Mastering the Omics Data Deluge
What is Omics Data Management?
Omics data management is the process of handling the vast, complex, and diverse data generated by technologies like genomics, proteomics, and metabolomics. It’s crucial for turning raw biological information into actionable insights for research and healthcare.
In simple terms, omics data management involves:
- Collecting & Storing massive datasets from various omics technologies.
- Processing raw data files into standardized, usable formats.
- Integrating different omics types for a holistic biological view.
- Analyzing complex information to uncover patterns and discoveries.
- Sharing data securely, efficiently, and ethically across organizations.
- Ensuring data meets FAIR principles (Findable, Accessible, Interoperable, Reusable).
Data is one of the most important assets for any company, especially in the life sciences. Pharmaceutical companies and research projects generate an enormous amount of omics data. This flood of information promises breakthroughs in understanding diseases and developing new treatments.
However, managing this “data deluge” is far from simple. It comes with unique challenges, from the sheer volume and complexity of the data to ensuring it can be used across different systems and shared responsibly. Effective omics data management is key to unlocking the full potential of these valuable datasets.
I’m Maria Chatzou Dunford, CEO and Co-founder of Lifebit. My background in computational biology and AI, including contributing to Nextflow, gives me a unique perspective on the complexities of omics data management and empowering precision medicine. In this guide, we’ll explore these challenges and the strategies to master them.

Omics data management glossary:
- clinical data portal
- data privacy research
- enterprise data governance solution
Solving the Primary Challenges in Omics Data Management
The transition from traditional biology to “Big Data” biology has been exhilarating, but it hasn’t come without growing pains. When we talk about omics data management, we aren’t just talking about big files; we are talking about a fundamental shift in how we approach scientific inquiry. The sheer scale of the data is staggering: a single whole-genome sequence (WGS) can generate 100-200 gigabytes of raw data. When multiplied by thousands of participants in a cohort study, we are looking at petabytes of information that must be stored, indexed, and queried.
One of the steepest hurdles is the existence of data silos. In many large pharmaceutical companies, data is scattered across different departments, often stored in incompatible formats or on isolated local servers. This fragmentation makes it nearly impossible for a researcher in oncology to benefit from insights discovered by a team in immunology, even if they are looking at the same underlying biological pathways. Breaking down these silos requires not just better hardware, but a cultural shift toward data democratization and the implementation of robust enterprise data governance solutions.
Then there is the “Curse of Dimensionality.” In multi-omics, we often face the High-Dimensional Low-Sample-Size (HDLSS) problem. This means we have thousands (or millions) of features—like gene expression levels, methylation sites, or protein abundances—but only a few hundred patient samples. This imbalance creates a massive risk of “noise” and overfitting, where AI models find patterns that aren’t actually there. To combat this, we need advanced analytics solutions for multi-omic data that utilize regularization techniques and dimensionality reduction (such as PCA or UMAP) to filter out the noise and focus on biologically relevant signals.
Furthermore, as we rely more on AI to process this data, we face the “black box” problem. If an algorithm identifies a potential drug target, we need to know why. This is why scientific research on explainable AI is so critical in biomedical data science. Without interpretability, clinical translation remains a distant dream because clinicians cannot trust a recommendation they don’t understand. We must move toward models that provide feature importance scores and biological context for their predictions.
Finally, we must address the sheer cost of storage and compute. Sequencing costs have plummeted, but the costs of managing, storing, and moving that data have not followed the same curve. Egress fees—the costs charged by cloud providers to move data out of their environment—can be astronomical. This is why we advocate for Lifebit CloudOS genomic data federation, which allows researchers to run analyses where the data resides, rather than moving petabytes of information across the globe. This “bring the code to the data” approach is the only sustainable way to manage global-scale omics projects.
Why Manual Data Access Takes 2-6 Weeks
One of the most frustrating bottlenecks in modern research is the time it takes just to get permission to look at data. Currently, Data Access Committees (DACs) typically respond to requests in two to six weeks. Why so long? It’s because of the manual evaluation of heterogeneous consent forms. Every patient sample comes with a set of rules: “You can use this for cancer research, but not for commercial gain,” or “This data cannot leave the UK.” Historically, these rules were written in plain, unstandardized text. A human has to read every single form to ensure compliance, which is a slow and error-prone process.
To fix this, we look to initiatives like the Global Alliance for Genomics and Health (GA4GH). They developed the DUO (Data Use Ontology), which allows users to tag datasets with standardized codes based on patient consent. Instead of a committee reading a 10-page legal document, a machine can instantly match a researcher’s request to the dataset’s allowed uses. For instance, a code like ‘DUO:0000007’ indicates ‘disease-specific research,’ allowing for automated filtering of requests. By adopting these standards, we move closer to the FAIR principles—ensuring data is Findable, Accessible, Interoperable, and Reusable. For more on how we implement these, see our white paper on Lifebit data standardisation.
Centralizing the Deluge: Data Lakes and Standardized Pipelines
To stop the data from overflowing, we need a better bucket. Traditional data warehouses are often too rigid for the messy, unstructured nature of raw omics data. This is where the concept of a data lake or, even better, a Trusted Data Lakehouse comes into play. A data lakehouse integrates the flexibility of a data lake with the management and performance of a data warehouse. It allows us to store raw files (like FASTQ, BAM, or VCF) alongside structured clinical data, such as Electronic Health Records (EHR) and phenotypic information.
But a lake is only useful if you know what’s in it. Effective omics data management requires rigorous metadata capture at the point of entry. Without metadata—information about the sample’s origin, the sequencing kit used, the date of extraction, and the technician involved—the data becomes a “data swamp.” For example, GSK implemented a sample request system using a single web interface entry point. This ensures that every sample comes with uniform metadata—sequencing platform, tissue type, and preparation protocol—regardless of which department generated it. This type of health data harmonization is what separates a functional research environment from a digital graveyard.
Achieving Functional Equivalence in Omics Data Management
When processing raw omics data, such as RNA-seq FASTQ files, we often encounter a hidden problem: tool variation. If two researchers use different versions of the same alignment tool (e.g., STAR or BWA), or even the same tool with slightly different parameters, they might get different results from the same raw data. This lack of reproducibility is a major hurdle for scaling genomics in clinical environments, where consistency is a regulatory requirement.
To solve this, the industry is moving toward “functional equivalence.” This means that even if different pipelines are used, they are tested to ensure the outputs are scientifically identical. We achieve this by using NF-core: standardised bioinformatics pipelines. These pipelines are built on Nextflow, which provides a domain-specific language (DSL) for writing workflows that are highly portable. These pipelines are containerized using Docker or Singularity and version-controlled, ensuring that an analysis run in London today will produce the exact same results as one run in New York next year. This level of standardization is essential for large-scale meta-analyses where data from multiple centers must be combined without introducing technical artifacts.
| Data Type | Format | Primary Use | Management Challenge |
|---|---|---|---|
| Raw Reads | FASTQ | Sequence alignment | Massive file size; storage costs |
| Aligned Reads | BAM/CRAM | Variant calling | High compute requirement for processing |
| Variants | VCF | Identifying mutations | Requires constant annotation updates |
| Expression | CSV/TSV | Quantifying gene activity | Batch effects and normalization |
| Proteomics | MZML | Protein identification | High complexity of mass spec data |
| Metabolomics | NMR/MS | Metabolic profiling | High chemical diversity and noise |
Leveraging Cloud Computing and Multi-Omics Integration
Cloud computing has been the ultimate game-changer for omics data management. Before the cloud, researchers were limited by the size of their local high-performance computing (HPC) clusters. If a project required more nodes, the institution had to purchase and install physical hardware, a process taking months. Today, we can spin up thousands of CPUs in minutes to process a new batch of genomes. This elasticity allows us to access petabyte-scale repositories like The Cancer Genome Atlas (TCGA) or the UK Biobank without needing to buy a single hard drive.
The cloud doesn’t just provide storage; it provides the scalability needed for multi-omics integration. Multi-omics is the practice of combining different “layers” of biological data—like DNA (genomics), RNA (transcriptomics), proteins (proteomics), and metabolites (metabolomics)—to see the full picture of a disease. This is critical because the genome is relatively static, while the proteome and metabolome are dynamic and reflect the current state of the organism.
A classic example is the TCGA study of stomach cancer. By using cloud-based analysis to integrate different omics types, researchers discovered that what we thought was one disease is actually four distinct molecular subtypes: EBV-infected, MSI (microsatellite instability), GS (genomically stable), and CIN (chromosomal instability). This discovery has massive implications for precision medicine, as each subtype may respond differently to chemotherapy or immunotherapy. Without a unified data management strategy, these cross-layer correlations would remain hidden.
Common Combinations in Multi-Omics Data Management
Not all omics are created equal. Depending on the disease, certain combinations are more powerful than others. According to a review of 388 multi-omics studies (2018–2021), transcriptomics is the most common omic type used, appearing in the majority of both cancer and non-cancer studies. This is because RNA-seq is relatively mature, cost-effective, and provides a high-resolution snapshot of cellular activity.
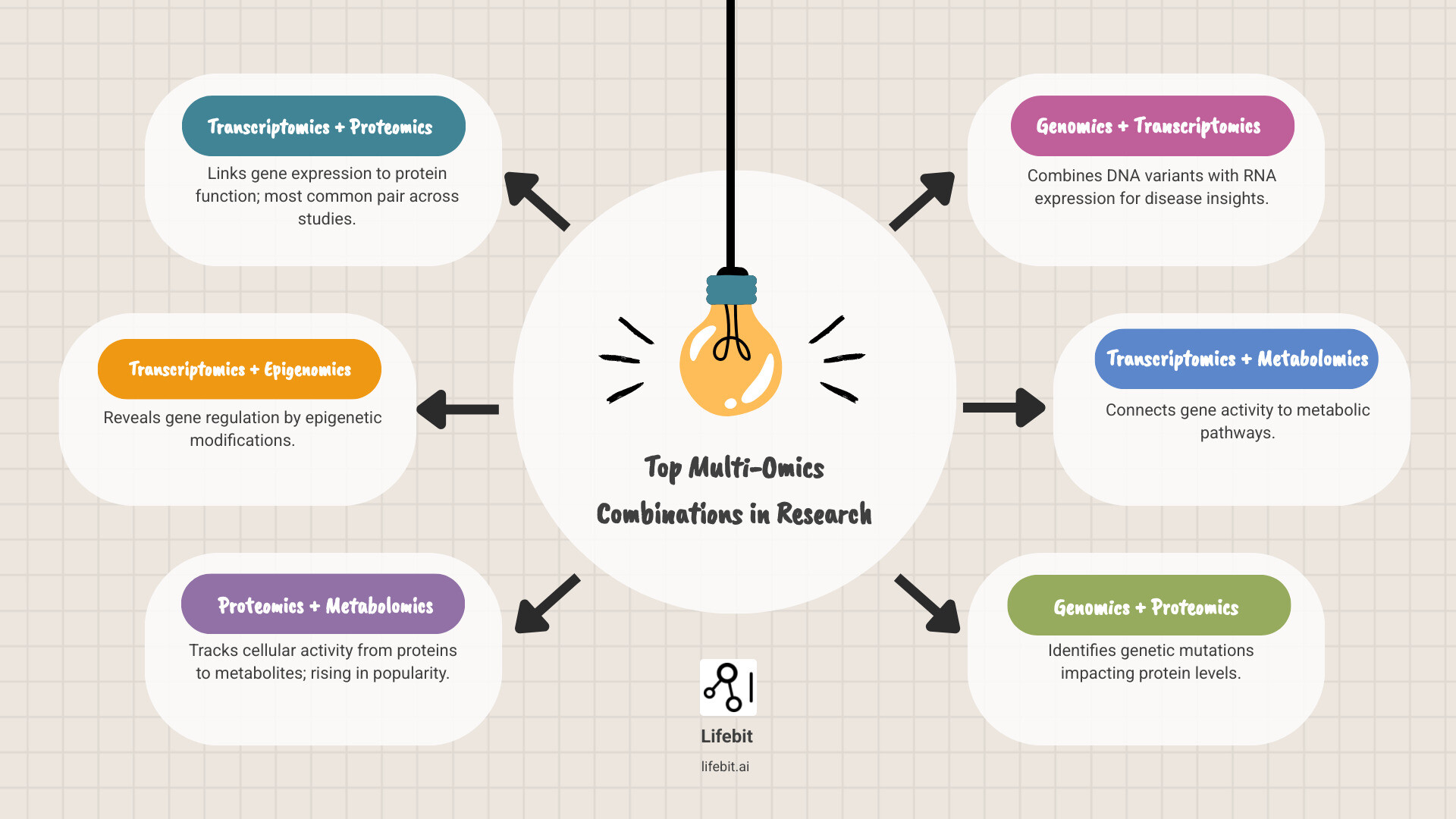
The most frequent “power couples” in the data are:
- Transcriptomics + Proteomics: The most popular pair, linking gene messages to the actual working proteins. This helps identify cases where high mRNA levels do not lead to high protein levels due to post-translational regulation.
- Transcriptomics + Epigenomics: Used to understand how gene expression is regulated by external factors, such as DNA methylation or histone modification. This is vital in fields like aging and environmental health.
- Proteomics + Metabolomics: A rising favorite for understanding the direct functional readout of cellular activity. Since proteins (enzymes) drive metabolic reactions, combining these layers provides a clear view of metabolic flux.
Among non-cancer diseases, nervous system disorders like Alzheimer’s, Parkinson’s, and ALS are the most frequently studied with multi-omics datasets. These complex conditions are rarely caused by a single genetic mutation; instead, they involve a cascade of failures across multiple biological systems. For more insights, explore our dedicated omics content portal.
Advanced Integration Methods for High-Dimensional Data
Once you have your various omics datasets, how do you actually “mash” them together? In omics data management, we generally use three strategies for integration, each with its own strengths and weaknesses:
- Early Integration (Concatenation): You simply stick all the data tables together into one giant matrix. For example, you might have 20,000 columns for genes and 500 columns for metabolites. It’s simple, but it often fails because one data type (like genomics) might overwhelm another (like metabolomics) due to scale differences. It also ignores the unique statistical distributions of different data types.
- Late Integration (Ensemble): You analyze each omic type separately—building a model for the genome, another for the proteome—and then combine the results at the end using voting or averaging. This is great for handling heterogeneity and allows for specialized pipelines, but it can miss the complex, non-linear interactions between layers that occur simultaneously.
- Intermediate Integration (Joint Modeling): This is the “Goldilocks” zone. It transforms all data types into a common mathematical space (a latent space) simultaneously. This allows the model to learn the shared variance across all omics layers while still accounting for the unique noise in each.
To do this, we use sophisticated tools. MOFA (Multi-Omics Factor Analysis) is excellent for identifying the main drivers of variation across datasets. It decomposes the data into factors that explain variation in one or more omics layers. SNF (Similarity Network Fusion) is a favorite for detecting disease-related clusters by creating a network of patient similarities for each omic type and then fusing them into a single, robust network. Meanwhile, iCluster is a robust probabilistic method for subtype identification that has been widely used in the TCGA projects.
The goal of these methods is to make multi-omics data accessible and actionable. Whether we are dealing with missing data (where a patient has a genome but no proteome) or the “curse of dimensionality,” these advanced algorithms are the engines that drive innovation in omics data analysis. As we move forward, we are seeing the rise of Deep Learning approaches, such as Variational Autoencoders (VAEs), which can learn even more complex representations of multi-omics data, though they require much larger sample sizes to be effective.
Frequently Asked Questions about Omics Data
What is the most common omics type used in studies?
Transcriptomics (the study of RNA) is the most common omic type. It acts as a bridge between the static genome and the dynamic proteome. In both cancer and non-cancer research, it is the foundational layer for almost all multi-omics studies because it captures the real-time response of a cell to its environment.
Common pairs include:
- Transcriptomics + Proteomics (Most frequent)
- Transcriptomics + Epigenomics
- Proteomics + Metabolomics
How does the Data Use Ontology (DUO) improve sharing?
DUO improves sharing by turning “legalese” into “code.” By tagging datasets with standardized terms, it allows for automated compliance checks. This can reduce the time for data access requests from weeks to just minutes, significantly accelerating research. It is a cornerstone of federated data governance, ensuring that data is used only in ways that respect the original patient consent.
Which diseases are most frequently studied with multi-omics?
Cancer remains the primary focus of multi-omics research due to its inherent genomic instability and the availability of large public datasets. However, among non-cancer diseases, nervous system diseases are the leaders. This includes:
- Alzheimer’s Disease: Integrating genomics and proteomics to find early biomarkers in cerebrospinal fluid.
- Parkinson’s Disease: Looking at metabolomics to understand mitochondrial dysfunction and oxidative stress.
- ALS (Amyotrophic Lateral Sclerosis): Large-scale resources like Answer ALS combine clinical, genomic, and transcriptomic data from iPSC-derived neurons to find new therapeutic targets.
- Pulmonary Fibrosis: Databases like Fibromine are being used for target discovery by integrating single-cell RNA-seq with clinical outcomes.
What is the difference between a Data Lake and a Data Warehouse?
A data warehouse stores highly structured data that has been processed for a specific purpose. A data lake stores data in its raw, natural format (structured or unstructured). For omics, a data lake is preferred because raw sequence files are too large and complex for traditional warehouses. A “Lakehouse” combines both, providing the storage of a lake with the analytical power of a warehouse.
Conclusion: The Future of Federated Omics
Mastering the omics data deluge is not just about having the biggest computer or the most storage. It’s about building a smarter ecosystem. The future of omics data management lies in federation—bringing the analysis to the data, rather than the other way around.
At Lifebit, we are pioneering this shift. Our federated AI platform enables secure, real-time access to global multi-omic data while maintaining strict privacy and compliance. By using components like our Trusted Research Environment (TRE) and OMOP harmonization, we help organizations turn scattered data into life-saving discoveries.
The potential of multi-omics in translational medicine is vast. From identifying new disease subtypes to predicting drug responses, the insights are there—we just need the right tools to find them. As we continue to refine our data harmonization techniques, we move closer to a world where precision healthcare is a reality for everyone.
For more on the latest trends, check out our white paper on forecasting the future of genomic management or visit our dedicated omics content portal.
Master your data today. Contact us to see how Lifebit can transform your omics data management.

