A Guide to Next-Generation Sequencing Technology.

The Dawn of High-Throughput Genomics
Next generation sequencing platform technology has revolutionized how we decode life’s genetic blueprint. These powerful systems can sequence millions of DNA fragments in parallel, making it possible to sequence everything from specific targeted regions to the entire human genome in a single day.
Key NGS Platform Categories:
- Benchtop sequencers – Ideal for small-scale studies, targeted panels, and single-cell analysis
- Production-scale sequencers – Built for large genome projects, population studies, and high-volume clinical testing
- Specialized platforms – Designed for specific applications like long-read sequencing or real-time analysis
The market is diverse, with various manufacturers offering a range of instruments custom to different research needs, from small-scale benchtop systems to large, production-scale sequencers.
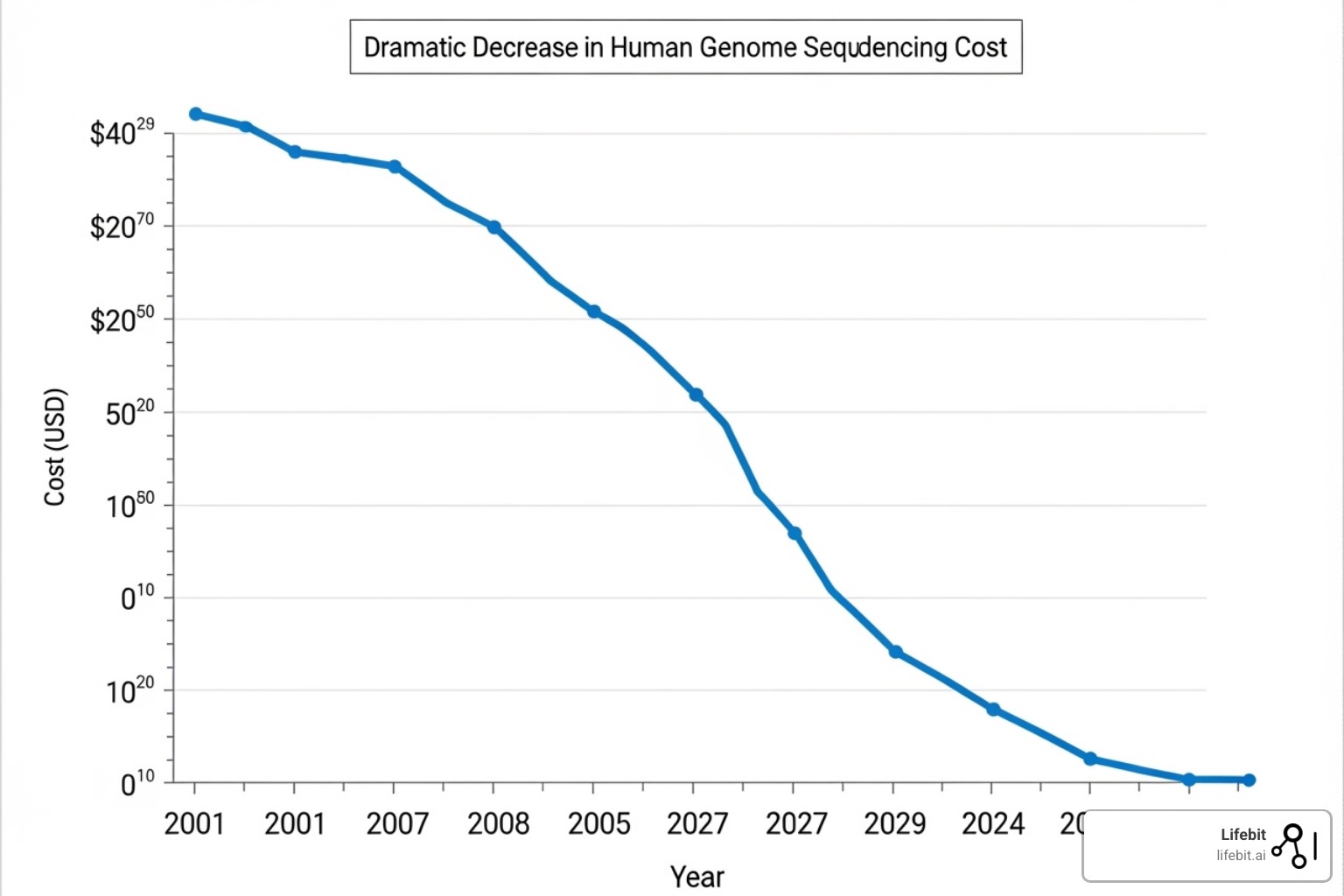
The impact has been staggering. The Human Genome Project took over 10 years and cost nearly $3 billion using traditional Sanger sequencing. Today’s next generation sequencing platform systems make large-scale whole-genome sequencing accessible and practical for the average researcher at a fraction of that cost and time.
This change didn’t happen overnight. NGS technology emerged from the critical need for faster, more accurate, and cost-effective DNA sequencing methods. The technology offers significant advantages over first-generation methods: improved sensitivity and coverage, cost effectiveness, and efficient workflow with faster turnaround times.
Modern NGS platforms can generate data outputs ranging from 300 kilobases up to multiple terabases in a single run, depending on the instrument type and configuration. Some systems can complete sequencing runs in as little as four hours, while others can process up to 6 TB of data daily for comprehensive genomic studies.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit, with over 15 years of experience in computational biology and genomics technology development. My work focuses on building next generation sequencing platform solutions that enable secure, federated analysis of genomic data across global healthcare organizations.
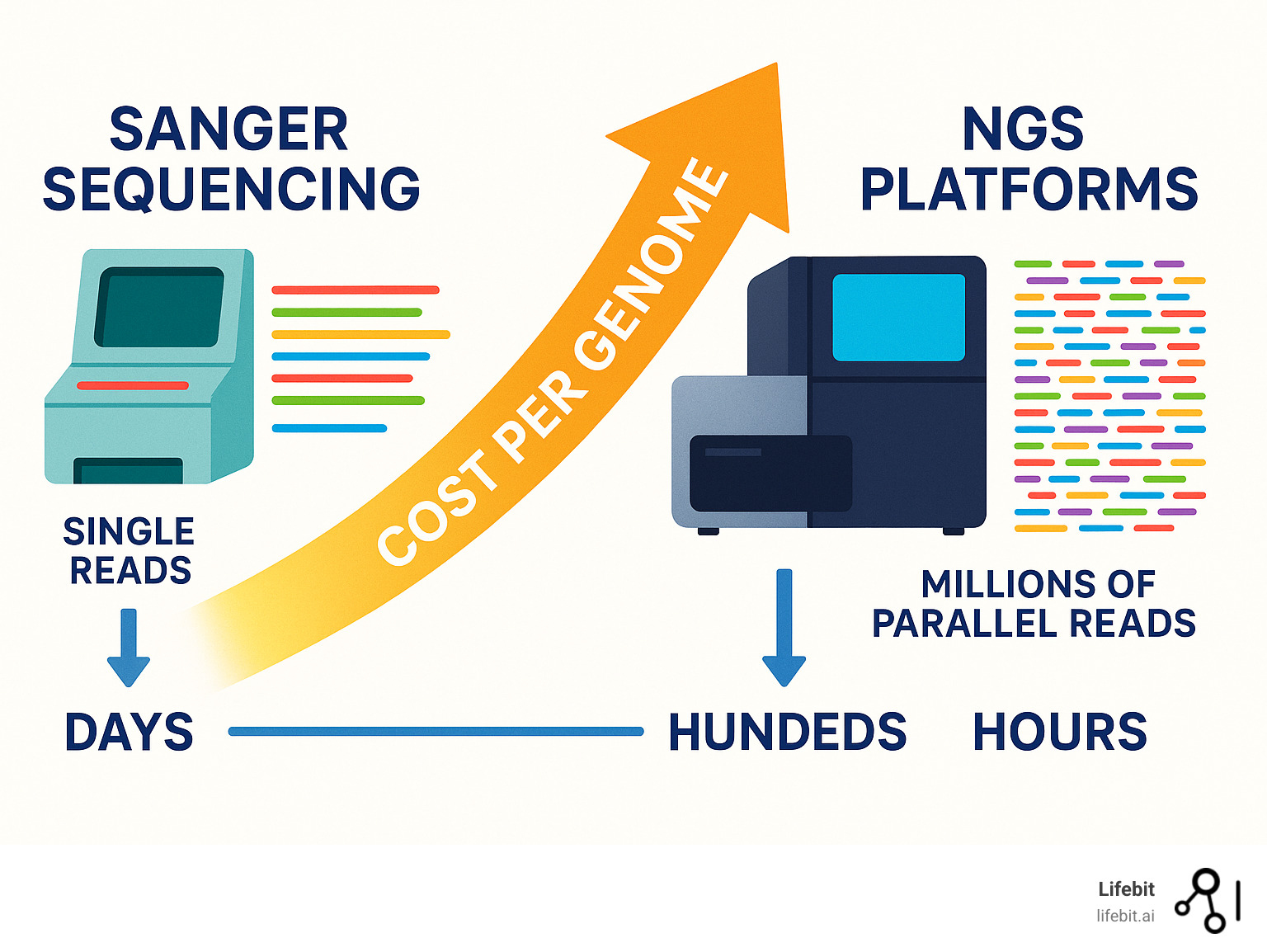
The Evolution of DNA Sequencing Technologies
Think of DNA sequencing like photography. First, we had film cameras that took one careful shot at a time. Then digital cameras let us snap hundreds of photos quickly. Now we have smartphones that can capture, edit, and share images instantly. That’s essentially what happened with DNA sequencing technology.
The story begins in the 1970s with first-generation sequencing. Frederick Sanger developed what became known as the Sanger method – a brilliant technique that used chain-termination to read DNA sequences one base at a time. Picture a DNA polymerase enzyme adding bases to a growing DNA chain until it hits a special “stop” base that ends the process. By running these different-length fragments through gel electrophoresis, scientists could read the sequence like a ladder.
This method was the workhorse of the Human Genome Project, but it was painfully slow. Sequencing a single human genome took over a decade and cost nearly $3 billion. Each sequence had to be read individually, making large-scale studies nearly impossible for most researchers.
Everything changed in the mid-2000s with second-generation sequencing – the birth of true Next-Generation Sequencing platforms. The first of these platforms was launched in 2005, followed by a rapid succession of other systems. These systems introduced massively parallel sequencing – imagine going from taking one photo at a time to suddenly having millions of cameras all clicking simultaneously.
The cost reduction was dramatic. What once cost billions now cost thousands. Speed increased exponentially too – projects that took years could be completed in days. The improved sensitivity meant researchers could detect genetic variations that were previously invisible. For a deeper dive into these methods, check out DNA Sequencing Methods.
Third-generation sequencing arrived to solve a different problem. While second-generation platforms were fast and cheap, they produced short reads that struggled with complex genomic regions. Innovators in the field developed single-molecule sequencing that could read incredibly long DNA fragments – sometimes spanning entire genes in one go.
These longer reads are like having a wide-angle lens instead of looking through a keyhole. They can span repetitive regions and structural variants that shorter reads simply can’t resolve. The trade-off is often higher error rates per individual base, but the long reads provide context that makes the overall picture much clearer.
This rapid evolution has been driven by one simple truth: researchers need more genetic information, faster and cheaper than ever before. The impact on research has been transformative, as detailed in this scientific research on NGS platforms and applications.
Today’s next generation sequencing platform technologies have made whole-genome sequencing accessible to researchers worldwide. What once required massive government funding can now be done in university labs, opening up entirely new fields of study and bringing personalized medicine within reach.
Deconstructing the NGS Workflow: From Sample to Sequence
The fundamental NGS workflow involves three stages: template preparation, sequencing and imaging, and data analysis. Understanding these steps is key to appreciating the technology’s power.
Template Preparation
This stage converts the biological sample (DNA or RNA) into a format ready for sequencing through several key steps:
- Nucleic Acid Extraction: DNA or RNA is extracted from the sample. High quality and quantity are paramount for success.
- Fragmentation: Long DNA or RNA molecules are fragmented into smaller, manageable pieces using methods like enzymatic digestion, sonication, or nebulization. Fragment size depends on the specific next generation sequencing platform and application.
- Library Preparation: Adaptors (short, known DNA sequences) are ligated to both ends of the fragmented DNA. These are crucial as they allow fragments to bind to the flow cell or beads, provide primer binding sites for amplification, and often contain unique barcodes (molecular identifiers) for multiplexing—pooling multiple samples in a single run.
- Amplification: The library fragments are amplified to generate a sufficient signal for sequencing. Common methods include Polymerase Chain Reaction (PCR), emulsion PCR (ePCR), and bridge amplification, which creates template clusters on a flow cell or microbeads.
Sequencing and Imaging
Once the library is prepared and amplified, it’s loaded onto the next generation sequencing platform for the actual sequencing process. Different platforms employ unique biochemical and detection methods:
- Sequencing by Synthesis (SBS): This is the predominant method used by many platforms. It involves adding fluorescently labeled reversible terminator nucleotides one at a time. After each nucleotide is incorporated, a camera captures the fluorescent signal, and the terminator is cleaved, allowing the next cycle to begin. This process generates millions of parallel reads simultaneously.
- Semiconductor Sequencing: This method detects changes in pH. When a nucleotide is incorporated into a growing DNA strand, a hydrogen ion is released. A semiconductor sequencer, for example, detects this change in pH using a semiconductor chip, directly converting chemical information into a digital signal. This allows for a fast and simple workflow without the need for optical detection.
During the sequencing reaction, the instrument captures raw data, typically in the form of images or electrical signals, which are then converted into base calls (A, T, C, G).
Data Analysis
The sequencing process generates an enormous amount of raw data, often in the terabytes range. This “data deluge” necessitates sophisticated bioinformatics pipelines to process, analyze, and interpret the information.
- Quality Control (QC): Raw reads undergo initial QC to assess their quality, remove low-quality bases, and trim adapter sequences.
- Alignment/Mapping: The cleaned reads are then aligned or mapped to a known reference genome (if available). This step identifies where each short read originated in the genome.
- Variant Calling: Once mapped, specialized algorithms identify variations (e.g., Single Nucleotide Polymorphisms (SNPs), insertions, deletions, structural variants) between the sequenced sample and the reference genome.
- Annotation and Interpretation: The identified variants are then annotated with information from various databases to determine their potential functional impact, clinical significance, or association with diseases.
- Downstream Analysis: Depending on the application, further analyses might include gene expression quantification (for RNA-Seq), microbial community profiling (for metagenomics), or epigenetic modification analysis (for ChIP-Seq).
This intricate data analysis requires powerful computational resources and expertise in bioinformatics. We often leverage advanced tools and standardized workflows, such as those found in Nextflow DeepVariant Tutorial and NF-Core Standardised Bioinformatics Pipelines, to manage and accelerate these complex processes.
Understanding the Modern Next-Generation Sequencing Platform
A modern next generation sequencing platform is like a molecular assembly line, running millions to billions of reactions simultaneously to reveal genetic secrets.

This is achieved through different sequencing chemistries. Sequencing by synthesis is the workhorse of many platforms. Fluorescently labeled nucleotides are added one by one to growing DNA strands. A camera captures the unique color of each base as it’s added. The fluorescent tags are then removed, and the cycle repeats hundreds of times to build the complete sequence.
Semiconductor sequencing uses a different approach. It detects the chemical changes that occur as DNA grows. When a nucleotide joins the chain, it releases a hydrogen ion, changing the local pH. Sensitive chips detect this pH change and convert it directly into a digital signal, eliminating the need for cameras or fluorescence.
Key Specifications of a Next-Generation Sequencing Platform
When you’re evaluating a next generation sequencing platform, several critical specifications determine whether it’s right for your research goals.
Data output is perhaps the most obvious metric – how much genetic information can the system generate in one run? Modern platforms range from producing 300 kilobases for focused studies up to multiple terabases for massive genome projects. High-end production-scale sequencers, for example, can deliver over 16 TB of data in a single run.
Quality scores accompany every single base call, giving you confidence in your results. These scores tell you the probability that each detected nucleotide is correct, helping you distinguish reliable data from potential errors.
Read length matters tremendously for different applications. Short reads of 50-150 base pairs work beautifully for gene expression studies, while longer reads extending into kilobases or even megabases are essential for assembling complex genomes or detecting large structural changes.
Paired-end reads provide valuable context by sequencing both ends of each DNA fragment. This gives you information about fragment size and orientation, making it easier to accurately map reads and spot structural variations.
The turnaround time encompasses everything from sample preparation through final results. Some benchtop systems can complete runs in just four hours, while comprehensive whole-genome projects might take several days. Some systems aim to automate the entire workflow, dramatically reducing hands-on time.
Scalability and cost per gigabase round out the key considerations. Can the platform grow with your research needs? And crucially, what’s the economic reality of generating each gigabase of data?
Choosing the Right Next-Generation Sequencing Platform for Your Research
Selecting the perfect next generation sequencing platform feels a bit like choosing the right vehicle for a journey. A sports car won’t help you move furniture, and a moving truck isn’t ideal for a quick trip to the grocery store.
| Application | Typical Throughput | Read Length | Key Advantage |
|---|---|---|---|
| Whole Genome Sequencing (WGS) | Very High (Gb – Tb) | Short to Long | Comprehensive genomic view |
| Targeted Sequencing / Gene Panels | Low to Medium (Mb – Gb) | Short | Cost-effective, high depth on specific regions |
| RNA Sequencing (RNA-Seq) | Medium to High (Gb) | Short | Measures gene expression levels |
Your application needs should drive every decision. Whole genome sequencing demands massive throughput and substantial computational resources. Targeted gene panels require precision and cost-effectiveness. RNA sequencing needs sensitivity to detect lowly expressed genes. Each application has its sweet spot.
Throughput requirements depend entirely on your research scale. Are you running a pilot study with a dozen samples, or managing a population study with thousands of participants? Your sample volume directly impacts which platform makes economic sense.
Budget considerations extend far beyond the initial instrument purchase. Factor in ongoing costs for reagents, consumables, maintenance contracts, and the computational infrastructure needed to analyze your data. It’s like buying a house – the purchase price is just the beginning.
Your data analysis infrastructure capabilities are absolutely critical. Generating terabytes of genomic data is exciting, but only if you can process and interpret it effectively. This is where platforms like ours at Lifebit become invaluable, providing the computational power and analytical tools needed to transform raw sequencing data into meaningful biological insights. Learn more about our approach to Genomics and how we help researchers maximize the value of their NGS investments.
The beauty of today’s next generation sequencing platform landscape is its diversity. Whether you need the raw power of production-scale sequencers or the convenience of benchtop systems, there’s likely a platform perfectly suited to your research goals and budget constraints.
The Broad Spectrum of NGS Applications
The versatility of a next generation sequencing platform continues to amaze me after all these years in genomics. What started as a tool for basic research has blossomed into something that touches nearly every corner of biology and medicine.
Whole Genome Sequencing (WGS) represents the most comprehensive approach, where we sequence every single base pair in an organism’s DNA. Think of it as reading the entire instruction manual of life, rather than just specific chapters. This complete genetic blueprint has become invaluable for understanding rare genetic disorders that might affect only a handful of families worldwide. We’re also seeing WGS transform preventive medicine through initiatives like Newborn Genomic Sequencing for Early Disease Detection, where catching genetic conditions early can literally save lives. The field of Precision Medicine: Newborn Genomic Screening exemplifies how next generation sequencing platform technology is reshaping healthcare from day one.
RNA Sequencing (RNA-Seq) takes a different approach by focusing on the active parts of our genome. While DNA tells us what could happen, RNA shows us what is happening right now in our cells. This technology captures the dynamic nature of gene expression, revealing which genes are turned on or off under different conditions. It’s particularly powerful for understanding how diseases develop and how treatments might work at the molecular level.
Targeted sequencing offers a more focused approach when researchers know exactly which genetic regions interest them most. Rather than sequencing everything, this method zooms in on specific genes or the exome (the protein-coding regions that make up about 2% of our genome but contain most disease-causing mutations). It’s like having a high-powered microscope instead of a wide-angle lens—perfect for clinical diagnostics where you’re looking for mutations in known disease genes.
The cancer research field has been completely transformed by next generation sequencing platform technology. Researchers can now identify the specific mutations driving each patient’s tumor, track how cancers evolve over time, and understand why some treatments work while others don’t. This has opened the door to precision oncology, where treatments are custom to each tumor’s unique genetic fingerprint.
Microbial sequencing and metagenomics have revealed an entirely hidden world around us. Traditional methods could only identify microbes that could be grown in laboratory cultures—which turns out to be less than 1% of all microbial species. NGS allows us to sequence DNA directly from environmental samples, uncovering the vast microbial communities in our gut, soil, oceans, and even extreme environments like hot springs or deep-sea vents. This has revolutionized our understanding of the human microbiome and its role in health and disease.
In human identification, forensic scientists now use NGS to analyze degraded DNA samples that would have been impossible to work with using older methods. This technology can extract genetic information from tiny, damaged samples, helping solve cold cases and identify remains in disaster scenarios.
The applications extend far beyond human health. Reproductive health benefits from expanded carrier screening and preimplantation genetic testing, helping families make informed decisions about family planning. Agrigenomics uses NGS to improve crop yields, develop disease-resistant plants, and improve livestock breeding. HLA typing for organ transplantation has become more precise, improving donor-recipient matching for better transplant outcomes.
What excites me most is how these diverse applications all rely on the same fundamental next generation sequencing platform technology, yet each reveals completely different aspects of life. From understanding the genetic basis of rare diseases to exploring microbial communities in extreme environments, NGS has truly become the Swiss Army knife of modern biology.
Navigating the Challenges and Future of NGS
The next generation sequencing platform has transformed how we understand genetics, but like any powerful technology, it comes with its own set of problems. The good news? These challenges are driving some of the most exciting innovations in genomics today.
The biggest headache facing researchers today is what we call the data deluge. When a single human genome can generate hundreds of gigabytes of information, and large population studies produce petabytes of data, you quickly realize that storage becomes a massive problem. It’s like trying to organize every grain of sand on a beach – the sheer volume is overwhelming.
This data explosion creates what’s known as the bioinformatics bottleneck. Having all this genetic information is fantastic, but making sense of it requires sophisticated computational pipelines and skilled bioinformaticians who can interpret the results. Often, the analysis takes longer than the actual sequencing – imagine taking a photo instantly but needing weeks to develop it.
Sequencing errors remain another persistent challenge. While modern platforms are remarkably accurate, mistakes can still creep in during library preparation, amplification, or the sequencing process itself. These errors might seem small, but they can lead to false positives or missed variants that could be clinically important.
Then there’s the question of data security and privacy. Genomic data is incredibly sensitive – it’s essentially your biological blueprint. When you’re dealing with large-scale studies or clinical applications, protecting this information becomes paramount. This challenge has led to innovative solutions like Federated Architecture in Genomics and Trusted Research Environments.
Looking ahead, the future of NGS is incredibly bright. Long-read sequencing technologies are getting better every year, with some platforms now generating reads that span millions of base pairs. This is crucial for understanding complex genomic regions that shorter reads simply can’t resolve.
Single-cell sequencing is opening up entirely new frontiers. Instead of looking at the average of millions of cells, we can now peer into individual cells to understand how they differ. This is particularly exciting for cancer research, where tumor heterogeneity plays a crucial role in treatment resistance.
The integration of AI in genomics is perhaps the most transformative trend we’re seeing. Machine learning algorithms are becoming incredibly sophisticated at identifying patterns in genomic data that humans might miss. They’re helping improve variant calling accuracy, predict gene function, and even accelerate drug findy.
Cloud computing and federated architectures are solving the data storage and analysis challenges. Rather than requiring every lab to have massive computing infrastructure, researchers can leverage cloud resources and analyze data where it lives, without compromising security.
The development of automated, turnkey solutions is making NGS more accessible to smaller labs and clinics. These systems can take you from sample to results with minimal hands-on time, democratizing access to genomic technologies.
At Lifebit, we’re tackling these challenges head-on with our federated AI platform. Our Trusted Research Environment (TRE), Trusted Data Lakehouse (TDL), and R.E.A.L. (Real-time Evidence & Analytics Layer) work together to provide secure, real-time access to genomic data while ensuring compliance with strict privacy regulations. We’re helping researchers collaborate across institutions without compromising data security, turning the challenges of today’s next generation sequencing platform technologies into tomorrow’s opportunities.
The future of genomics isn’t just about better sequencing – it’s about smarter, more secure, and more collaborative ways to open up the insights hidden in our DNA.
Conclusion: Using NGS Data for Future Findies
The change from Sanger sequencing to today’s sophisticated next generation sequencing platform technologies tells one of science’s most remarkable stories. We’ve witnessed an incredible shift from spending billions of dollars and over a decade to sequence a single human genome to now completing entire genome sequences in just one day at a fraction of the cost.
This revolution has fundamentally changed how we understand life itself. Next generation sequencing platform systems have become the backbone of modern genomic medicine, enabling researchers and clinicians to dive deep into the genetic underpinnings of rare diseases, cancer, infectious diseases, and countless other conditions with unprecedented precision.
The applications span far beyond what we initially imagined. From detailed cancer research that guides personalized treatments to comprehensive microbial community analysis that reveals the hidden world of our microbiome, NGS technology touches nearly every corner of biological research. It’s helping solve crimes through human identification, improving crop yields in agriculture, and even enabling early disease detection in newborns.
But with this incredible power comes equally significant challenges. The data deluge created by these platforms is staggering – we’re talking about petabytes of information that need secure storage, careful management, and sophisticated analysis. The bioinformatics bottleneck remains real, requiring advanced computational tools and skilled professionals who can transform raw genetic data into meaningful insights that actually help patients.
Perhaps most importantly, we must handle the privacy and security of sensitive genomic data with the utmost care. People trust us with their most personal information – their genetic blueprint – and we have a responsibility to protect it.
This is exactly why we built Lifebit’s federated AI platform. We recognized that the future of genomics depends on our ability to securely manage and analyze genomic data with a federated platform while enabling collaboration across institutions and borders. Our platform brings together the Trusted Research Environment (TRE), Trusted Data Lakehouse (TDL), and R.E.A.L. (Real-time Evidence & Analytics Layer) to deliver real-time insights while maintaining the highest security standards.
Our commitment to data security isn’t just talk – it’s backed by rigorous standards, as demonstrated by our Lifebit ISO Certification for Genomic Data Security.
The future of genomics looks incredibly bright. We’re seeing continued advances in sequencing technology, the integration of powerful AI and machine learning tools, and the development of secure, collaborative data environments that enable researchers worldwide to work together safely. Long-read sequencing is getting better, single-cell analysis is revealing cellular secrets we never knew existed, and artificial intelligence is helping us make sense of the massive amounts of data we’re generating.
At Lifebit, we’re not just watching this future unfold – we’re actively building it. We’re helping researchers, pharmaceutical companies, and public health agencies open up the full potential of genomic data while ensuring it remains secure and compliant with global regulations.
The next generation sequencing platform has given us the tools. Now it’s up to us to use them wisely to improve human health and advance scientific knowledge for generations to come.

