Who is Powering the Genomic Medicine Revolution?
Genomic Medicine Centre Technology Providers: How to Cut Diagnostic Time by 80%
Genomic medicine centre technology providers are the backbone of modern precision healthcare. These specialized companies supply the sequencing platforms, bioinformatics tools, AI analytics, and secure data infrastructure that enable genomic medicine centres worldwide to transform raw DNA sequences into life-saving clinical insights. The global genomics market is projected to reach $85.09 billion by 2030, up from $47.07 billion in 2025, driven by advances in next-generation sequencing and AI-powered data interpretation.
Key categories of genomic medicine centre technology providers include:
- Sequencing platform providers – Companies offering high-throughput NGS, WGS, and WES systems
- Bioinformatics and AI analytics – Software for variant calling, clinical interpretation, and automated reporting
- Data management and security – Cloud infrastructure, federated platforms, and GDPR/HIPAA-compliant storage solutions
- Specialized diagnostics – Precision oncology tools, liquid biopsy platforms, and pharmacogenomics solutions
- Population genomics infrastructure – National-scale platforms supporting initiatives like NHS Genomic Medicine Service and Genomic Medicine Sweden
These providers enable genomic medicine centres to deliver faster diagnoses for rare diseases, match patients to targeted cancer therapies, and integrate genomic insights into routine clinical care. Programs like the NHS Genomic Medicine Service now aim to offer whole genome sequencing as part of standard care for all children with cancer or serious genetic conditions—a change that would be impossible without robust technology partnerships.
The challenge for healthcare systems and research institutions is selecting partners who can deliver not just cutting-edge sequencing, but also secure data access, AI-driven interpretation, and seamless integration across siloed datasets.
I’m Dr. Maria Chatzou Dunford, CEO and Co-founder of Lifebit, where we’ve spent over a decade building federated data analysis platforms for genomic medicine centre technology providers and their healthcare partners. Our work powering secure, compliant genomic research environments across public sector institutions and pharmaceutical organizations has given me insight into what makes these technology partnerships succeed—and where they often fall short.
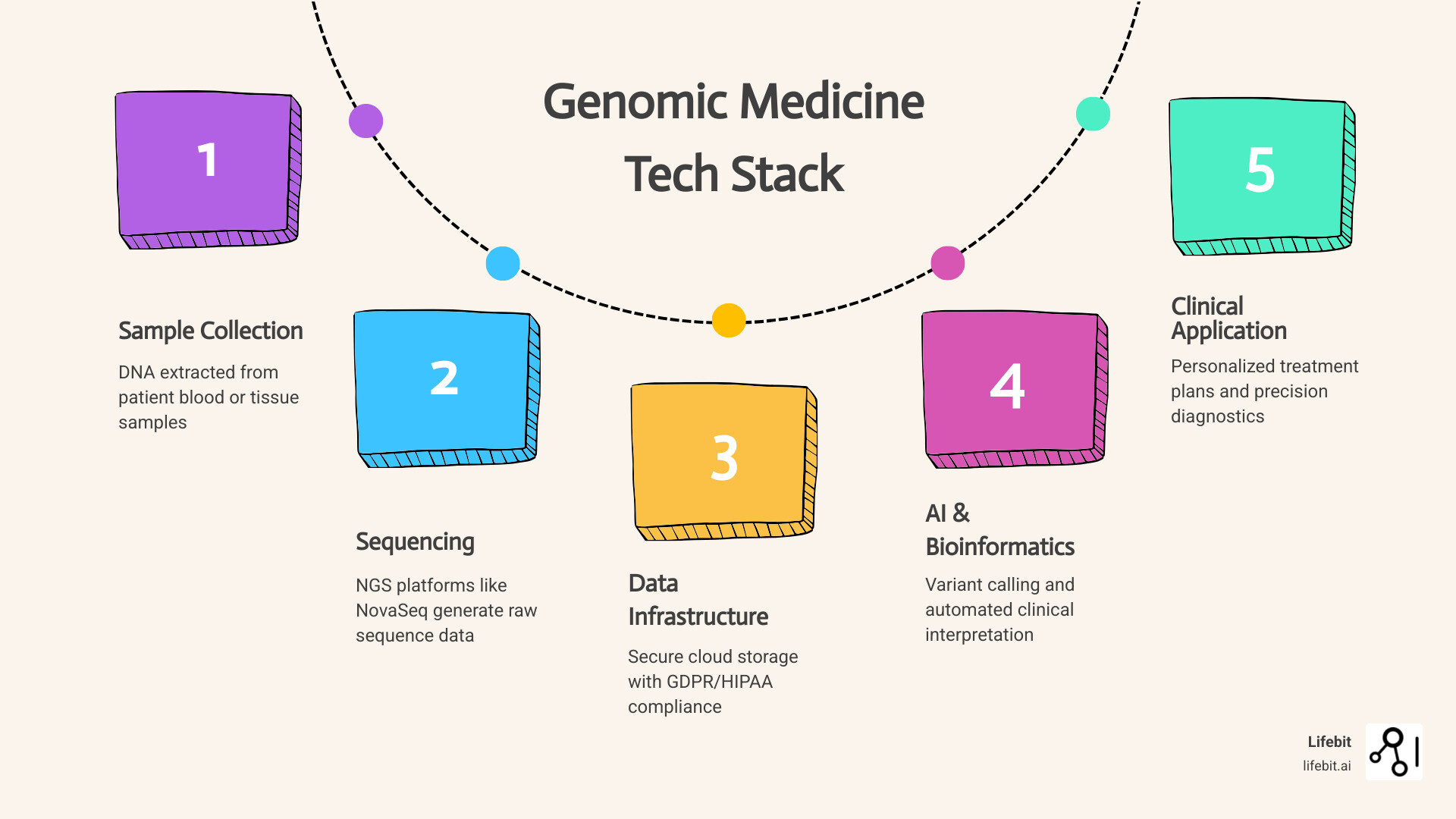
Simple guide to genomic medicine centre technology providers terms:
Scaling Precision Care: The Role of Genomic Medicine Centre Technology Providers
The transition from “bench to bedside” is no longer a slow crawl; it’s a high-speed race powered by genomics. To move from a patient’s blood sample to a personalized treatment plan, a genomic medicine centre relies on a complex ecosystem of technology. This ecosystem is not merely a collection of hardware; it is a sophisticated stack of integrated services that must operate with 99.9% uptime and clinical-grade precision.
At the heart of this revolution is the ability to read and understand genomes at scale. Whether it’s clinical genetics for pediatric rare diseases, cancer genetics for targeted oncology, or reproductive screening for expectant parents, the technology must be flawless. The NHS Genomic Medicine Service in the UK is a prime example of this in action, providing a national framework where technology providers must meet rigorous standards to serve 55 million people. This service relies on a network of Genomic Laboratory Hubs (GLHs) that utilize standardized technology stacks to ensure that a patient in Cornwall receives the same quality of genomic analysis as a patient in Newcastle.

High-Throughput NGS for Genomic Medicine Centre Technology Providers
Next-generation sequencing (NGS) is the engine room of the genomic centre. Leading technology providers offer FDA-cleared IVD NGS systems, ensuring that every “read” is accurate enough to base a medical decision on. This level of regulatory clearance is vital for clinical labs that need to maintain the highest standards of patient care. The evolution of these platforms has seen a shift from “short-read” sequencing, which is excellent for identifying small variants, to “long-read” sequencing, which can resolve complex structural variations in the genome that were previously invisible.
We see a diverse range of platforms being used in top-tier facilities. These systems allow for a variety of applications, including:
- Whole Genome Sequencing (WGS): Looking at the entire DNA blueprint, including the non-coding regions that regulate gene expression. This is increasingly used for undiagnosed rare diseases where the cause may lie outside the known exons.
- Whole Exome Sequencing (WES): Focusing on the protein-coding regions where 80% of disease-causing mutations hide. This remains a cost-effective workhorse for many clinical applications.
- Targeted Panels: Focusing on specific genes known to be associated with certain cancers or inherited conditions, such as the BRCA1/2 genes for breast cancer risk or panels for hereditary cardiomyopathies.
- RNA-Seq (Transcriptomics): Measuring gene expression to understand how the genome is actually behaving in real-time, which is crucial for understanding tumor microenvironments.
Choosing the right next-generation sequencing platform depends on the centre’s specific needs—balancing cost, speed, and the depth of data required. For instance, a rapid-turnaround NICU setting requires different technology than a large-scale population health study.
| Feature | Whole Genome Sequencing (WGS) | Whole Exome Sequencing (WES) |
|---|---|---|
| Coverage | Entire genome (coding & non-coding) | Coding regions (exons) only |
| Diagnostic Yield | Highest (detects structural variants) | High for known genetic disorders |
| Data Size | Massive (~100-120 GB per sample) | Moderate (~5-10 GB per sample) |
| Typical Use Case | Rare disease, complex research | Clinical diagnostics, targeted screening |
| Cost per Sample | Higher ($600 – $1,000+) | Lower ($200 – $400) |
Secure Data Management for Genomic Medicine Centre Technology Providers
As we generate more data, we face a massive “data gravity” problem. You can’t simply email a whole genome; a single WGS file can be over 100GB. This is where federated technology in population genomics becomes critical. Instead of moving massive, sensitive datasets to the researchers (which risks security breaches and violates GDPR), we move the analysis to the data. This “data-centric” approach is the only way to scale genomic medicine globally.
Security isn’t just a checkbox; it’s the foundation of patient trust. Technology providers must ensure:
- GDPR and HIPAA Compliance: Protecting patient privacy across borders. This includes implementing “Right to be Forgotten” protocols and strict audit trails for every data access event.
- Data Harmonization: Ensuring that data from a lab in Singapore can be compared with data from a lab in New York. This requires adherence to GA4GH (Global Alliance for Genomics and Health) standards for data formats and API protocols.
- Cybersecurity: Implementing robust encryption (both at rest and in transit) and multi-factor authentication to prevent unauthorized access to the most personal information a human possesses—their DNA.
- Interoperability with EHRs: The genomic data must flow seamlessly into Electronic Health Records (EHRs) like Epic or Cerner so that a primary care physician can see a pharmacogenomic alert when prescribing medication.
Our Genomic Medicine Accessibility Report highlights that the biggest bottleneck in precision medicine today isn’t the sequencing—it’s the secure, compliant access to the data once it’s been generated. Without a robust data infrastructure, a genomic medicine centre is just a library where the books are locked in separate, unreadable rooms.
AI and Bioinformatics: Turning Raw Data into Clinical Insights
Raw DNA data is just a long string of A, C, T, and G. Without AI for genomics, it’s virtually useless in a clinical setting. Advanced bioinformatics tools act as the “translator,” identifying which variants are benign and which are pathogenic. The sheer volume of data—3 billion base pairs per person—makes manual interpretation impossible.
The Bioinformatics Pipeline: From FastQ to VCF
The process begins with primary analysis (base calling on the sequencer), followed by secondary analysis. This involves aligning the millions of short DNA fragments to a reference genome and “variant calling”—identifying where the patient’s DNA differs from the norm. Technology providers now offer “accelerated” pipelines using FPGAs or GPUs that can process a whole genome in under 30 minutes, a task that used to take days.
Providers are now integrating machine learning to speed up tertiary analysis—the clinical interpretation. In the Genomics England research environment, researchers use advanced tools to sift through thousands of variants to find the one responsible for a child’s rare disease. This automated interpretation reduces the “diagnostic odyssey” from years to just days. AI models like DeepVariant use image-recognition techniques to identify genetic variants with higher accuracy than traditional statistical methods.
Multi-Omics and the Holistic View
The best AI for genomics doesn’t just look at DNA in isolation. It incorporates multi-omics—looking at RNA (transcriptomics), proteins (proteomics), and metabolites (metabolomics)—to provide a holistic view of a patient’s health. For example, a genetic variant might suggest a risk for a disease, but proteomics can confirm if that disease is actually active in the body. This is particularly transformative in “pharmacogenomics,” where AI can predict if a patient will have a bad reaction to a specific drug before they ever take it, by analyzing how their specific genetic makeup affects drug metabolism enzymes like the CYP450 family.
Furthermore, Large Language Models (LLMs) are being adapted to “read” the vast scientific literature, automatically linking a newly discovered variant in a patient to the latest research paper published only hours earlier. This ensures that clinicians are always working with the most current knowledge available.
Specialized Solutions for Oncology and Rare Disease Diagnosis
Precision oncology is perhaps the most visible success story for genomic medicine centre technology providers. Comprehensive genomic profiling (CGP) allows doctors to see the exact mutations driving a tumor’s growth. The CGP market, valued at $9.9 billion in 2022, is exploding as we move toward liquid biopsies. Unlike traditional tissue biopsies, which are invasive and may not capture the full heterogeneity of a tumor, CGP provides a comprehensive molecular map.
Liquid Biopsy: The Frontier of Monitoring
Liquid biopsies analyze circulating tumor DNA (ctDNA) from a simple blood draw. This is a game-changer for patients who cannot undergo invasive tissue biopsies or for monitoring treatment response in real-time. Technology providers are developing ultra-sensitive assays that can detect a single cancer-related mutation among thousands of healthy DNA fragments. This allows for the detection of “minimal residual disease” (MRD), identifying if a cancer is returning months before it would show up on a traditional CT scan.
Technology providers are also helping clinicians steer the NHS National Genomic Test Directory, which specifies which tests are funded for specific clinical indications. This directory is a living document, constantly updated as new evidence emerges, and technology providers must ensure their assays are validated against these national standards.
Ending the Diagnostic Odyssey in Rare Disease
For rare diseases, tools like the Genomics England PanelApp allow the global scientific community to collaborate on defining which genes should be tested for specific conditions. This crowdsourced expertise ensures that rare disease diagnosis genomics stays at the absolute cutting edge.
In many cases, children with undiagnosed developmental delays undergo years of testing—the “diagnostic odyssey.” By implementing WGS early in the clinical pathway, technology providers are helping centres achieve diagnostic yields of 30-40%, providing answers to families who have often spent a decade searching for a cause. These diagnoses often lead to immediate changes in clinical management, such as starting a specific diet, avoiding certain medications, or qualifying for a clinical trial for a novel gene therapy.
Future Trends: Population Genomics and Real-Time Analytics
We are moving away from small, isolated studies toward massive, population-scale initiatives. Whether it’s the Population Genomics Platform Hong Kong Genome project, the All of Us Research Program in the US, or the collaborative efforts of Genomic Medicine Sweden, the goal is the same: sequence hundreds of thousands of individuals to understand the health of an entire nation. These projects require technology providers to build “Trusted Research Environments” (TREs) where data can be analyzed without being downloaded.
The Shift to Preventive Healthcare
The ultimate goal of these large-scale projects is to move from reactive medicine (treating the sick) to preventive medicine (keeping people healthy). By calculating Polygenic Risk Scores (PRS), technology providers can help healthcare systems identify individuals at high risk for common diseases like Type 2 diabetes or coronary artery disease long before symptoms appear. This allows for targeted interventions, such as increased screening or lifestyle changes, which can significantly reduce the long-term burden on healthcare budgets.
Real-Time Evidence and Global Collaboration
The future lies in “Real-Time Evidence.” Imagine a world where a new virus emerges, and within hours, omics data from around the world is analyzed via a Latin America genomics research partnership to identify the most effective vaccine targets. This requires a level of EHDS (European Health Data Space) compliance and secure data sharing that we are only just beginning to realize.
Technology providers are now focusing on “federated learning,” where AI models are trained across multiple hospitals without the patient data ever leaving the hospital’s firewall. This preserves privacy while allowing the AI to learn from a much more diverse dataset, reducing the bias that often occurs when models are trained on limited, non-representative populations. This is essential for ensuring that the benefits of genomic medicine are equitable across all ethnicities and backgrounds.
Frequently Asked Questions about Genomic Technology
How do providers ensure regulatory compliance?
Technology providers must steer a complex web of regulations. This includes obtaining FDA clearance in the US and CE-IVD marking in Europe under the new In Vitro Diagnostic Regulation (IVDR). They also adhere to ISO 15189 standards for medical laboratories and ISO 27001 for information security management. For a deep dive into how national services manage this, see our Genomics England complete guide. Compliance is an ongoing process involving regular audits and rigorous validation of every software update.
What is the difference between WGS and WES in clinical settings?
Whole Exome Sequencing (WES) is often the first choice because it is more cost-effective and focuses on the coding regions where most known mutations occur. However, Whole Genome Sequencing (WGS) is becoming the gold standard for undiagnosed rare diseases because it can catch structural variants, copy number variations, and intronic mutations that WES misses. Furthermore, WGS provides more uniform coverage and doesn’t suffer from the “capture bias” inherent in WES. You can compare various DNA sequencing methods to see which fits your lab’s budget and goals.
How does AI improve genomic data interpretation?
AI removes the human bottleneck. Instead of a geneticist manually reviewing thousands of variants, AI algorithms prioritize the most likely candidates based on massive global databases like ClinVar and gnomAD. AI can also predict the functional impact of a variant on protein folding or splicing. This increases speed, reduces errors, and is the primary tool for finally ending the diagnostic odyssey for families with rare diseases. It allows a single clinician to process ten times as many cases as they could five years ago.
What is the role of cloud computing in genomic medicine?
Cloud computing provides the elastic scalability required to process bursts of genomic data. While some centres prefer on-premise storage for perceived security, the major cloud providers (AWS, Azure, Google Cloud) now offer specialized “Health Omics” services that meet the highest security standards. Cloud-native solutions allow for easier collaboration between international research teams and provide the massive compute power needed for AI training and complex bioinformatics pipelines without the need for expensive, depreciating hardware on-site.
How is genomic data integrated into clinical workflows?
Integration is achieved through standardized APIs and HL7 FHIR (Fast Healthcare Interoperability Resources) protocols. This ensures that genomic findings are not just a PDF report buried in a file, but actionable data points that can trigger automated alerts in a clinician’s workflow. For example, if a patient is prescribed Abacavir, the system can automatically check for the HLA-B*5701 variant to prevent a potentially fatal hypersensitivity reaction.
Conclusion: Choosing the Right Partner for Precision Health
The “Genomic Revolution” isn’t a future event—it’s happening right now in clinics and hospitals across the globe. But the revolution is only as strong as the technology powering it.
At Lifebit, we believe that the key to unlocking the power of genomics isn’t just more data—it’s more accessible and secure data. Our federated AI platform provides the “Airlock” that allows researchers to work with sensitive biomedical data without it ever leaving its secure home. By providing a Trusted Research Environment (TRE) and a Trusted Data Lakehouse (TDL), we enable the real-time insights and secure collaboration that modern medicine demands.
If you are looking to scale your genomic initiatives while maintaining the highest standards of security and compliance, let’s talk. We are proud to power the next generation of Lifebit Federated Biomedical Data Platform solutions that are truly reshaping healthcare for everyone.

