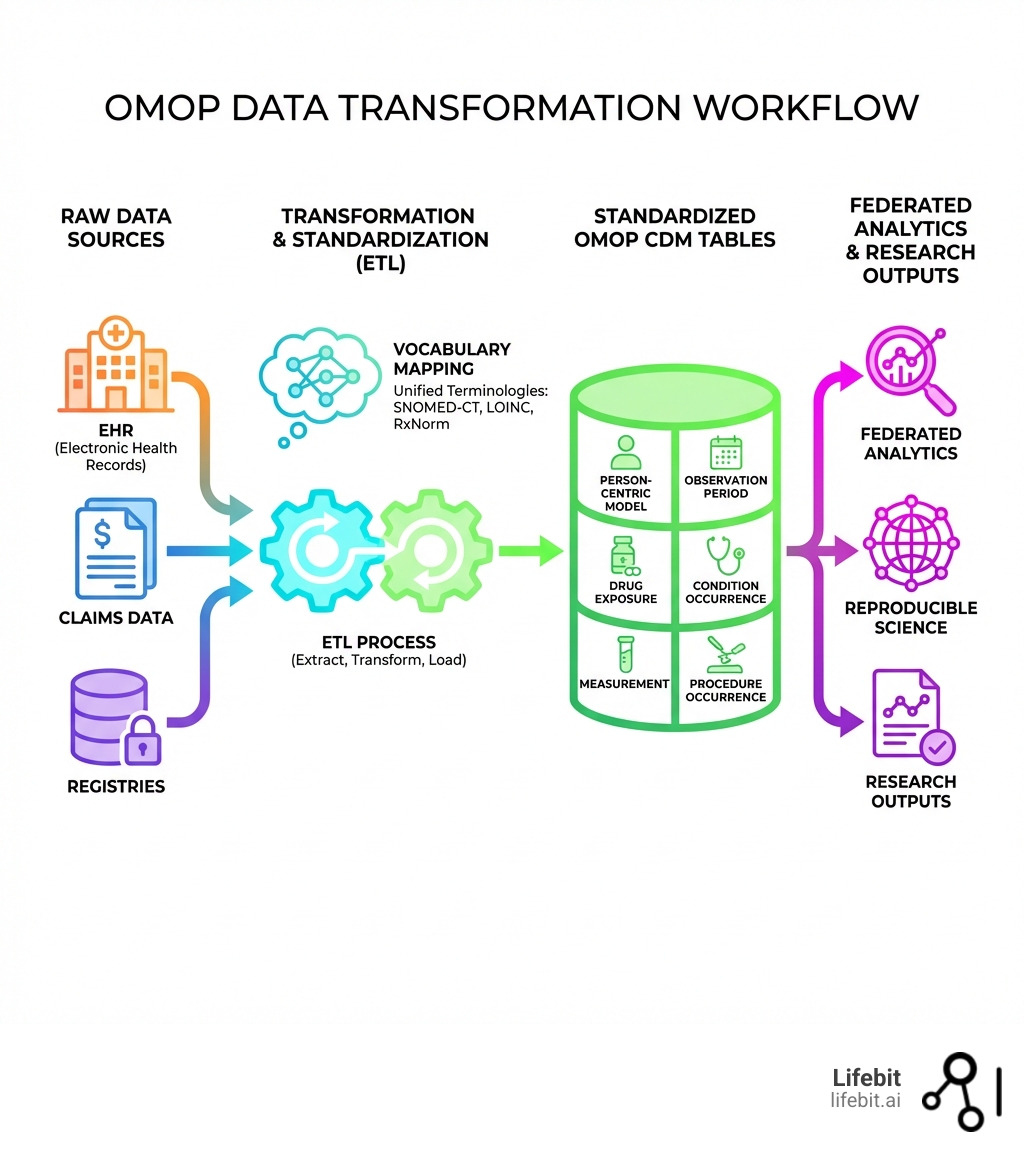
OMOP data transformation is the process of converting diverse healthcare datasets—such as electronic health records (EHR), claims data, and registries—into the standardized OMOP Common Data Model format, enabling large-scale, reproducible research across institutions and geographies.
Key aspects of OMOP data transformation:
- Standardization: Converts disparate data sources into a common format with unified terminologies (SNOMED-CT, LOINC, RxNorm)
- ETL Process: Involves Extract, Transform, Load steps to map source data to OMOP tables
- Person-Centric Model: Structures data around individual patients with required person and date fields
- Quality Assurance: Uses tools like the Data Quality Dashboard to validate transformation accuracy
- Privacy-by-Design: Enables transformation through metadata processing without exposing sensitive data
Healthcare data is notoriously chaotic. Every hospital, clinic, and insurer records patient information differently—using unique codes, formats, and terminologies. This fragmentation creates a “Tower of Babel” that makes collaborative research nearly impossible. When a researcher wants to study treatment outcomes across multiple institutions, they face months of manual data wrangling, custom integrations, and inconsistent results. In fact, industry estimates suggest that data scientists spend up to 80% of their time simply cleaning and preparing data, leaving only 20% for actual analysis and insight generation.
The OMOP Common Data Model (CDM) solves this problem by acting as a universal translator. By transforming raw healthcare data into a standardized format, organizations can run the same analytical queries across datasets from different sources, enabling reproducible science at scale. The transformation process is both technical and strategic—requiring careful vocabulary mapping, data quality checks, and governance frameworks to ensure the converted data remains accurate, complete, and privacy-compliant. This is not merely a database migration; it is a fundamental shift in how clinical knowledge is represented and shared.
Real-world adoption proves the model works. The OHDSI (Observational Health Data Sciences and Informatics) community now represents standardized health data for approximately 800 million patients across 74 countries, with literature on OMOP doubling from 2019 to 2020. Tools like Carrot have converted over 5 million patient records using more than 45,000 transformation rules, while achieving a 97% data quality passing rate. This global network allows for rapid-response studies, such as those seen during the COVID-19 pandemic, where researchers were able to characterize patient populations across continents in a matter of weeks rather than years.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where I’ve spent over 15 years building federated platforms that enable secure OMOP data transformation across pharmaceutical, public sector, and regulatory organizations. Our work powers data-driven drug discovery and real-world evidence generation across compliant, privacy-preserving environments. We believe that by standardizing the world’s health data, we can accelerate the delivery of life-saving treatments to patients who need them most.
OMOP data transformation glossary:
Why Your EHR Data is Useless Without OMOP Data Transformation
The Observational Medical Outcomes Partnership (OMOP) Common Data Model (CDM) is the foundation of modern, collaborative health research. Managed by the Observational Health Data Sciences and Informatics (OHDSI) initiative, the OMOP CDM provides a standardized structure for storing and analyzing disparate observational databases. Without this standardization, EHR data remains trapped in proprietary silos, limited by the specific software version or local coding practices of the institution that collected it.
At its heart, OMOP is a person-centric relational model. This means that every record is linked to an individual patient, and every clinical event is anchored in time. For a record to exist in OMOP, it must, at a minimum, include a person_id and a date. This logical structure ensures that a patient’s story is told chronologically, moving from demographics to visits, and then to specific clinical events like diagnoses, drug exposures, and procedures. This longitudinal view is critical for understanding disease progression and the long-term effects of medical interventions.
The latest OMOP Common Data Model version v5.4 continues this tradition of excellence, offering a robust framework that accommodates both administrative claims and EHR data. It includes specialized tables for oncology, genomic data, and medical devices, ensuring that the model evolves alongside medical technology. By adopting this model, organizations can move away from siloed, proprietary formats and join a global network of researchers. For those new to the field, we offer a comprehensive OMOP guide to help you steer these basics.
Solving the Heterogeneity of EHR and Claims Data
One of the biggest problems in healthcare is data heterogeneity. Electronic Health Records (EHR) are designed for clinical care at the point of service, focusing on real-time documentation and billing. Administrative claims data, on the other hand, are built for insurance reimbursement and often lack the clinical granularity of EHRs. These two worlds speak different languages. A “blood glucose” measurement might be recorded as a LOINC code in one system, a local proprietary string in another, or simply as a line item on a bill.
OMOP data transformation bridges this gap by mapping these varied representations into a common format and concept. This process is essential because, without it, large-scale analytics are impossible. For example, identifying all patients with “Type 2 Diabetes” across ten different hospitals would require ten different queries if the data weren’t standardized. With OMOP, a single query using the standard concept ID for Type 2 Diabetes (SNOMED code 44054006) will return results from all ten sites. We’ve detailed the importance of this in our whitepaper on clinical data harmonisation and AI readiness, which explains how standardized data acts as the fuel for advanced machine learning models.
Aligning with FAIR Data Principles
The OMOP CDM is more than just a database schema; it is a vehicle for the FAIR principles: Findable, Accessible, Interoperable, and Reusable.
- Findable: Standardized metadata allows researchers to find relevant datasets across a federated network. By using common vocabularies, data catalogs become searchable across institutional boundaries.
- Accessible: By using common protocols, data owners can grant secure access without moving sensitive information. Access is governed by standardized roles and permissions that align with the CDM structure.
- Interoperable: This is OMOP’s superpower. Because the data structure and vocabularies are identical across sites, a query written in New York can run perfectly on a dataset in London or Singapore. This eliminates the need for site-specific code adjustments.
- Reusable: Once transformed, data can be used for multiple studies—from safety surveillance to comparative effectiveness—without needing to be rebuilt from scratch. This dramatically reduces the marginal cost of subsequent research projects.
Master OMOP Data Transformation: A 3-Step ETL Guide to Error-Free Mapping
The transition from raw data to a research-ready OMOP dataset happens through a process called ETL: Extract, Transform, and Load. This is where the “art and science” of data engineering meet. It requires a deep understanding of both the source data’s quirks and the target CDM’s strict requirements.
| Step | Action | Tools Often Used |
|---|---|---|
| Extract | Pull raw data from source systems (SQL, NoSQL, CSV) | White Rabbit, Airflow |
| Transform | Map source fields to OMOP tables and local codes to Standard Concepts | Usagi, Rabbit-in-a-Hat |
| Load | Execute SQL scripts to move data into the CDM schema | Carrot-CDM, SQL Compilers, Spark |
The first critical step is metadata profiling. You cannot map what you do not understand. Profiling involves scanning your source data to identify table structures, column types, and the frequency of values. This step often reveals “data debt”—such as dates in the future, misspelled gender entries, or impossible lab values. For a deeper look at how this fits into a broader pipeline, see our guide on how a Data Factory maps to OMOP.
Prioritizing the Person and Visit Tables
In a successful omop data transformation, there is a logical order of operations. You must build the foundation before the walls. This is due to the relational nature of the CDM, where foreign key constraints ensure data integrity.
- PERSON Table: This is always created first. It establishes the unique identifiers for every patient in the dataset. It includes demographic information like year of birth, gender, and race. Without a record here, no other data for that patient can be loaded.
- VISIT_OCCURRENCE Table: This must be created before clinical event tables. It provides the “where and when” context for every diagnosis or medication. Was the drug administered during an inpatient stay or an outpatient visit? This context is vital for health economics and outcomes research (HEOR).
- OBSERVATION_PERIOD Table: This defines the span of time during which we can reasonably expect to see data for a patient. It is essential for defining the “at-risk” period in epidemiological studies. If a patient has no records for five years, we cannot assume they were healthy; they may have simply moved to a different healthcare system.
Failing to follow this order can lead to orphaned records—clinical events that aren’t linked to a person or a visit—making the data useless for longitudinal analysis. Recent scientific research on OMOP ETL Frameworks emphasizes that these dependencies are critical for maintaining the “provenance” or history of the data, allowing researchers to trace a standardized record back to its raw source.
Mapping Vocabularies for OMOP Data Transformation
Vocabularies are the engine of OMOP. The model uses “Standard Concepts” to ensure everyone means the same thing when they say “Type 2 Diabetes.” This process involves mapping local “source values” (like a hospital’s internal code for a blood test) to a globally recognized concept ID.
- SNOMED-CT: The primary vocabulary used for conditions, procedures, and observations. It provides a hierarchical structure that allows researchers to search for broad categories (e.g., “all respiratory infections”) or specific diagnoses.
- LOINC: The standard for laboratory tests and measurements. It captures the component measured, the property, the timing, and the system (e.g., blood vs. urine).
- RxNorm: The go-to for medications and drug exposures. It normalizes drug names and dosages, making it possible to compare medication use across different countries that may use different brand names for the same active ingredient.
The Standardized Vocabularies are managed through the Athena Data Set, which contains millions of concepts and their hierarchical relationships. During the transformation, your local “source values” are mapped to these “concept IDs.” This ensures that a researcher doesn’t have to know your local hospital’s internal coding system to find all patients with a specific condition, effectively removing the language barrier in clinical data.
Cut Mapping Time by 75% With These OMOP Data Transformation Tools
The OHDSI community provides a suite of open-source tools that make omop data transformation manageable, even for large, complex datasets. These tools are designed to handle the specific challenges of healthcare data, from messy terminologies to complex temporal relationships.
- White Rabbit: Scans your source database to produce a “Scan Report” (metadata profile). It identifies which tables and columns contain the most data and highlights potential quality issues early in the process.
- Rabbit-in-a-Hat: Uses the Scan Report to create a graphical mapping between source tables and OMOP tables. It allows data engineers to visualize the flow of data and document the logic behind each transformation.
- Usagi: Helps map local codes to Standard Concepts using term similarity algorithms. It is particularly useful for mapping local laboratory strings or custom medication lists that don’t use standard codes.
- Achilles: Provides data characterization and visualization of the transformed CDM. It generates a dashboard that allows users to explore the demographics, condition prevalence, and drug exposures in their standardized dataset.
- Data Quality Dashboard (DQD): Runs thousands of automated checks to ensure the transformation is accurate. It is the final gatekeeper before data is released for research.
Automating Mappings with Usagi and Carrot
Manual mapping is slow, expensive, and prone to human error. Modern tools now automate much of this heavy lifting using machine learning and rule-based engines. Usagi uses term similarity (such as Levenshtein distance) to suggest mappings, providing a “Match Score” to help curators prioritize their review. This allows experts to focus on the 20% of complex clinical terms that require human judgment, rather than the 80% of straightforward mappings.
Even more advanced is Carrot, a tool developed to handle sensitive data through a privacy-by-design approach. Carrot has been used to convert over 5 million patient records and has created more than 45,000 mapping rules. One of its most powerful features is rule reuse: once a mapping for “Gender: M” to “Male (Concept ID 8507)” is verified, it can be automatically applied to future datasets. This creates a “network effect” where the transformation process becomes faster and more accurate with every new dataset processed. In fact, research shows that over 75% of mappings can be automatically generated from existing vocabularies and previous projects. You can find more details in our OMOP complete guide.
Best Practices for OMOP Data Transformation Projects
While tools provide the “how,” best practices provide the “why.” To achieve a 97% quality passing rate, keep these principles in mind:
- Human Oversight: Automation is a head start, not a finish line. Always have a domain expert (such as a clinician or pharmacist) review mappings, especially for complex clinical terms where subtle differences in meaning can impact research results.
- Incremental Updates: Healthcare data isn’t static. Your ETL process must handle new data arriving daily or weekly without rebuilding the entire database. This requires a robust “delta loading” strategy that only processes new or changed records.
- Provenance Tracking: Maintain a clear link between the transformed record and its original source value. This is vital for auditing, troubleshooting, and “drilling down” into the raw data when an anomaly is found in the standardized results.
- Technical Debt: Avoid “spaghetti queries” in your SQL scripts. Use structured frameworks like YAML-based DML or modular SQL to keep your mapping logic readable, maintainable, and version-controlled.
Achieve 97% Data Quality: Secure OMOP Data Transformation Without Privacy Risks
When dealing with sensitive health information, privacy isn’t just a feature—it’s a legal and ethical requirement. OMOP data transformation must align with strict regulations like GDPR in Europe, HIPAA in the US, and the UK Data Protection Act 2018. The challenge is how to perform complex data engineering without exposing patient identities to the engineers or researchers involved.
A “Privacy-by-Design” workflow involves processing metadata only. By using tools like White Rabbit to extract table structures and value frequencies rather than patient names or social security numbers, external experts can design the transformation logic without ever seeing the raw, sensitive data. The actual transformation (the “Load” step) then happens within the data partner’s secure environment, often behind a firewall. This “federated” approach ensures that sensitive data never leaves its home institution, significantly reducing the risk of data breaches.
Leveraging the Data Quality Dashboard (DQD)
The OHDSI Data Quality Dashboard (DQD) is the gold standard for quality assurance in the OMOP community. It automates thousands of checks across three critical categories, providing a transparent report on the “health” of the dataset:
- Conformance: Does the data follow the OMOP schema rules? This includes checking for correct data types (e.g., ensuring a date field doesn’t contain text), verifying primary key uniqueness, and ensuring that all foreign keys point to valid records in parent tables.
- Completeness: Are there missing values where there shouldn’t be? For example, every record in the
DRUG_EXPOSUREtable should ideally have a start date. The DQD identifies columns with high rates of null values that might indicate a failure in the ETL process. - Plausibility: Does the data make sense in a clinical context? This includes temporal checks (e.g., a patient cannot have a doctor’s visit before they were born or after they died) and value checks (e.g., a human body temperature of 150 degrees Fahrenheit is implausible).
By setting strict thresholds for these checks, organizations can ensure their transformed data is “research-grade.” A typical high-quality OMOP dataset demonstrates a 97% overall passing rate on these checks, giving researchers the confidence that their findings are based on solid, reliable data.
The Role of Federated Governance in Data Sharing
In the era of global collaboration, moving large volumes of sensitive data to a central location is often too risky, expensive, or legally prohibited. This is where federated governance comes in. Instead of centralizing data, we bring the analysis to the data.
Through Trusted Research Environments (TREs) and platforms like Lifebit, researchers can run queries across multiple standardized OMOP databases simultaneously. The sensitive data remains behind the data owner’s firewall, while only the aggregated, non-identifiable results (such as “how many patients in this cohort took Drug X?”) are shared. This enables secure collaboration across public health agencies, academic institutions, and biopharma companies without compromising patient privacy. It is the only way to achieve the scale required for studying rare diseases or long-term drug safety across diverse populations.
OMOP Data Transformation: Answers to the 3 Biggest Conversion Hurdles
What are the primary challenges in OMOP conversion?
The biggest hurdle is almost always vocabulary mapping. Translating thousands of local, proprietary codes into standard OHDSI concepts requires both clinical expertise and technical skill. Many institutions use “homegrown” codes for labs or medications that have no direct equivalent in standard vocabularies. The second challenge is the quality of source data, which often contains inconsistencies, missing values, or illogical entries (like a male patient with a pregnancy diagnosis) that must be identified and “cleaned” or excluded during the ETL process.
How does OMOP handle non-standard local vocabularies?
If a local term has no direct match in the standard vocabularies, it is mapped to Concept ID 0 (Non-standard). However, the OMOP CDM is designed so that the original “source value” is always preserved in a dedicated column (e.g., drug_source_value). This ensures no information is lost. Curators can revisit these mappings as vocabularies evolve or as new mapping rules are developed. This preservation of source data is a key feature that allows for the “provenance” and auditability of the transformed dataset.
Why is the Person table created first in ETL?
The PERSON table is the “parent” of almost every other table in the OMOP CDM. Because of referential integrity (a database concept that ensures relationships between tables remain consistent), you cannot add a diagnosis or a medication record for a patient who doesn’t yet exist in the PERSON table. Establishing the patient identity first ensures that all subsequent clinical events are correctly linked. This structure prevents “orphan records” and ensures that every piece of clinical data can be traced back to a specific individual’s longitudinal record, which is the core requirement for person-centric research.
Is OMOP better than FHIR for research?
OMOP and FHIR (Fast Healthcare Interoperability Resources) serve different purposes. FHIR is designed for the exchange of individual patient records in real-time (e.g., sharing a patient’s chart between two doctors). OMOP is designed for large-scale analytics and population-level research. While FHIR is excellent for clinical operations, its nested structure makes it difficult to run complex queries across millions of patients. Most modern data architectures use FHIR for data transport and OMOP for the final research database, treating them as complementary rather than competitive standards.
Scale Your Research: Start Your OMOP Data Transformation Today
OMOP data transformation is the key to unlocking the true value of healthcare data. By turning fragmented, chaotic records into a standardized, interoperable format, we enable the kind of large-scale research that leads to better treatments, faster drug discovery, and improved patient outcomes. The journey from raw data to a research-ready CDM is complex, but the rewards—in terms of scientific reproducibility and global collaboration—are immense.
At Lifebit, we specialize in making this complex journey simple, secure, and scalable. Our next-generation federated AI platform provides the tools you need for automated harmonization, real-time insights, and compliant data sharing. We help you navigate the technical hurdles of ETL, the clinical nuances of vocabulary mapping, and the legal requirements of data privacy. Whether you are a biopharma company looking for real-world evidence to support a new drug application or a government agency managing national health data for public safety, we are here to help you power your research with the strength of OMOP.
Get started with Lifebit today to see how our Trusted Data Lakehouse can transform your research capabilities and help you join the global community of OHDSI researchers.

